Abstract
We determined the prevalence rate and risk of infection of Trypanosoma cruzi and other trypanosomatids in Peruvian non-human primates (NHPs) in the wild (n = 126) and in different captive conditions (n = 183). Blood samples were collected on filter paper, FTA cards, or EDTA tubes and tested using a nested PCR protocol targeting the 24Sα rRNA gene. Main risk factors associated with trypanosomatid and T. cruzi infection were genus and the human–animal context (wild vs captive animals). Wild NHPs had higher prevalence of both trypanosomatids (64.3 vs 27.9%, P < 0.001) and T. cruzi (8.7 vs 3.3%, P = 0.057), compared to captive NHPs, suggesting that parasite transmission in NHPs occurs more actively in the sylvatic cycle. In terms of primate family, Pitheciidae had the highest trypanosomatid prevalence (20/22, 90.9%) and Cebidae had the highest T. cruzi prevalence (15/117, 12.8%). T. cruzi and trypanosomatids are common in Peruvian NHPs and could pose a health risk to human and animals that has not been properly studied.
Similar content being viewed by others
Introduction
Trypanosoma cruzi, etiological agent of Chagas disease, infects approximately 8 million people worldwide and more than 25 million people are at risk of infection (WHO 2014). This neglected tropical disease is endemic in 21 Latin American countries, including Peru with 964 cases reported between 2000 and 2015 (MINSA 2015). However, the number of cases is probably underestimated as the disease is often asymptomatic. Transmission typically occurs via a triatomine bug vector, but it can also occur via blood transfusion, organ transplantation, congenital transmission, the oral route, and accidental infection (Luquetti and Schmuñis 2010). Recent outbreaks reported in Latin America have been caused by foodborne transmission of T. cruzi due to ingestion of contaminated beverages (Yoshida 2008; Coura and Junqueira 2015). The consumption of infected animal blood or raw or undercooked meat also represents a risk of infection due to the presence of blood trypanomastigotes and tissue amastigotes (Pereira et al. 2010; Rueda et al. 2014). This route of infection may be more common in tropical forests where wild animals are consumed or traded into urban areas.
The diagnosis of Chagas disease is complex due to the dynamics of parasitemia in the two successive phases of the disease: an acute phase diagnosed by the direct detection of the parasite circulating in the bloodstream (microscopy) where the parasite is abundant and a chronic phase where the parasite is hidden in host tissues and diagnosis is usually made via the detection of antibodies against T. cruzi (serology) (WHO 2014). In many prevalence studies, microscopy has commonly been used to detect trypanosomes in non-human primates (NHPs) (Sousa et al. 1974; D’Alessandro et al. 1986; Ziccardi and Lourenco-de-Oliveira 1997). However, this technique has a low sensitivity in the chronic phase, and species identification may be difficult due to the interspecific morphological similarities and the high intraspecific variability of trypanosomes (Ziccardi and Lourenco-de-Oliveira 1998). Alternatively, although PCR shows a higher specificity and sensitivity (Kirchhoff et al. 1996; Ndao et al. 2000; Cox et al. 2010), most PCR protocols for the detection of trypanosomes focus specifically on T. cruzi in human blood, overlooking other trypanosomatids probably present in wild animals that may have veterinary and public health importance. Thus, a nested PCR protocol is a convenient option that enables the detection of trypanosomatids in a first reaction, while a second reaction allows for the identification of T. cruzi.
Since non-human primates (NHPs) can act as reservoirs for human pathogens, they may serve as sentinels for epidemiologic studies (Wolfe et al. 1998), providing important information for the understanding of the epidemiology, transmission dynamics, and emergence risk of various anthropozoonoses. Close contact between these mammals and human beings may result in transmission risks in both urban and rural settings, possibly leading to emerging zoonoses. Communities in the Amazon frequently hunt and poach wildlife and may inadvertently introduce infectious parasites into new areas. Thus, epidemiological and prevalence studies that address these issues are necessary. By estimating the prevalence rate and risk of infection of trypanosomatids and T. cruzi from different genera of neotropical peruvian NHPs, we aimed to identify which NHP families and species may pose the highest risk of infection to humans, and to contribute with relevant data as studies in Peruvian primates is scarce.
Materials and Methods
Sample Collection
Convenience sampling was used based on the availability of captive (n = 183) and wild (n = 126) NHPs. From November 2011 to May 2012, captive NHPs from seven Peruvian cities were sampled from zoos, wildlife rescue centers, wet markets, and households (Fig. 1). A 1–3-ml blood sample was collected from the femoral vein of 183 primates in EDTA-containing tubes and an aliquot was applied to Flinders Technology Associates (FTA) classical cards. Blood samples were refrigerated during field collection and stored at −80°C with a silica gel desiccant until PCR analysis. In 82 of these NHPs, blood smears were also collected for microscopy screening of hemoparasites.
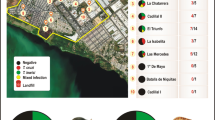
Map of Peru showing the locations where captive (n = 183, circles) and wild (n = 126, triangles) NHPs were sampled: Nueva Esperanza (n = 118), Yurimaguas (n = 68), Lima (n = 28), Pucallpa (n = 26), Puerto Maldonado (n = 26), Cuzco (n = 18), Moyobamba (n = 16), Sol Naciente (n = 8) and Iquitos (n = 1). Biological samples from wild NHPs were collected in two indigenous communities in the Northern Peruvian Amazon.
From 2007 to 2012, blood samples were collected from 126 wild NHPs by subsistence hunters from two remote indigenous communities in the Northeastern Peruvian Amazon: Nueva Esperanza (S 04°19.53 W 71°57.33) and Sol Naciente (S 03°38.26 W 73°12.57) (Fig. 1). A Whatman filter paper No. 3 was impregnated with blood from the jugular vein during the postmortem management of each NHP. This material was sealed in individual plastic bags with desiccant and kept in indoor conditions before transfer to storage at −80°C in Lima.
The research protocol was approved by the Peruvian Forestry and Wildlife Agency (0411-2010-AG-DGFFS-DGEFFS, 0618-2011-AG-DGFFS-DGEFFS, 0350-2012-AG-DGFFS-DGEFFS) and the Institutional Animal Care and Use Committee (IACUC) of UC Davis School of Veterinary Medicine (IACUC PREDICT-WCS #16027).
Nested PCR Protocol
DNA from parasite cultures and filter paper samples was extracted using QIAamp DNA Mini kit (Qiagen), while DNA from whole blood in EDTA tubes was extracted using QIAmp Blood Mini kit (Qiagen) and quantified using a NanoDrop 1000 spectrophotometer. Blood samples applied to FTA cards were processed following the manufacturer’s instructions (GE Healthcare Life Sciences).
The nested PCR protocol was optimized using primers for the 24S alpha subunit rRNA gene. Primers D75/D76 target the conserved flanking sequences of the D7 alpha domain in trypanosomatids, while primers D71/D72 target an internal region in the same domain that is specific to T. cruzi (Souto and Zingales 1993; Souto et al. 1999). PCR was performed in a Gene Amp PCR System 9700 thermocycler (Applied Biosystems, Foster City, CA) in a final volume of 30 µl.
The first PCR reaction had 3 µl of template DNA, and a final concentration of 1X PCR buffer (Invitrogen), 0.1 mM of each primer (D75 and D76), 1 U Taq DNA Polymerase (Thermo Scientific), 2.0 mM MgCl2, and 0.15 mM of each dNTP. The PCR protocol included an initial denaturation step at 94°C for 5 min; 35 cycles at 94°C for 40 s to denature the template, annealing at 57°C for 40 s, extension at 72°C for 40 s; and a final extension step at 72°C for 5 min. Amplicons were analyzed on a 2% agarose gel with a 100 bp DNA ladder (Invitrogen).
The second PCR reaction contained 1 µl of PCR product from the first reaction, and a final concentration of 1× PCR buffer (Invitrogen), 0.2 mM of each primer (D71 and D72), 1 U Taq DNA Polymerase (Thermo Scientific), 1.5 mM MgCl2, and 0.25 mM of each dNTP. The initial denaturation step of 94°C for 4 min, was followed by 30 cycles of denaturation at 94°C for 1 min, annealing at 62°C for 1 min, extension at 72°C for 1 min; and a final extension step at 72°C for 5 min. PCR products were analyzed in 2.5% agarose gels with a 50 bp size standard (Thermo Scientific).
Twelve parasite strains were used as reference, including T. rangeli, four T. cruzi strains (Tulahuen, Y, AQP 300, Viera Roque), and seven Leishmania species (L. (V.) braziliensis, L. (V.) peruviana, L. (V.) guyanensis, L. (V.) panamensis, L. (V.) lainsoni, L (L.) amazonensis, and L (L.) mexicana). DNA extracted from human and primate (Aotus nancymaae) blood infected with Plasmodium falciparum was used to assess cross-reactions with the host and Plasmodium. The analytical sensitivity of the protocol was evaluated by determining the minimum amount of T. cruzi DNA required for its successful PCR amplification. Serial 1:10 dilutions of T. cruzi DNA in AE buffer were prepared and a total of nine different concentrations were assayed (ranging from 0.5 ng/µl to 0.15 fg/µl).
Statistical Analyses
Diagnosis was based on microscopy and PCR. Results were grouped as correct positive diagnosis (a), false positive diagnosis (b), false negative diagnosis (c), correct negative diagnosis incorrect (d). From these values, the sensitivity (100 × a/(a + c)) and specificity (100 × d/(c + d)) of trypanosomatids and T. cruzi diagnosis were calculated (Ndao et al. 2000).
Prevalence rates of infection were calculated for both trypanosomatids and T. cruzi by expressing the percentage of infected individuals with respect to the number of analyzed monkeys. This analysis was performed considering primate taxonomy and the human-animal context in which infection could occur (wild and captive interfaces). Prevalences were compared with Chi-squared and Fisher’s exact tests (significance level P < 0.05). We used a recursive modeling approach with classification trees to identify major risk factors linked to the presence of trypanosomatids and T. cruzi using the package “rpart” version 4.1-9 of the statistical software R version 3.2.1 (Therneau et al. 2015). Recursive partitioning is the most appropriate technique for epidemiological studies based on complex relationships including nonlinear and interacting risk factors (Zhang and Singer 2010). The presence of trypanosomatids or T. cruzi was the dependent variable, and genus and the human–animal context (interface) were the independent categorical variables considered in the study as possible risk factors for prior trypanosomatid or T. cruzi infection.
Results
Nested PCR Optimization
All trypanosomatids generated bands of approximately 280 bp with primers D75/D76, while T. cruzi strains produced bands of approximately 120 bp with primers D71/D72. The nested PCR protocol did not show evidence of cross-reactions or non-specific amplification with A. nancymaae nor P. falciparum DNA control samples. Primers D71/D72 were specific for T. cruzi and did not amplify neither T. rangeli nor any of the seven Leishmania species assayed.
The detection limit of the PCR using primers D75/D76 for detecting trypanosomatids was 1.5 pg of parasite DNA, while the detection limit for the PCR using primers D71/D72 for detecting T. cruzi was 15 fg. Compared to PCR, microscopy had a sensitivity of 82.9% (29/35, 95% CI 66–93.0%) and a specificity of 97.9% (46/47, CI 89.1–99.9%).
Prevalence of Trypanosomatids and T. cruzi
Tables 1 and 2 show the prevalence of trypanosomatids and T. cruzi in the 16 species (11 genera, 5 families) of the 309 sampled NHPs screened. Table 3 shows the prevalence of trypanosomatids and T. cruzi in the NHPs, according to their holding facility.
Overall prevalence was 42.7% (132/309) for trypanosomatids and 5.5% (17/309) for T. cruzi. In decreasing order, the prevalence of trypanosomatids by NHP family was: Pitheciidae (90.9%), Atelidae (42.2%), Aotidae (40%) and Cebidae (35.9%) (P < 0.001). The highest prevalence of T. cruzi by family was observed in Cebidae (12.8%) and Pitheciidae (4.5%) (P < 0.001). Wild NHPs had a higher prevalence of both trypanosomatids (64.3 vs 27.9%, P < 0.001) and T. cruzi (8.7 vs 3.3%, P = 0.057) compared to captive NHPs (Table 1). Within the captive interface, wet markets were the most important holding facility for the presence of both trypanosomatids (P < 0.001) and T. cruzi (P < 0.008).
In NHPs hosts, the most parsimonious tree model explained 35.1% and 23.1% of the observed variability of presence of trypanosomatids (Fig. 2) and T. cruzi (Fig. 3), respectively. The best tree size for each model was selected using cross-validation: for the trypanosomatids model, a tree of size 5 (N splits = 4) with a complexity parameter CP = 0.0065; and for the T. cruzi model, a tree of size 6 (N splits = 5) with a complexity parameter CP = 0.0165. The trees showed that genus and interface are risk factors associated with trypanosomatids and T. cruzi.
Most parsimonious tree build using recursive partitioning to identify major risk factors linked to the presence of trypanosomatids in NHPs. Wild NHPs and NHPs from wet markets, and genera Alouatta, Lagothrix, Cacajao, Pithecia, Cebus, Samiri and Callicebus have a significantly higher risk of trypanosomatid infection. The most parsimonious tree model explained 35.1% of observed variability.
Most parsimonious tree build using recursive partitioning to identify major risk factors linked to the presence of Trypanosoma cruzi in NHPs. Wild NHPs, and genus Cebus, Saimiri and Callicebus have a significantly higher risk of T. cruzi infection. The most parsimonious tree model explained 23.1% of observed variability.
Discussion
Since the close contact between humans and NHPs infected with trypanosomes poses an important risk of infection, this study aimed to determine the prevalence rate of both trypanosomatids and T. cruzi in different genera of Neotropical primates in Peru. Trypanosomatid and T. cruzi prevalence rates were significantly higher in wild primates compared to captive animals, suggesting that parasite transmission in NHPs occurs more actively in the sylvatic cycle. Primates in the wild are probably exposed to a greater number of infection routes, such as triatomine vectors that may dwell and feed on primate resting sites, but also via oral route (i.e., ingesting infected insects) and other mechanisms such as vertical transmission or sexual contact (Jansen and Roque 2010; Rocha et al. 2013). In addition, there is a greater number of potential reservoir species in the wild which increases the probability of transmission within wild NHP populations. T. cruzi is a multihost parasite that exhibits remarkable biological and genetic diversity, as well as complex transmission cycles (Xavier et al. 2012; Rocha et al. 2013). Therefore, sylvatic cycles should be understood as dynamic webs, with parasite transmission occurring by different routes in cycles in the forest, understory, or canopy that may or may not overlap (Jansen and Roque 2010).
Neotropical primates can be naturally infected with several species of trypanosomes, including T. cruzi, T. rangeli, T. evansi, T. simiae, T. minasense, T. saimirii, T. lambrechti, T. devei, T. sanmartini, T. diasi, T. prowazecki, and T. mycetae (Ziccardi and Lourenco-de-Oliveira 1997; Ndao et al. 2000). Trypanosomes have been commonly reported in the NHP families Callitrichidae (tamarins and marmosets), Cebidae (squirrel monkeys and capuchins), Aotidae (night monkeys), and Atelidae (spider monkeys and howler monkeys) (Sousa et al. 1974; Ziccardi and Lourenco-de-Oliveira 1997; Ziccardi et al. 2000; Maia da Silva et al. 2008).
In our study, the observed variability in presence of trypanosomatids and T. cruzi was mainly explained by intrinsic (genus) and extrinsic (interface) factors. Among wild NHPs, Pitheciidae (Titi Monkeys, Sakis, and Uakaris) had the highest trypanosomatid prevalence (95.2%), and Cebidae had the highest T. cruzi prevalence (23.1%). Within captive NHPs, specimens from wet markets had the highest trypanosomatid and T. cruzi prevalences, which underscores the possibility that extracted primates from the wild may introduce the parasite into urban areas. The wet market interface probably has the closest prevalence rate to the one found in the wild, since recently captured animals from the jungle are brought into cities for trade in markets (Shanee et al. 2015). Primates recovered from this illegal traffic and from wildlife and biodiversity conservation programs are usually reintroduced to the wild without an exhaustive disease screening. Thus, availability of efficient protocols for trypanosome detection could mitigate risks to naïve resident populations exposed to reintroduced or translocated animals (Lisboa et al. 2004).
High mortality during transport and captivity could explain the lower rate of infection observed in captive NHPs. Another possibility for low prevalence is the absence of vectors in the area. However, transmission can still occur in closed NHP colonies by vector-independent mechanisms such as trauma, blood exposure, saliva, sexual activity and transplacentary transmission (Ndao et al. 2000; Kunz et al. 2002; Lisboa et al. 2004; Dorn et al. 2012).
Feeding behavior of NHPs is an intrinsic factor associated with genus that might contribute to the rate of infection. Oral transmission of T. cruzi is probably the most frequent mechanism among animals in the wild cycle, considering that several species of wild mammals frequently ingest insects, including infected triatomines (Coura 2015). In addition, stercorarial transmission may not be as efficient as the oral route because infected feces of triatomines are probably deposited in dense animal fur and not directly on skin (Roque et al. 2008).
Most human Chagas cases in Peru occur in the region of Arequipa (682 cases since 2004), where we could not sample NHPs. However, we did find T. cruzi positive animals in regions with very low numbers of human cases reported, such as Ucayali (6 cases since 2004) and Amazonas (13 cases) (MINSA 2015). This could represent the introduction of T. cruzi by NHPs to new areas, and future vectorial transmission cannot be ruled out since Peru has at least 19 species (9 genera) of triatomine bugs, which are present in all the regions sampled in this study (Lima, Cuzco, Loreto, San Martin, Madre de Dios, and Ucayali) (Chávez 2006; Náquira and Cabrera 2009).
Prevalence studies in Peruvian NHPs are scarce, and T. cruzi infection rates varied from 3.6 to 12.9% (Dunn et al. 1963; Sullivan et al. 1993), although some studies reported a difference in prevalence depending on the method used: 6.1% using microscopy, 10.4% using ELISA, and 26.5% using PCR (Ndao et al. 2000). Studies in free ranging primates of Central and South America show variable but usually high trypanosome prevalence. In Brazil, trypanosome prevalences of 47–51% (D’Alessandro et al. 1986), 67.9% (Ziccardi and Lourenco-de-Oliveira 1997), and 46.8–87.5% have been reported (Maia da Silva et al. 2008); in French Guiana, trypanosome prevalence varied between 16 and 51% depending on the primate species (de Thoisy et al. 2001); and in Panama, the trypanosome prevalence was 10.1% (Sousa et al. 1974). However, there is need for caution when comparing prevalences from different studies because of the different sampling techniques and the different diagnostic methods used (with varying sensitivities). For instance, the infection rates of T. cruzi in Amazonian monkeys ranged from 10.3% using parasitological methods (Ziccardi and Lourenco-de-Oliveira 1997; Ziccardi et al. 2000) to 46% using serology (Lisboa et al. 2006). Furthermore, in studies using PCR several other aspects must be taken into account, such as sample type, sample volume and conservation, DNA extraction method, parasite sequences targeted, primers, reagents, and thermocycling conditions (Luquetti and Schmuñis 2010).
In our study, whole blood was collected in EDTA tubes whenever possible, but samples were also collected on FTA cards and filter paper. Dried blood spot sampling overcame the limitations of cold chain in rural areas and remote locations (Luquetti and Schmuñis 2010). However, natural inhibitors present in samples and the uneven distribution of parasite DNA on FTA cards may decrease the assay’s sensitivity (Pizarro et al. 2007; Ahmed et al. 2011). We were able to analyze only one disk (punch) per sample, which could have resulted in an underestimation of the real prevalences, as it has been observed that Trypanosoma spp. prevalence increases dramatically when a higher number of FTA sections are analyzed (Cox et al. 2010). Further studies should consider analyzing multiple FTA sections and a complementary PCR protocol that targets host DNA to avoid false-negatives (Molaei et al. 2008; Jercic et al. 2010; Schijman et al. 2011).
In terms of sensitivity, the first PCR reaction amplifies up to 1.5 pg of parasite DNA, whereas the second reaction detects up to 15 fg of parasite DNA. Thus, the second step is more sensitive and may detect T. cruzi infections that are not evident with only the first step of the protocol. For this reason, even when a sample is negative for trypanosomatids, T. cruzi infection cannot be discarded and the second step should be performed. Estimating the exact number of parasites detected with the nested PCR is not completely straightforward. Preparing samples with very few parasites may not be accurate, as parasite DNA from partially degraded organisms may be present in the liquid portion of the samples (Moser et al. 1989; Kirchhoff et al. 1996). Thus, we opted to prepare serial dilutions of known amounts of parasite DNA. However, the total amount of DNA in T. cruzi cells varies among different strains (Santos et al. 1997), and the absolute amount of total DNA (nuclear + kinetoplast) varies from 0.12 to 0.33 pg per T. cruzi cell (Souza et al. 2011). Considering this range, the nested PCR protocol detects from 4.5 to 12.5 parasites in the first reaction, and 0.045 to 0.12 parasites in the second step.
In our study, we found the highest T. cruzi prevalence in NHPs in the wild and wet markets. Since NHPs may act as reservoirs for this parasite, their handlers are at risk of accidental infection (Ndao et al. 2000; Strait et al. 2012). Trade of wildlife for exotic pet markets results in close contact between humans and NHPs, providing new opportunities for disease emergence (Wolfe et al. 1998). Accidental infections have been reported to occur not only by needlestick injuries, but by exposure of mucous membranes to contaminated fluids and skin lesions while handling infected animals (Hofflin et al. 1987; Kinoshita-Yanaga et al. 2009, Pereira et al. 2010; Kirchhoff 2011). This may be a frequent threat in rural communities where wild animals are hunted for subsistence, because carcasses are handled with bare hands. Indeed, wildlife subsistence hunting in the Amazon region is a traditional source of food for rural populations (FitzGibbon 1998), and primates represent an important source of meat in the region (Puertas 2004; Bowler et al. 2014; Mayor et al. 2015). Furthermore, meat in these rural settings may be ingested raw or undercooked, and parasite pseudocysts and blood form trypomastigotes in the hunted animals could represent a high-risk of infection (Pereira et al. 2009).
In summary, T. cruzi and trypanosomatids frequently infect both wild and captive Peruvian NHPs, posing a risk to human and animal health that has not been properly studied. The main risk factors associated with trypanosomatids and T. cruzi infections were genus and interface. Prevalence rates were also especially high in wild primates, suggesting that parasite transmission in NHPs occurs more actively in the sylvatic cycle. NHPs are regularly hunted, manipulated, consumed, and introduced into urban areas; hence, further studies are necessary to better understand pathogen transmission and public health implications.
References
Ahmed HA, MacLeod ET, Hide G, Welburn SC and Picozzi K (2011). “The Best Practice for Preparation of Samples from FTA Cards for Diagnosis of Blood Borne Infections Using African Trypanosomes as a Model System.” Parasit Vectors 4: 68.
Bowler M, Anderson M, Montes D, Pérez P and Mayor P (2014). “Refining Reproductive Parameters for Modelling Sustainability and Extinction in Hunted Primate Populations in the Amazon.” PLoS ONE 9(4): e93625.
Coura JR (2015). “The Main Sceneries of Chagas Disease Transmission. The Vectors, Blood and Oral Transmissions - a Comprehensive Review.” Memórias do Instituto Oswaldo Cruz 110(3): 277–282.
Coura JR and Junqueira ACV (2015). “Ecological Diversity of Trypanosoma cruzi Transmission in the Amazon Basin. The Main Scenaries in the Brazilian Amazon.” Acta Trop 151: 51–57.
Cox AP, Tosas O, Tilley A, Picozzi K, Coleman P, Hide G and Welburn SC (2010). “Constraints to Estimating the Prevalence of Trypanosome Infections in East African Zebu Cattle.” Parasit Vectors 3: 82.
Chávez J (2006). “Contribución Al Estudio De Los Triatominos Del Perú: Distribución Geográfica, Nomenclatura Y Notas Taxonómicas.” Anales de la Facultad de Medicina 67: 65–76.
D’Alessandro A, Eberhard M, de Hincapie O and Halstead S (1986). “Trypanosoma cruzi and Trypanosoma rangeli in Saimiri Sciureus from Bolivia and Saguinus Mistax from Brazil.” Am J Trop Med Hyg 35(2): 285–289.
de Thoisy B, Vogel I, Reynes JM, Pouliquen JF, Carme B, Kazanji M and Vie JC (2001). “Health Evaluation of Translocated Free-Ranging Primates in French Guiana.” Am J Primatol 54(1): 1–16.
Dorn PL, Daigle ME, Combe CL, Tate AH, Stevens L and Phillippi-Falkenstein KM (2012). “Low Prevalence of Chagas Parasite Infection in a Nonhuman Primate Colony in Louisiana.” J Am Assoc Lab Anim Sci 51(4): 443–447.
Dunn FL, Lambrecht FL and Duplessis R (1963). “Trypanosomes of South American Monkeys and Marmosets.” Am J Trop Med Hyg 12: 524–534.
FitzGibbon C (1998). The Management of Subsistence Harvesting: Behavioural Ecology of Hunter and Their Mammalian Prey. New York, Oxford University Press.
Hofflin JM, Sadler RH, Araujo FG, Page WE and Remington JS (1987). “Laboratory-Acquired Chagas Disease.” Trans R Soc Trop Med Hyg 81(3): 437–440.
Jansen AM and Roque ALR (2010). 11 - Domestic and Wild Mammalian Reservoirs. American Trypanosomiasis. J. Telleria and M. Tibayrenc. London, Elsevier: 249–276.
Jercic MI, Mercado R and Villarroel R (2010). “Congenital Trypanosoma cruzi Infection in Neonates and Infants from Two Regions of Chile Where Chagas’ Disease Is Endemic.” J Clin Microbiol 48(10): 3824–3826.
Kinoshita-Yanaga AT, Toledo MJ, Araujo SM, Vier BP and Gomes ML (2009). “Accidental Infection by Trypanosoma cruzi Follow-up by the Polymerase Chain Reaction: Case Report.” Rev Inst Med Trop Sao Paulo 51(5): 295–298.
Kirchhoff LV (2011). Chapter 1 - Epidemiology of American Trypanosomiasis (Chagas Disease). In: H. B. T. Louis, M. Weiss and V. K. Louis (editor) Advances in Parasitology, vol 75. Academic Press, New York, pp 1–18.
Kirchhoff LV, Votava JR, Ochs DE and Moser DR (1996). “Comparison of PCR and Microscopic Methods for Detecting Trypanosoma cruzi” J Clin Microbiol 34(5): 1171–1175.
Kunz E, Matz-Rensing K, Stolte N, Hamilton PB and Kaup FJ (2002). “Reactivation of a Trypanosoma cruzi Infection in a Rhesus Monkey (Macaca Mulatta) Experimentally Infected with Siv.” Vet Pathol 39(6): 721–725.
Lisboa CV, Mangia RH, Luz SL, Kluczkovski A, Jr., Ferreira LF, Ribeiro CT, Fernandes O and Jansen AM (2006). “Stable Infection of Primates with Trypanosoma cruzi I and Ii.” Parasitology 133(Pt 5): 603–611.
Lisboa CV, Mangia RH, Rubiao E, de Lima NR, das Chagas Xavier SC, Picinatti A, Ferreira LF, Fernandes O and Jansen AM (2004). “Trypanosoma cruzi Transmission in a Captive Primate Unit, Rio De Janeiro, Brazil.” Acta Trop 90(1): 97–106.
Luquetti AO and Schmuñis GA (2010). 28 - Diagnosis of Trypanosoma cruzi Infection. American Trypanosomiasis. J. Telleria and M. Tibayrenc. London, Elsevier: 743–792.
Maia da Silva F, Naiff RD, Marcili A, Gordo M, D’Affonseca Neto JA, Naiff MF, Franco AM, Campaner M, Valente V, Valente SA, Camargo EP, Teixeira MM and Miles MA (2008). “Infection Rates and Genotypes of Trypanosoma rangeli and T. cruzi Infecting Free-Ranging Saguinus Bicolor (Callitrichidae), a Critically Endangered Primate of the Amazon Rainforest.” Acta Trop 107(2): 168–173.
Mayor P, Pérez-Peña P, Bowler M, Puertas PE, Kirkland M and Bodmer R (2015). “Effects of Selective Logging on Large Mammal Populations in a Remote Indigenous Territory in the Northern Peruvian Amazon.” Ecology and Society 20(4): 36.
MINSA (2015) “Sala Situacional Para El Análisis De Situacion De Salud - Se 38, 2015.” 2015, from http://www.dge.gob.pe/portal/index.php?option=com_content&view=article&id=14&Itemid=121.
Molaei G, Andreadis TG, Armstrong PM and Diuk-Wasser M (2008). “Host-Feeding Patterns of Potential Mosquito Vectors in Connecticut, U.S.A.: Molecular Analysis of Bloodmeals from 23 Species of Aedes, Anopheles, Culex, Coquillettidia, Psorophora, and Uranotaenia.” J Med Entomol 45(6): 1143–1151.
Moser DR, Kirchhoff LV and Donelson JE (1989). “Detection of Trypanosoma cruzi by DNA Amplification Using the Polymerase Chain Reaction.” J Clin Microbiol 27(7): 1477–1482.
Náquira C and Cabrera R (2009). “Breve Reseña Histórica De La Enfermedad De Chagas, a Cien Años De Su Descubrimiento Y Situación Actual En El Perú.” Revista Peruana de Medicina Experimental y Salud Publica 26: 494–504.
Ndao M, Kelly N, Normandin D, Maclean JD, Whiteman A, Kokoskin E, Arevalo I and Ward BJ (2000). “Trypanosoma cruzi Infection of Squirrel Monkeys: Comparison of Blood Smear Examination, Commercial Enzyme-Linked Immunosorbent Assay, and Polymerase Chain Reaction Analysis as Screening Tests for Evaluation of Monkey-Related Injuries.” Comp Med 50(6): 658–665.
Pereira KS, Schmidt FL, Barbosa RL, Guaraldo AMA, Franco RMB, Dias VL and Passos LAC (2010). Chapter 3 - Transmission of Chagas Disease (American Trypanosomiasis) by Food. Advances in Food and Nutrition Research. L. T. Steve, Academic Press. Volume 59: 63–85.
Pereira KS, Schmidt FL, Guaraldo AM, Franco RM, Dias VL and Passos LA (2009). “Chagas’ Disease as a Foodborne Illness.” J Food Prot 72(2): 441–446.
Pizarro JC, Lucero DE and Stevens L (2007). “PCR Reveals Significantly Higher Rates of Trypanosoma cruzi Infection Than Microscopy in the Chagas Vector, Triatoma Infestans: High Rates Found in Chuquisaca, Bolivia.” BMC Infect Dis 7: 66.
Puertas PE (2004). Hunting Effort as a Tool for Community-Based Wildlife Management in Amazonia. People in Nature, Columbia University Press: 123–136.
Rocha FL, Roque AL, de Lima JS, Cheida CC, Lemos FG, de Azevedo FC, Arrais RC, Bilac D, Herrera HM, Mourao G and Jansen AM (2013). “Trypanosoma cruzi Infection in Neotropical Wild Carnivores (Mammalia: Carnivora): At the Top of the T. cruzi Transmission Chain.” PLoS ONE 8(7): e67463.
Roque AL, Xavier SC, da Rocha MG, Duarte AC, D’Andrea PS and Jansen AM (2008). “Trypanosoma cruzi Transmission Cycle among Wild and Domestic Mammals in Three Areas of Orally Transmitted Chagas Disease Outbreaks.” Am J Trop Med Hyg 79(5): 742–749.
Rueda K, Trujillo JE, Carranza JC and Vallejo GA (2014). “Transmisión Oral De Trypanosoma cruzi: Una Nueva Situación Epidemiológica De La Enfermedad De Chagas En Colombia Y Otros Países Suramericanos.” Biomédica 34: 631–641.
Santos MR, Cano MI, Schijman A, Lorenzi H, Vázquez M, Levin MJ, Ramirez JL, Brandão A, Degrave WM and Silveira JFd (1997). “The Trypanosoma cruzi Genome Project: Nuclear Karyotype and Gene Mapping of Clone Cl Brener.” Memórias do Instituto Oswaldo Cruz 92: 821–828.
Schijman AG, Bisio M, Orellana L, Sued M, Duffy T, Mejia Jaramillo AM, Cura C, Auter F, Veron V, Qvarnstrom Y, Deborggraeve S, Hijar G, Zulantay I, Lucero RH, Velazquez E, Tellez T, Sanchez Leon Z, Galvao L, Nolder D, Monje Rumi M, Levi JE, Ramirez JD, Zorrilla P, Flores M, Jercic MI, Crisante G, Anez N, De Castro AM, Gonzalez CI, Acosta Viana K, Yachelini P, Torrico F, Robello C, Diosque P, Triana Chavez O, Aznar C, Russomando G, Buscher P, Assal A, Guhl F, Sosa Estani S, DaSilva A, Britto C, Luquetti A and Ladzins J (2011). “International Study to Evaluate Pcr Methods for Detection of Trypanosoma cruzi DNA in Blood Samples from Chagas Disease Patients.” PLoS Negl Trop Dis 5(1): e931.
Shanee N, Mendoza AP and Shanee S (2015) “Diagnostic Overview of the Illegal Trade in Primates and Law Enforcement in Peru.” Am J Primatol.
Sousa OE, Rossan RN and Baerg DC (1974). “The Prevalence of Trypanosomes and Microfilariae in Panamanian Monkeys.” Am J Trop Med Hyg 23(5): 862–868.
Souto RP, Vargas N and Zingales B (1999). “Trypanosoma rangeli: Discrimination from Trypanosoma cruzi Based on a Variable Domain from the Large Subunit Ribosomal Rna Gene.” Exp Parasitol 91(4): 306–314.
Souto RP and Zingales B (1993). “Sensitive Detection and Strain Classification of Trypanosoma cruzi by Amplification of a Ribosomal Rna Sequence.” Mol Biochem Parasitol 62(1): 45–52.
Souza RT, Lima FM, Barros RM, Cortez DR, Santos MF, Cordero EM, Ruiz JC, Goldenberg S, Teixeira MM and da Silveira JF (2011). “Genome Size, Karyotype Polymorphism and Chromosomal Evolution in Trypanosoma cruzi.” PLoS ONE 6(8): e23042.
Strait K, Else JG and Eberhard ML (2012). Chapter 4 - Parasitic Diseases of Nonhuman Primates. In: C Abee, K Mansfield, S Tardif, T Morris (ed) Nonhuman Primates in Biomedical Research (2nd Edition). Boston, Academic Press: 197–297.
Sullivan JJ, Steurer F, Benavides G, Tarleton RL, Eberhard ML and Landry S (1993). “Trypanosomes and Microfilariae in Feral Owl and Squirrel Monkeys Maintained in Research Colonies.” Am J Trop Med Hyg 49(2): 254–259.
Therneau T, Atkinson B, Ripley B (2015) Recursive Partitioning and Regression Trees. https://cran.r-project.org/web/packages/rpart/index.html. Accessed January 27, 2016.
WHO (2014) “Chagas Disease (American Trypanosomiasis).” 2016, from http://www.who.int/chagas/home_more/en/.
Wolfe ND, Escalante AA, Karesh WB, Kilbourn A, Spielman A and Lal AA (1998). “Wild Primate Populations in Emerging Infectious Disease Research: The Missing Link?” Emerg Infect Dis 4(2): 149–158.
Xavier SC, Roque AL, Lima Vdos S, Monteiro KJ, Otaviano JC, Ferreira da Silva LF and Jansen AM (2012). “Lower Richness of Small Wild Mammal Species and Chagas Disease Risk.” PLoS Negl Trop Dis 6(5): e1647.
Yoshida N (2008). “Trypanosoma cruzi Infection by Oral Route: How the Interplay between Parasite and Host Components Modulates Infectivity.” Parasitol Int 57(2): 105–109.
Zhang H and Singer B (2010). Recursive Partitioning and Applications, Springer, New York.
Ziccardi M and Lourenco-de-Oliveira R (1997). “The Infection Rates of Trypanosomes in Squirrel Monkeys at Two Sites in the Brazilian Amazon.” Mem Inst Oswaldo Cruz 92(4): 465–470.
Ziccardi M and Lourenco-de-Oliveira R (1998). “Morphological Features of Trypanosomes from Squirrel Monkeys from the Brazilian Amazon.” Mem Inst Oswaldo Cruz 93(1): 45–55.
Ziccardi M, Lourenco-De-Oliveira R, Lainson R, Brigido MC and Muniz JA (2000). “Trypanosomes of Non-Human Primates from the National Centre of Primates, Ananindeua, State of Para, Brazil.” Mem Inst Oswaldo Cruz 95(2): 157–159.
Acknowledgements
The authors thank the Peruvian Forestry Service and its regional offices for their cooperation in the management of research permits and access to confiscated animals. This project was possible thanks to financial support of the LA Zoo, USAID through the Emerging Pandemic Threats PREDICT Program, and NIH/FIC Training Grant D43 TW007393. The field work and sample collection was supported by the Cooperative Agreement # GHN-A-00-09-00010-00 “USAID Avian and Pandemic Influenza and Zoonotic Disease Program—PREDICT,” between USAID, Wildlife Conservation Society (WCS), and the Universitat Autònoma de Barcelona. We are also grateful to zoos and rescue centers for allowing us to work with animals from their collections, and to the villagers of Nueva Esperanza and Sol Naciente who kindly received us in the field. We thank the many volunteers, field assistants, veterinarians, and laboratory technicians who made this work possible. T. cruzi strains were provided by the Infectious Diseases Laboratory of Universidad Peruana Cayetano Heredia (UPCH), and T. rangeli was provided by Franklin Vargas from the Microbiology and Tropical Parasitology Research Institute of Trujillo National University (UNT).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that no competing interests exist.
Additional information
Disclaimers: The views expressed in this article are those of the authors and do not necessarily reflect the official policy or position of the Department of the Navy, Department of Defense, nor the U.S. Government. The experiments reported herein were conducted in compliance with the Animal Welfare Act and in accordance with the principles set forth in the “Guide for the Care and Use of Laboratory Animals,” Institute of Laboratory Animals Resources, National Research Council, National Academy Press, 1996.
Rights and permissions
About this article
Cite this article
Aysanoa, E., Mayor, P., Mendoza, A.P. et al. Molecular Epidemiology of Trypanosomatids and Trypanosoma cruzi in Primates from Peru. EcoHealth 14, 732–742 (2017). https://doi.org/10.1007/s10393-017-1271-8
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10393-017-1271-8