Abstract
Accurate quantitative methods are needed to determine the amount of transgenic material in ingredients and comply with labelling GMO thresholds. Quantitative real-time PCR methods are usually applied for GMO quantification, but since a few years, digital PCR (dPCR) has been described as a potential alternative by quantifying DNA molecules directly without any standard curves. In this study, the performance of dPCR to quantify 9 GM-soya events and 15 GM-maize events was assessed. Following GMO validation guidelines, the trueness and precision were determined on high, medium and low levels of transgenic content. Results showed biases below ± 25% and satisfactory precision data. Limits of quantification were determined for each GM-event and were between 12 and 31 target copies. The reliability of GMO quantification by dPCR was further confirmed by analysing several proficiency test samples. Overall, dPCR showed accurate and precise GMO quantification on all the tested GM-events, from high to low transgenic amount. With its ease-of-use, dPCR was found to be an appealing alternative technology for routine GMO testing laboratories.

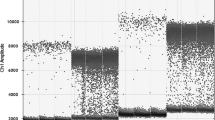
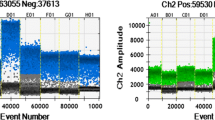
Graphical abstract
Similar content being viewed by others
Abbreviations
- GMO:
-
Genetically modified organisms
- PCR:
-
Polymerase chain reaction
- dPCR:
-
Digital PCR
- CRM:
-
Certified reference material
- NTC:
-
No template control
- LOQ:
-
Limit of quantification
References
Cottenet G, Blancpain C, Sonnard V, Chuah PF. Two FAST multiplex real-time PCR reactions to assess the presence of genetically modified organisms in food. Food Chem. 2019;274:760–5.
Cottenet G, Blancpain C, Sonnard V, Chuah PF. Development and validation of a multiplex real-time PCR method to simultaneously detect 47 targets for the identification of genetically modified organisms. Anal Bioanal Chem. 2013;405:6831–44.
Bonfini L, Van Den Bulcke MH, Mazzara M, Ben E, Patak A. GMOMETHODS: the European Union database of reference methods for GMO analysis. J AOAC Int. 2012;95:1713–9.
Quan PL, Sauzade M, Brouzes E. dPCR : a technology review. Sensors. 2018. https://doi.org/10.3390/s18041271.
Ren J, Deng T, Huang W, Chen Y, Ge Y. A digital PCR method for identifying and quantifying adulteration of meat species in raw and processed food. PLoS One. 2017. https://doi.org/10.1371/journal.pone.0173567.
Corbisier P, Bhat S, Partis L, Xie VR, Emslie KR. Absolute quantification of genetically modified MON810 maize (Zea mays L.) by digital polymerase chain reaction. Anal Bioanal Chem. 2010;396:2143–50.
Iwobi A, Gerdes L, Busch U, Pecoraro S. Droplet digital PCR for routine analysis of genetically modified organisms (GMO) – a comparison with real-time quantitative PCR. Food Control. 2016;69:205–13.
Köppel R, Bucher T. Rapid establishment of droplet digital PCR for quantitative GMO analysis. Eur Food Res Technol. 2015;241:427–39.
Whale AS, Huggett JF, Tzonev S. Fundamentals of multiplexing with digital PCR. Biomol Detect Quantif. 2016;10:15–23.
Dobnik D, Spilsberg B, Kosir AB, Holst-Jensen A, Zel J. Multiplex quantification of 12 European Union authorized genetically modified maize lines with droplet digital polymerase chain reaction. Anal Chem. 2015;87:8218–26.
Kosir AB, Spilsberg B, Holst-Jensen A, Zel J, Dobnik D. Development and inter-laboratory assessment of droplet digital PCR assays for multiplex quantification of 15 genetically modified soybean lines. Sci Rep. 2017;7:8601.
Köppel R, Bucher T, Bär D, Van Velsen F, Ganeshan A. Validation of 13 duplex droplet digital PCR systems for quantitative GMO analysis of most prevalent GMO traits. Eur Food Res Technol. 2017;244:313–21.
GMOMETHODS. EU Database of Reference Methods for GMO Analysis. 2018. http://gmo-crl.jrc.ec.europa.eu/gmomethods/. Accessed 16 February 2018.
Demeke T, Dobnik D. Critical assessment of digital PCR for the detection and quantification of genetically modified organisms. Anal Bioanal Chem. 2018;410:4039–50.
Jacchia S, Kagkli D-M, Lievens A, Angers-Loustau A, Savini C, Emons H, et al. Identification of single target taxon-specific reference assays for the most commonly genetically transformed crops using digital droplet PCR. Food Control. 2018;93:191–200.
ERM, European reference materials. Application note 4: use of certified reference materials for the quantification of GMO in food and feed. 2006. https://ec.europa.eu/jrc/sites/jrcsh/files/erm_application_note_4_en.pdf Accessed on 26 January 2018.
ENGL, European network of GMO laboratories. Definition of minimum performance requirements for analytical methods of GMO testing. 2015. http://gmo-crl.jrc.ec.europa.eu/doc/MPR%20Report%20Application%2020_10_2015.pdf Accessed on 26 January 2018.
Corbisier P, Barbante A, Berben G, Broothaerts W, De Loose M, Emons H, Georgieva TZ, Lievens A, Mazzara M, Papazova N, Perri E, Sowa S, Stebih D, Terzi V, Trapmann S. Recommendation for the unit of measurement and the measuring system to report traceable and comparable results expressing GM content in accordance with EU legislation. EUR28536 EN. 2017; https://doi.org/10.2760/177516
Arumuganathan K, Earle ED. Nuclear content of some important plant species. Plant Mol Biol Rep. 1991;9:208–18.
Majumdar N, Wessel T, Marks J. Digital PCR modeling for maximal sensitivity, dynamic range and measurement precision. PLOSone. 2015. https://doi.org/10.1371/journal.pone.0118833.
Dobnik D, Demsar T, Huber I, Gerdes L, Broeders S, Roosens N, et al. Inter-laboratory analysis of selected genetically modified plant reference materials with digital PCR. Anal Bioanal Chem. 2018;410:211–21.
Chaouachi M, Berard A, Saïd K. Relative quantification in seed GMO analysis: state of art and bottlenecks. Transgenic Res. 2013;22:461–76.
Acknowledgements
The authors would like to thank Pia Scheu and Cyril Dubuck from Bio-Rad for their technical and scientific support during our study.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Cottenet, G., Blancpain, C. & Chuah, P.F. Performance assessment of digital PCR for the quantification of GM-maize and GM-soya events. Anal Bioanal Chem 411, 2461–2469 (2019). https://doi.org/10.1007/s00216-019-01692-7
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00216-019-01692-7