Abstract
Purpose
There is a need for biomarkers of drug efficacy for targeted therapies in triple-negative breast cancer (TNBC). As a step toward this, we identify multi-omic molecular determinants of anti-TNBC efficacy in cell lines for a panel of oncology drugs.
Methods
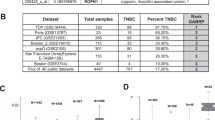
Using 23 TNBC cell lines, drug sensitivity scores (DSS3) were determined using a panel of investigational drugs and drugs approved for other indications. Molecular readouts were generated for each cell line using RNA sequencing, RNA targeted panels, DNA sequencing, and functional proteomics. DSS3 values were correlated with molecular readouts using a FDR-corrected significance cutoff of p* < 0.05 and yielded molecular determinant panels that predict anti-TNBC efficacy.
Results
Six molecular determinant panels were obtained from 12 drugs we prioritized based on their efficacy. Determinant panels were largely devoid of DNA mutations of the targeted pathway. Molecular determinants were obtained by correlating DSS3 with molecular readouts. We found that co-inhibiting molecular correlate pathways leads to robust synergy across many cell lines.
Conclusions
These findings demonstrate an integrated method to identify biomarkers of drug efficacy in TNBC where DNA predictions correlate poorly with drug response. Our work outlines a framework for the identification of novel molecular determinants and optimal companion drugs for combination therapy based on these correlates.




Similar content being viewed by others
References
Pareja F, Reis-Filho JS (2018) Triple-negative breast cancers—a panoply of cancer types. Nat Rev Clin Oncol 15(6):347–348
Wahba HA, El-Hadaad HA (2015) Current approaches in treatment of triple-negative breast cancer. Cancer Biol Med 12(2):106–116
Kalimutho M, Parsons K, Mittal D, Lopez JA, Srihari S, Khanna KK (2015) Targeted therapies for triple-negative breast cancer: combating a stubborn disease. Trends Pharmacol Sci 36(12):822–846
Le Du F, Ueno NT (2015) Targeted therapies in triple-negative breast cancer: failure and future. Womens Health (Lond) 11(1):1–5
Yam C, Mani SA, Moulder SL (2017) Targeting the molecular subtypes of triple negative breast cancer: understanding the diversity to progress the field. Oncologist 22(9):1086–1093
Pellegrino B, Mateo J, Serra V, Balmana J (2019) Controversies in oncology: are genomic tests quantifying homologous recombination repair deficiency (HRD) useful for treatment decision making? Esmo Open. https://doi.org/10.1136/esmoopen-2018-000480
Mayer EL, Abramson VG, Jankowitz RC, Falkson CI, Marcom PK, Traina TA, Carey LA, Rimawi MF, Specht JM, Miller K et al (2019) TBCRC 030: a randomized phase II study of preoperative cisplatin versus paclitaxel in TNBC—evaluating the homologous recombination deficiency (HRD) biomarker. J Clin Oncol 37(15_suppl):507
Schmid P, Adams S, Rugo HS, Schneeweiss A, Barrios CH, Iwata H, Dieras V, Hegg R, Im SA, Shaw Wright G et al (2018) Atezolizumab and Nab-paclitaxel in advanced triple-negative breast cancer. N Engl J Med 379(22):2108–2121
Heimes AS, Schmidt M (2019) Atezolizumab for the treatment of triple-negative breast cancer. Expert Opin Investig Drugs 28(1):1–5
Griguolo G, Dieci MV, Guarneri V, Conte P (2018) Olaparib for the treatment of breast cancer. Expert Rev Anticancer Ther 18(6):519–530
Romero D (2019) Benefit in patients with PD-L1-positive TNBC. Nat Rev Clin Oncol 16(1):6
Bareche Y, Venet D, Ignatiadis M, Aftimos P, Piccart M, Rothe F, Sotiriou C (2018) Unravelling triple-negative breast cancer molecular heterogeneity using an integrative multiomic analysis. Ann Oncol 29(4):895–902
Chiu AM, Mitra M, Boymoushakian L, Coller HA (2018) Integrative analysis of the inter-tumoral heterogeneity of triple-negative breast cancer. Sci Rep 8(1):11807
Davis SL, Eckhardt SG, Tentler JJ, Diamond JR (2014) Triple-negative breast cancer: bridging the gap from cancer genomics to predictive biomarkers. Ther Adv Med Oncol 6(3):88–100
Lander ES, Linton LM, Birren B, Nusbaum C, Zody MC, Baldwin J, Devon K, Dewar K, Doyle M, FitzHugh W et al (2001) Initial sequencing and analysis of the human genome. Nature 409(6822):860–921
Saito M, Momma T, Kono K (2018) Targeted therapy according to next generation sequencing-based panel sequencing. Fukushima J Med Sci 64(1):9–14
Lynch TJ, Bell DW, Sordella R, Gurubhagavatula S, Okimoto RA, Brannigan BW, Harris PL, Haserlat SM, Supko JG, Haluska FG et al (2004) Activating mutations in the epidermal growth factor receptor underlying responsiveness of non-small-cell lung cancer to gefitinib. N Engl J Med 350(21):2129–2139
Garrido-Castro AC, Lin NU, Polyak K (2019) Insights into molecular classifications of triple-negative breast cancer: improving patient selection for treatment. Cancer Discov 9(2):176–198
Mardis E (2018) Many mutations in one clinical-trial basket. Nature 554(7691):173–175
Renfro LA, Sargent DJ (2017) Statistical controversies in clinical research: basket trials, umbrella trials, and other master protocols: a review and examples. Ann Oncol 28(1):34–43
Wheler J, Lee JJ, Kurzrock R (2014) Unique molecular landscapes in cancer: implications for individualized, curated drug combinations. Cancer Res 74(24):7181–7184
Roerink SF, Sasaki N, Lee-Six H, Young MD, Alexandrov LB, Behjati S, Mitchell TJ, Grossmann S, Lightfoot H, Egan DA et al (2018) Intra-tumour diversification in colorectal cancer at the single-cell level. Nature 556(7702):457–462
Hodson R (2016) Precision medicine. Nature 537(7619):S49
Dugger SA, Platt A, Goldstein DB (2018) Drug development in the era of precision medicine. Nat Rev Drug Discov 17(3):183–196
Yadav B, Pemovska T, Szwajda A, Kulesskiy E, Kontro M, Karjalainen R, Majumder MM, Malani D, Murumagi A, Knowles J et al (2014) Quantitative scoring of differential drug sensitivity for individually optimized anticancer therapies. Sci Rep 4:5193
Chou TC (2010) Drug combination studies and their synergy quantification using the Chou–Talalay method. Cancer Res 70(2):440–446
Chou TC (2006) Theoretical basis, experimental design, and computerized simulation of synergism and antagonism in drug combination studies. Pharmacol Rev 58(3):621–681
Morpheus
Chou TC, Talalay P (1984) Quantitative analysis of dose–effect relationships: the combined effects of multiple drugs or enzyme inhibitors. Adv Enzyme Regul 22:27–55
Brower V (2015) NCI-MATCH pairs tumor mutations with matching drugs. Nat Biotechnol 33(8):790–791
Fan J, Upadhye S, Worster A (2006) Understanding receiver operating characteristic (ROC) curves. CJEM 8(1):19–20
Hammerman PS, Sos ML, Ramos AH, Xu C, Dutt A, Zhou W, Brace LE, Woods BA, Lin W, Zhang J et al (2011) Mutations in the DDR2 kinase gene identify a novel therapeutic target in squamous cell lung cancer. Cancer Discov 1(1):78–89
Thota R, Johnson DB, Sosman JA (2015) Trametinib in the treatment of melanoma. Expert Opin Biol Ther 15(5):735–747
Hassan B, Akcakanat A, Sangai T, Evans KW, Adkins F, Eterovic AK, Zhao H, Chen K, Chen H, Do KA et al (2014) Catalytic mTOR inhibitors can overcome intrinsic and acquired resistance to allosteric mTOR inhibitors. Oncotarget 5(18):8544–8557
Guo Y, Kwiatkowski DJ (2013) Equivalent benefit of rapamycin and a potent mTOR ATP-competitive inhibitor, MLN0128 (INK128), in a mouse model of tuberous sclerosis. Mol Cancer Res 11(5):467–473
Tetsu O, McCormick F (2017) ETS-targeted therapy: can it substitute for MEK inhibitors? Clin Transl Med 6(1):16
Rai K, Akdemir KC, Kwong LN, Fiziev P, Wu CJ, Keung EZ, Sharma S, Samant NS, Williams M, Axelrad JB et al (2015) Dual roles of RNF2 in melanoma progression. Cancer Discov 5(12):1314–1327
Xu J, Pfarr N, Endris V, Mai EK, Md Hanafiah NH, Lehners N, Penzel R, Weichert W, Ho AD, Schirmacher P et al (2017) Molecular signaling in multiple myeloma: association of RAS/RAF mutations and MEK/ERK pathway activation. Oncogenesis 6(5):e337
Diamond JR, Eckhardt SG, Pitts TM, van Bokhoven A, Aisner D, Gustafson DL, Capasso A, Sams S, Kabos P, Zolman K et al (2018) A phase II clinical trial of the Aurora and angiogenic kinase inhibitor ENMD-2076 for previously treated, advanced, or metastatic triple-negative breast cancer. Breast Cancer Res 20(1):82
Chang Q, Jorgensen C, Pawson T, Hedley DW (2008) Effects of dasatinib on EphA2 receptor tyrosine kinase activity and downstream signalling in pancreatic cancer. Br J Cancer 99(7):1074–1082
Shi H, Zhang CJ, Chen GY, Yao SQ (2012) Cell-based proteome profiling of potential dasatinib targets by use of affinity-based probes. J Am Chem Soc 134(6):3001–3014
Yang W, Soares J, Greninger P, Edelman EJ, Lightfoot H, Forbes S, Bindal N, Beare D, Smith JA, Thompson IR et al (2013) Genomics of drug sensitivity in cancer (GDSC): a resource for therapeutic biomarker discovery in cancer cells. Nucleic Acids Res 41(Database issue):D955–D961
Ben-David U, Siranosian B, Ha G, Tang H, Oren Y, Hinohara K, Strathdee CA, Dempster J, Lyons NJ, Burns R et al (2018) Genetic and transcriptional evolution alters cancer cell line drug response. Nature 560(7718):325–330
Kim SB, Dent R, Im SA, Espie M, Blau S, Tan AR, Isakoff SJ, Oliveira M, Saura C, Wongchenko MJ et al (2017) Ipatasertib plus paclitaxel versus placebo plus paclitaxel as first-line therapy for metastatic triple-negative breast cancer (LOTUS): a multicentre, randomised, double-blind, placebo-controlled, phase 2 trial. Lancet Oncol 18(10):1360–1372
Tamura S, Wang Y, Veeneman B, Hovelson D, Bankhead A 3rd, Broses LJ, Lorenzatti Hiles G, Liebert M, Rubin JR, Day KC et al (2018) molecular correlates of in vitro responses to dacomitinib and afatinib in bladder cancer. Bladder Cancer 4(1):77–90
Nagamine A, Araki T, Nagano D, Miyazaki M, Yamamoto K (2019) l-Lactate dehydrogenase B may be a predictive marker for sensitivity to anti-EGFR monoclonal antibodies in colorectal cancer cell lines. Oncol Lett 17(5):4710–4716
Oztop S, Isik A, Guner G, Gurdal H, Karabulut E, Yilmaz E, Akyol A (2019) Class III beta-tubulin expression in colorectal neoplasms is a potential predictive biomarker for paclitaxel response. Anticancer Res 39(2):655–662
Wallden B, Storhoff J, Nielsen T, Dowidar N, Schaper C, Ferree S, Liu S, Leung S, Geiss G, Snider J et al (2015) Development and verification of the PAM50-based Prosigna breast cancer gene signature assay. BMC Med Genomics 8:54
Costa RLB, Han HS, Gradishar WJ (2018) Targeting the PI3 K/AKT/mTOR pathway in triple-negative breast cancer: a review. Breast Cancer Res Treat 169(3):397–406
Ganesan P, Moulder S, Lee JJ, Janku F, Valero V, Zinner RG, Naing A, Fu S, Tsimberidou AM, Hong D et al (2014) Triple-negative breast cancer patients treated at MD Anderson Cancer Center in phase I trials: improved outcomes with combination chemotherapy and targeted agents. Mol Cancer Ther 13(12):3175–3184
Ibrahim YH, Garcia-Garcia C, Serra V, He L, Torres-Lockhart K, Prat A, Anton P, Cozar P, Guzman M, Grueso J et al (2012) PI3 K inhibition impairs BRCA1/2 expression and sensitizes BRCA-proficient triple-negative breast cancer to PARP inhibition. Cancer Discov 2(11):1036–1047
Shimoi T, Hamada A, Yamagishi M, Hirai M, Yoshida M, Nishikawa T, Sudo K, Shimomura A, Noguchi E, Yunokawa M et al (2018) PIK3CA mutation profiling in patients with breast cancer, using a highly sensitive detection system. Cancer Sci 109(8):2558–2566
Basho RK, Gilcrease M, Murthy RK, Helgason T, Karp DD, Meric-Bernstam F, Hess KR, Herbrich SM, Valero V, Albarracin C et al (2017) Targeting the PI3 K/AKT/mTOR pathway for the treatment of mesenchymal triple-negative breast cancer: evidence from a phase 1 trial of mTOR inhibition in combination with liposomal doxorubicin and bevacizumab. JAMA Oncol 3(4):509–515
Fouque A, Jean M, Weghe P, Legembre P (2016) Review of PI3 K/mTOR inhibitors entering clinical trials to treat triple negative breast cancers. Recent Pat Anticancer Drug Discov 11(3):283–296
Stanfield Z, Coskun M, Koyuturk M (2017) Corrigendum: drug response prediction as a link prediction problem. Sci Rep 7:44961
Groenendijk FH, Bernards R (2014) Drug resistance to targeted therapies: deja vu all over again. Mol Oncol 8(6):1067–1083
Wein L, Loi S (2017) Mechanisms of resistance of chemotherapy in early-stage triple negative breast cancer (TNBC). Breast 34(Suppl 1):S27–S30
Mundt F, Rajput S, Li S, Ruggles KV, Mooradian AD, Mertins P, Gillette MA, Krug K, Guo Z, Hoog J et al (2018) Mass spectrometry-based proteomics reveals potential roles of NEK9 and MAP2K4 in resistance to PI3 K inhibition in triple-negative breast cancers. Cancer Res 78(10):2732–2746
Wu ZH, Lin C, Liu MM, Zhang J, Tao ZH, Hu XC (2016) Src inhibition can synergize with gemcitabine and reverse resistance in triple negative breast cancer cells via the AKT/c-Jun pathway. PLoS ONE 11(12):e0169230
Mumin NH, Drobnitzky N, Patel A, Lourenco LM, Cahill FF, Jiang Y, Kong A, Ryan AJ (2019) Overcoming acquired resistance to HSP90 inhibition by targeting JAK-STAT signalling in triple-negative breast cancer. BMC Cancer 19(1):102
Miles D, von Minckwitz G, Seidman AD (2002) Combination versus sequential single-agent therapy in metastatic breast cancer. Oncologist 7(Suppl 6):13–19
Lee J, Yesilkanal AE, Wynne JP, Frankenberger C, Liu J, Yan J, Elbaz M, Rabe DC, Rustandy FD, Tiwari P et al (2019) Effective breast cancer combination therapy targeting BACH1 and mitochondrial metabolism. Nature 568(7751):254–258
Acknowledgements
We kindly thank Rork Kuick (University of Michigan) for his insight on statistical analysis and careful editing of this manuscript. We thank the MD Anderson RPPA core for protein analysis, TIGEM in Naples, Italy for RNA sequencing, and the University of Michigan sequencing core for DNA sequencing.
Funding
This research was supported by the National Institutes of Health (1R21CA218498 to M.B.S. and S.D.M.), the Breast Cancer Research Foundation (to S.D.M.), Tempting Tables (to S.D.M.), The Rose Run (to S.D.M.), and the Kathy Bruk Pearce Research Fund of the University of Michigan Rogel Cancer Center (to M.B.S.).
Author information
Authors and Affiliations
Contributions
Study concept and design: M.B.S., N.M.M. Acquisition of data: N.M.M., N.M.V., E.J.L., L.W.B. Drafting the manuscript: N.M.M., J.A.Y., M.B.S., S.D.M. Analysis and interpretation of data: N.M.M., M.B.S., P.J.L, J.P.L., S.D.M. Experimentation: N.M.M., N.M.V., E.J.L., L.W.B. Statistical analysis: N.M.M., P.U.L., J.P.L. Administrative, technical, or material support: J.A.Y., A.M. Study supervision: M.B.S., S.D.M. All authors reviewed and approved the final version of the manuscript.
Corresponding authors
Ethics declarations
Conflicts of interest
All authors declare that they have no conflicts of interest related to this work.
Research involving human and animal participants
This article does not contain any studies with human participants or animals performed by any of the authors.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
10549_2019_5473_MOESM2_ESM.eps
Supplementary material 2 Candidate molecular correlates in TNBC cell lines. Examples of compounds with significant correlates include: A. temsirolimus and B. docetaxel. All molecular marker readouts have a FDR-adjusted p*-value < 0.05. Top bars depict DSS3, with dark green representing the most sensitive lines and white representing the least sensitive. Orange labeled readouts are derived from DNA variants. Black labeled readouts are derived from targeted RNA expression levels (EPS 656 kb)
10549_2019_5473_MOESM4_ESM.docx
Supplementary material 4 Cell culture conditions. When comparing the Nanostring readouts with RNA sequencing, we observed strong correlations for A. IL7R, B. ERBB2, C. KIF2C, D. CD164, E. PRR15L. F. CACNB3, G. MYCL, H. EPHA2, I. PIK3CD, J. IGFBP4, and K. IRAK2. We observed poor correlations for L. FOSL1 (DOCX 314 kb)
10549_2019_5473_MOESM5_ESM.pdf
Supplementary material 5 Table showing IC50 values, hill-slopes, max response, dose range, and DSS3 values in individual cell lines. (PDF 312 kb)
10549_2019_5473_MOESM6_ESM.pdf
Supplementary material 6 Individual correlating DNA non-synonymous mutations in the molecular correlate panel. Genes marked with “*” indicate a roll-up of mutations was used for correlations. SIFT, Polyphen2, MutationTaster, MutationAssessor, and FATHMM were used to generate functional predictions. (PDF 25 kb)
Rights and permissions
About this article
Cite this article
Merrill, N.M., Lachacz, E.J., Vandecan, N.M. et al. Molecular determinants of drug response in TNBC cell lines. Breast Cancer Res Treat 179, 337–347 (2020). https://doi.org/10.1007/s10549-019-05473-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10549-019-05473-9