Abstract
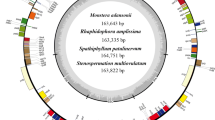
Eugenia uniflora is a plant native to tropical America that holds great ecological and economic importance. The complete chloroplast (cp) genome sequence of Eugenia uniflora, a member of the Neotropical Myrtaceae family, is reported here. The genome is 158,445 bp in length and exhibits a typical quadripartite structure of the large (LSC, 87,459 bp) and small (SSC, 18,318 bp) single-copy regions, separated by a pair of inverted repeats (IRs, 26,334 bp). It contains 111 unique genes, including 77 protein-coding genes, 30 tRNAs and 4 rRNAs. The genome structure, gene order, GC content and codon usage are similar to the typical angiosperm cp genomes. Comparison of the entire cp genomes of E. uniflora L. and three other Myrtaceae revealed an expansion of 43 bp in the intergenic spacer located between the IRA/large single-copy (LSC) border and the first gene of LSC region. Simple sequence repeat (SSR) analysis revealed that most SSRs are AT rich, which contribute to the overall AT richness of the cp genome. Additionally, fewer SSRs are distributed in the protein-coding sequences compared to the noncoding regions. Phylogenetic analysis among 58 species based on 57 cp genes demonstrated a closer relationship between E. uniflora L. and Syzygium cumini (L). Skeels compared to the Eucalyptus clade in the Myrtaceae family. The complete cp genome sequence of E. uniflora reported here has importance for population genetics, as well as phylogenetic and evolutionary studies in this species and other Myrtaceae species from Neotropical regions.




Similar content being viewed by others
References
Altschul SF, Gish W, Miller W, Myers EW, Lipman DJ (1990) Basic local alignment search tool. J Molec Biol 215:403–410. doi:10.1016/S0022-2836(05)80360-2
Asif H, Khan A, Iqbal A, Khan IA, Heinze B, Azim MK (2013) The chloroplast genome sequence of Syzygium cumini (L.) and its relationship with other angiosperms. Tree Genet Genomes 9:867–877. doi:10.1007/s11295-013-0604-1
Bayly MJ, Rigault P, Spokevicius A, Ladiges PY, Ades PK, Anderson C, Bossinger G, Merchant A, Udovicic F, Woodrow IE, Tibbits J (2013) Chloroplast genome analysis of Australian eucalypts—Eucalyptus, Corymbia, Angophora, Allosyncarpia and Stockwellia (Myrtaceae). Molec Phylogen Evol 69:704–716. doi:10.1016/j.ympev.2013.07.006
Benson G (1999) Tandem repeats finder: a program to analyze DNA sequences. Nucl Acids Res 27:573–580. doi:10.1093/nar/27.2.573
Berger BA, Kriebel R, Spalink D, Sytsma KJ (2016) Divergence times, historical biogeography, and shifts in speciation rates of Myrtales. Molec Phylogen Evol 95:116–136. doi:10.1016/j.ympev.2015.10.001
Biffin E, Lucas EJ, Craven LA, Da Costa IR, Harrington MG, Crisp MD (2010) Evolution of exceptional species richness among lineages of fleshy-fruited Myrtaceae. Ann Bot (Oxford) 106:79–93. doi:10.1093/aob/mcq088
De Almeida DJ, Faria MV, Da Silva PR (2012) Biologia experimental em Pitangueira: uma revisão de cinco décadas de publicações científicas/Experimental biology in pitangueira: a review of five decades of scientific publications. Rev Ambiência 8:159–175. doi:10.5777/ambiencia.2012.01.02rb
Dong W, Liu J, Yu J, Wang L, Zhou S (2012) Highly variable chloroplast markers for evaluating plant phylogeny at low taxonomic levels and for DNA barcoding. PLOS One 7:e35071. doi:10.1371/journal.pone.0035071
Doyle J, Doyle J (1990) Isolation of plant DNA from fresh tissue. Focus (Madison) 12:13–15
Ferreira-Ramos R, Laborda PR, De Oliveira Santos M, Mayor MS, Mestriner MA, De Souza AP, Alzate-Marin AL (2008) Genetic analysis of forest species Eugenia uniflora L. through of newly developed SSR markers. Conservation Genet 9:1281–1285. doi:10.1007/s10592-007-9458-0
Frazer KA, Pachter L, Poliakov A, Rubin EM, Dubchak I (2004) VISTA: computational tools for comparative genomics. Nucl Acids Res 32(Web Server issue): W273–W279. doi:10.1093/nar/gkh458
Govaerts R, Sobral M, Ashton P, Barrie F, Holst BK, Landrum LL, Matsumoto K, Mazine FF, Lughadha EN, Proenca C, Soares-Silva LH, Wilson PG, Lucas E (2015) World checklist of Myrtaceae. Royal Botanic Gardens, Kew
Ibrahim RIH, Azuma J-I, Sakamoto M (2006) Complete nucleotide sequence of the cotton (Gossypium barbadense L.) chloroplast genome with a comparative analysis of sequences among 9 dicot plants. Genes Genet Syst 81:311–321
Jansen RK, Cai Z, Raubeson LA, Daniell H, Depamphilis CW, Leebens-Mack J, Müller KF, Guisinger-Bellian M, Haberle RC, Hansen AK, Chumley TW, Lee S-B, Peery R, McNeal JR, Kuehl JV, Boore JL (2007) Analysis of 81 genes from 64 plastid genomes resolves relationships in angiosperms and identifies genome-scale evolutionary patterns. Proc Natl Acad Sci USA 104:19369–19374. doi:10.1073/pnas.0709121104
Kurtz S, Choudhuri JV, Ohlebusch E, Schleiermacher C, Stoye J, Giegerich R (2001) REPuter: the manifold applications of repeat analysis on a genomic scale. Nucl Acids Res 29:4633–4642. doi:10.1093/nar/29.22.4633
Langmead B (2010) Aligning short sequencing reads with Bowtie. Curr Protoc Bioinf 32:11.7.1–11.7.14. doi:10.1002/0471250953.bi1107s32
Leebens-Mack J, Raubeson LA, Cui L, Kuehl JV, Fourcade MH, Chumley TW, Boore JL, Jansen RK, depamphilis CW (2005) Identifying the basal angiosperm node in chloroplast genome phylogenies: sampling one’s way out of the Felsenstein zone. Molec Biol Evol 22:1948–1963. doi:10.1093/molbev/msi191
Leseberg CH, Duvall MR (2009) The complete chloroplast genome of coix lacryma-jobi and a comparative molecular evolutionary analysis of plastomes in cereals. J Molec Evol 69:311–318. doi:10.1007/s00239-009-9275-9
Lim TK (2012) Eugenia uniflora. In: Lim TK, Edible Medicinal and Non Medicinal Plants, vol. 3, Fruits. Springer, Netherlands, pp 620–630. doi:10.1007/978-94-007-2534-8_85
Lohse M, Drechsel O, Kahlau S, Bock R (2013) OrganellarGenomeDRAW–a suite of tools for generating physical maps of plastid and mitochondrial genomes and visualizing expression data sets. Nucl Acids Res 41(Web Server issue):W575–W581. doi:10.1093/nar/gkt289
Lucas EJ, Harris SA, Mazine FF, Belsham SR, Nic Lughadha EM, Telford A, Gasson PE, Chase MW (2007) Suprageneric phylogenetics of Myrteae, the generically richest tribe in Myrtaceae (Myrtales). Taxon 56:1105–1128. doi:10.2307/25065906
Machado LO, Vieira LD, Stefenon VM, Pedrosa OF, De Souza EM, Guerra MP, Nodari RO (2017) Phylogenomic relationship of feijoa (Acca sellowiana (O.Berg) Burret) with other Myrtaceae based on complete chloroplast genome sequences. Genetica 145:1–12. doi:10.1007/s10709-017-9954-1
Margis R, Felix D, Caldas JF, Salgueiro F, De Araujo DSD, Breyne P, Van Montagu M, De Oliveira D, Margis-Pinheiro M (2002) Genetic differentiation among three neighboring Brazil-cherry (Eugenia uniflora L.) populations within the Brazilian Atlantic rain forest. Biodivers & Conservation 11:149–163. doi:10.1023/A:1014028026273
Mazine FF, Souza VC, Sobral M, Forest F, Lucas E (2014) A preliminary phylogenetic analysis of Eugenia (Myrtaceae: Myrteae), with a focus on Neotropical species. Kew Bull 69:1–14. doi:10.1007/s12225-014-9497-x
Moore MJ, Soltis PS, Bell CD, Burleigh JG, Soltis DE (2010) Phylogenetic analysis of 83 plastid genes further resolves the early diversification of eudicots. Proc Natl Acad Sci USA 107:4623–4628. doi:10.1073/pnas.0907801107
Pennington RT, Lavin M, Oliveira-Filho A (2009) Woody Plant Diversity, Evolution, and Ecology in the Tropics: perspectives from Seasonally Dry Tropical Forests. Annual Rev Ecol Evol Syst 40:437–457. doi:10.1146/annurev.ecolsys.110308.120327
Posada D, Crandall KA (1998) MODELTEST: testing the model of DNA substitution. Bioinformatics 14:817–818. doi:10.1093/bioinformatics/14.9.817
Provan J, Powell W, Hollingsworth PM (2001) Chloroplast microsatellites: new tools for studies in plant ecology and evolution. Trends Ecol Evol 16:142–147. doi:10.1016/S0169-5347(00)02097-8
Ravi V, Khurana JP, Tyagi AK, Khurana P (2008) An update on chloroplast genomes. Pl Syst Evol 271:101–122. doi:10.1007/s00606-007-0608-0
Reginato M, Neubig KM, Majure LC, Michelangeli FA (2016) The first complete plastid genomes of Melastomataceae are highly structurally conserved. PeerJ 4:e2715. doi:10.7717/peerj.2715
Rohde W, Gramstat A, Schmitz J, Tacke E, Prufer D (1994) Plant viruses as model systems for the study of non-canonical translation mechanisms in higher plants. J Gen Virol 75:2141–2149. doi:10.1099/0022-1317-75-9-2141
Ronquist F, Huelsenbeck JP (2003) MrBayes 3: Bayesian phylogenetic inference under mixed models. Bioinformatics 19:1572–1574. doi:10.1093/bioinformatics/btg180
Ruhfel BR, Gitzendanner MA, Soltis PS, Soltis DE, Burleigh JG (2014) From algae to angiosperms - inferring the phylogeny of green plants (Viridiplantae) from 360 plastid genomes. BMC Evol Biol 14:23. doi:10.1186/1471-2148-14-23
Salgueiro F, Felix D, Caldas JF, Margis-Pinheiro M, Margis R (2004) Even population differentiation for maternal and biparental gene markers in Eugenia uniflora, a widely distributed species from the Brazilian coastal Atlantic rain forest. Diversity & Distrib 10:201–210. doi:10.1111/j.1366-9516.2004.00078.x
Sasaki T (2003) Identification of RNA editing sites in chloroplast transcripts from the maternal and paternal progenitors of tobacco (Nicotiana tabacum): comparative analysis shows the involvement of distinct trans-factors for ndhB editing. Molec Biol Evol 20:1028–1035. doi:10.1093/molbev/msg098
Shinozaki K, Ohme M, Tanaka M, Wakasugi T, Hayashida N, Matsubayashi T, Zaita N, Chunwongse J, Obokata J, Yamaguchi-Shinozaki K, Ohto C, Torazawa K, Meng BY, Sugita M, Deno H, Kamogashira T, Yamada K, Kusuda J, Takaiwa F, Kato A, Tohdoh N, Shimada H, Sugiura M (1986) The complete nucleotide sequence of the tobacco chloroplast genome: its gene organization and expression. EMBO J 5:2043–2049
Simpson JT, Wong K, Jackman SD, Schein JE, Jones SJM, Birol I (2009) ABySS: a parallel assembler for short read sequence data. Genome Res 19:1117–1123. doi:10.1101/gr.089532.108
Spada PDS, De Souza GGN, Bortolini GV, Henriques JAP, Salvador M (2008) Antioxidant, mutagenic, and antimutagenic activity of frozen fruits. J Med Food 11:144–151. doi:10.1089/jmf.2007.598
Stamatakis A (2014) RAxML version 8: A tool for phylogenetic analysis and post-analysis of large phylogenies. Bioinformatics 30:1312–1313. doi:10.1093/bioinformatics/btu033
Steane DA (2005) Complete nucleotide sequence of the chloroplast genome from the Tasmanian blue gum, Eucalyptus globulus (Myrtaceae). DNA Res 12:215–220. doi:10.1093/dnares/dsi006
Straub SCK, Fishbein M, Livshultz T, Foster Z, Parks M, Weitemier K, Cronn RC, Liston A (2011) Building a model: developing genomic resources for common milkweed (Asclepias syriaca) with low coverage genome sequencing. BMC Genomics 12:211. doi:10.1186/1471-2164-12-211
Su H-J, Hogenhout SA, Al-Sadi AM, Kuo C-H (2014) Complete Chloroplast Genome Sequence of Omani Lime (Citrus aurantiifolia) and Comparative Analysis within the Rosids. PLOS One 9:e113049. doi:10.1371/journal.pone.0113049
Tamura K, Dudley J, Nei M, Kumar S (2007) MEGA4: molecular evolutionary genetics analysis (MEGA) software version 4.0. Molec Biol Evol 24:1596–1599. doi:10.1093/molbev/msm092
Tangphatsornruang S, Sangsrakru D, Chanprasert J, Uthaipaisanwong P, Yoocha T, Jomchai N, Tragoonrung S (2010) The chloroplast genome sequence of mungbean (Vigna radiata) determined by high-throughput pyrosequencing: structural organization and phylogenetic relationships. DNA Res 17:11–22. doi:10.1093/dnares/dsp025
Thornhill AH, Ho SYW, Külheim C, Crisp MD (2015) Interpreting the modern distribution of Myrtaceae using a dated molecular phylogeny. Molec Phylogen Evol 93:29–43. doi:10.1016/j.ympev.2015.07.007
Wilson PG, O’Brien MM, Heslewood MM, Quinn CJ (2005) Relationships within Myrtaceae sensu lato based on a matK phylogeny. Pl Syst Evol 251:3–19. doi:10.1007/s00606-004-0162-y
Wolfe KH, Li WH, Sharp PM (1987) Rates of nucleotide substitution vary greatly among plant mitochondrial, chloroplast, and nuclear DNAs. Proc Natl Acad Sci USA 84:9054–9058. doi:10.1073/pnas.84.24.9054
Wyman SK, Jansen RK, Boore JL (2004) Automatic annotation of organellar genomes with DOGMA. Bioinformatics 20:3252–3255. doi:10.1093/bioinformatics/bth352
Acknowledgements
We would like to thank Prof. Andreia Turchetto Zolet for providing very helpful suggestions to our manuscript and Steven Clipman for correcting the English.
Funding
This study was carried out with the support of FAPERGS and the Conselho Nacional de Desenvolvimento Científico e Tecnológico (CNPq).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Handling editor: Marcus Koch.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Information on Electronic Supplementary material
Information on Electronic Supplementary material
Online Resource 1 List of accession numbers of the chloroplast genome sequences used to filter reads from cp genomes.
Online Resource 2 List of the 57 genes used in the ML and Bayesian phylogenetic analysis.
Online Resource 3 List of plastome sequences of Rosids included in the ML and Bayesian phylogenetic analyses.
Online Resource 4 Phylogenetic tree among 58 eurosids based on 57 protein-coding genes reconstructed by the maximum likelihood method.
Rights and permissions
About this article
Cite this article
Eguiluz, M., Rodrigues, N.F., Guzman, F. et al. The chloroplast genome sequence from Eugenia uniflora, a Myrtaceae from Neotropics. Plant Syst Evol 303, 1199–1212 (2017). https://doi.org/10.1007/s00606-017-1431-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00606-017-1431-x