Abstract
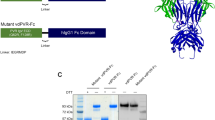
Hematopoietic human colony-stimulating factor 1 (hCSF-1) is essential for innate and adaptive immunity against viral and microbial infections and cancer. The human pathogen Epstein-Barr virus secretes the lytic-cycle protein BARF1 that neutralizes hCSF-1 to achieve immunomodulation. Here we show that BARF1 binds the dimer interface of hCSF-1 with picomolar affinity, away from the cognate receptor–binding site, to establish a long-lived complex featuring three hCSF-1 at the periphery of the BARF1 toroid. BARF1 locks dimeric hCSF-1 into an inactive conformation, rendering it unable to signal via its cognate receptor on human monocytes. This reveals a new functional role for hCSF-1 cooperativity in signaling. We propose a new viral strategy paradigm featuring an allosteric decoy receptor of the competitive type, which couples efficient sequestration and inactivation of the host growth factor to abrogate cooperative assembly of the cognate signaling complex.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
References
Young, L.S. & Rickinson, A.B. Epstein-Barr virus: 40 years on. Nat. Rev. Cancer 4, 757–768 (2004).
Kutok, J.L. & Wang, F. Spectrum of Epstein-Barr virus-associated diseases. Annu. Rev. Pathol. 1, 375–404 (2006).
Alcami, A. Viral mimicry of cytokines, chemokines and their receptors. Nat. Rev. Immunol. 3, 36–50 (2003).
Rivailler, P., Cho, Y.G. & Wang, F. Complete genomic sequence of an Epstein-Barr virus-related herpesvirus naturally infecting a new world primate: a defining point in the evolution of oncogenic lymphocryptoviruses. J. Virol. 76, 12055–12068 (2002).
Seto, E., Ooka, T., Middeldorp, J. & Takada, K. Reconstitution of nasopharyngeal carcinoma-type EBV infection induces tumorigenicity. Cancer Res. 68, 1030–1036 (2008).
Seto, E. et al. Epstein-Barr virus (EBV)-encoded BARF1 gene is expressed in nasopharyngeal carcinoma and EBV-associated gastric carcinoma tissues in the absence of lytic gene expression. J. Med. Virol. 76, 82–88 (2005).
Orlova, N., Fogg, M.H., Carville, A. & Wang, F. Antibodies to lytic infection proteins in lymphocryptovirus-infected rhesus macaques: a model for humoral immune responses to Epstein-Barr virus infection. Clin. Vaccine Immunol. 18, 1427–1434 (2011).
Wei, M.X. & Ooka, T. A transforming function of the BARF1 gene encoded by Epstein-Barr virus. EMBO J. 8, 2897–2903 (1989).
Wei, M.X., Moulin, J.C., Decaussin, G., Berger, F. & Ooka, T. Expression and tumorigenicity of the Epstein-Barr virus BARF1 gene in human Louckes B-lymphocyte cell line. Cancer Res. 54, 1843–1848 (1994).
Wei, M.X., deTurenne-Tessier, M., Decaussin, G., Benet, G. & Ooka, T. Establishment of a monkey kidney epithelial cell line with the BARF1 open reading frame from Epstein-Barr virus. Oncogene 14, 3073–3081 (1997).
Sheng, W., Decaussin, G., Ligout, A., Takada, K. & Ooka, T. Malignant transformation of Epstein-Barr virus-negative Akata cells by introduction of the BARF1 gene carried by Epstein-Barr virus. J. Virol. 77, 3859–3865 (2003).
Sall, A. et al. Mitogenic activity of Epstein-Barr virus-encoded BARF1 protein. Oncogene 23, 4938–4944 (2004).
Cohen, J.I. & Lekstrom, K. Epstein-Barr virus BARF1 protein is dispensable for B-cell transformation and inhibits alpha interferon secretion from mononuclear cells. J. Virol. 73, 7627–7632 (1999).
Ohashi, M., Orlova, N., Quink, C. & Wang, F. Cloning of the Epstein-Barr virus-related rhesus lymphocryptovirus as a bacterial artificial chromosome: a loss-of-function mutation of the rhBARF1 immune evasion gene. J. Virol. 85, 1330–1339 (2011).
Strockbine, L.D. et al. The Epstein-Barr virus BARF1 gene encodes a novel, soluble colony-stimulating factor-1 receptor. J. Virol. 72, 4015–4021 (1998).
Warren, M.K. & Ralph, P. Macrophage growth factor CSF-1 stimulates human monocyte production of interferon, tumor necrosis factor, and colony stimulating activity. J. Immunol. 137, 2281–2285 (1986).
Lin, J.C., Zhang, Z.X., Chou, T.C., Sim, I. & Pagano, J.S. Synergistic inhibition of Epstein-Barr virus: transformation of B lymphocytes by α and γ interferon and by 3′-azido-3′-deoxythymidine. J. Infect. Dis. 159, 248–254 (1989).
Hislop, A.D., Taylor, G.S., Sauce, D. & Rickinson, A.B. Cellular responses to viral infection in humans: lessons from Epstein-Barr virus. Annu. Rev. Immunol. 25, 587–617 (2007).
Hamilton, J.A. Colony-stimulating factors in inflammation and autoimmunity. Nat. Rev. Immunol. 8, 533–544 (2008).
Elegheert, J. et al. Extracellular complexes of the hematopoietic human and mouse CSF-1 receptor are driven by common assembly principles. Structure 19, 1762–1772 (2011).
Tarbouriech, N., Ruggiero, F., de Turenne-Tessier, M., Ooka, T. & Burmeister, W.P. Structure of the Epstein-Barr virus oncogene BARF1. J. Mol. Biol. 359, 667–678 (2006).
Gorba, C., Miyashita, O. & Tama, F. Normal-mode flexible fitting of high-resolution structure of biological molecules toward one-dimensional low-resolution data. Biophys. J. 94, 1589–1599 (2008).
Chen, X., Liu, H., Focia, P.J., Shim, A.H. & He, X. Structure of macrophage colony stimulating factor bound to FMS: diverse signaling assemblies of class III receptor tyrosine kinases. Proc. Natl. Acad. Sci. USA 105, 18267–18272 (2008).
Velazquez-Campoy, A. & Freire, E. Isothermal titration calorimetry to determine association constants for high-affinity ligands. Nat. Protoc. 1, 186–191 (2006).
Roussel, M.F., Downing, J.R., Rettenmier, C.W. & Sherr, C.J. A point mutation in the extracellular domain of the human CSF-1 receptor (c-fms proto-oncogene product) activates its transforming potential. Cell 55, 979–988 (1988).
Lupardus, P.J., Birnbaum, M.E. & Garcia, K.C. Molecular basis for shared cytokine recognition revealed in the structure of an unusually high affinity complex between IL-13 and IL-13Rα2. Structure 18, 332–342 (2010).
Taylor, E.W., Fear, A.L., Bohm, A., Kim, S.H. & Koths, K. Structure-function studies on recombinant human macrophage colony-stimulating factor (M-CSF). J. Biol. Chem. 269, 31171–31177 (1994).
Mantovani, A., Locati, M., Vecchi, A., Sozzani, S. & Allavena, P. Decoy receptors: a strategy to regulate inflammatory cytokines and chemokines. Trends Immunol. 22, 328–336 (2001).
Morgan, C., Pollard, J.W. & Stanley, E.R. Isolation and characterization of a cloned growth factor dependent macrophage cell line, BAC1.2F5. J. Cell. Physiol. 130, 420–427 (1987).
Yuzawa, S. et al. Structural basis for activation of the receptor tyrosine kinase KIT by stem cell factor. Cell 130, 323–334 (2007).
Verstraete, K. et al. Structural insights into the extracellular assembly of the hematopoietic Flt3 signaling complex. Blood 118, 60–68 (2011).
Ma, X. et al. Structural basis for the dual recognition of helical cytokines IL-34 and CSF-1 by CSF-1R. Structure 20, 676–687 (2012).
Sheng, W., Decaussin, G., Sumner, S. & Ooka, T. N-terminal domain of BARF1 gene encoded by Epstein-Barr virus is essential for malignant transformation of rodent fibroblasts and activation of BCL-2. Oncogene 20, 1176–1185 (2001).
Lemmon, M.A., Pinchasi, D., Zhou, M., Lax, I. & Schlessinger, J. Kit receptor dimerization is driven by bivalent binding of stem cell factor. J. Biol. Chem. 272, 6311–6317 (1997).
Krumm, B., Meng, X., Li, Y., Xiang, Y. & Deng, J. Structural basis for antagonism of human interleukin 18 by poxvirus interleukin 18-binding protein. Proc. Natl. Acad. Sci. USA 105, 20711–20715 (2008).
Liu, H. et al. Structural basis of semaphorin-plexin recognition and viral mimicry from Sema7A and A39R complexes with PlexinC1. Cell 142, 749–761 (2010).
Nuara, A.A. et al. Structure and mechanism of IFN-gamma antagonism by an orthopoxvirus IFN-gamma-binding protein. Proc. Natl. Acad. Sci. USA 105, 1861–1866 (2008).
Yang, Z., West, A.P. Jr. & Bjorkman, P.J. Crystal structure of TNFα complexed with a poxvirus MHC-related TNF binding protein. Nat. Struct. Mol. Biol. 16, 1189–1191 (2009).
Yoon, S.I., Jones, B.C., Logsdon, N.J. & Walter, M.R. Same structure, different function: crystal structure of the Epstein-Barr virus IL-10 bound to the soluble IL-10R1 chain. Structure 13, 551–564 (2005).
Calderwood, M.A. et al. Epstein-Barr virus and virus human protein interaction maps. Proc. Natl. Acad. Sci. USA 104, 7606–7611 (2007).
Uetz, P. et al. Herpesviral protein networks and their interaction with the human proteome. Science 311, 239–242 (2006).
Finger, C., Escher, C. & Schneider, D. The single transmembrane domains of human receptor tyrosine kinases encode self-interactions. Sci. Signal. 2, ra56 (2009).
Janowska-Wieczorek, A. et al. Increased circulating colony-stimulating factor-1 in patients with preleukemia, leukemia, and lymphoid malignancies. Blood 77, 1796–1803 (1991).
Chang, V.T. et al. Glycoprotein structural genomics: solving the glycosylation problem. Structure 15, 267–273 (2007).
Aricescu, A.R. et al. Eukaryotic expression: developments for structural proteomics. Acta Crystallogr. D. Biol. Crystallogr. 62, 1114–1124 (2006).
Winter, G. xia2: an expert system for macromolecular crystallography data reduction. J. Appl. Crystallogr. 43, 186–190 (2010).
McCoy, A.J. et al. Phaser crystallographic software. J. Appl. Crystallogr. 40, 658–674 (2007).
Pandit, J. et al. Three-dimensional structure of dimeric human recombinant macrophage colony-stimulating factor. Science 258, 1358–1362 (1992).
Langer, G., Cohen, S.X., Lamzin, V.S. & Perrakis, A. Automated macromolecular model building for X-ray crystallography using ARP/wARP version 7. Nat. Protoc. 3, 1171–1179 (2008).
Emsley, P., Lohkamp, B., Scott, W.G. & Cowtan, K. Features and development of Coot. Acta Crystallogr. D. Biol. Crystallogr. 66, 486–501 (2010).
Adams, P.D. et al. PHENIX: a comprehensive Python-based system for macromolecular structure solution. Acta Crystallogr. D. Biol. Crystallogr. 66, 213–221 (2010).
Nicholls, R.A., Long, F. & Murshudov, G.N. Low-resolution refinement tools in REFMAC5. Acta Crystallogr. D. Biol. Crystallogr. 68, 404–417 (2012).
Petoukhov, M.V. et al. New developments in the ATSAS program package for small-angle scattering data analysis. J. Appl. Crystallogr. 45, 342–350 (2012).
Lindahl, E., Azuara, C., Koehl, P. & Delarue, M. NOMAD-Ref: visualization, deformation and refinement of macromolecular structures based on all-atom normal mode analysis. Nucleic Acids Res. 34, W52–W56 (2006).
Ludtke, S.J., Baldwin, P.R. & Chiu, W. EMAN: Semiautomated software for high-resolution single-particle reconstructions. J. Struct. Biol. 128, 82–97 (1999).
Mindell, J.A. & Grigorieff, N. Accurate determination of local defocus and specimen tilt in electron microscopy. J. Struct. Biol. 142, 334–347 (2003).
van Heel, M., Harauz, G., Orlova, E.V., Schmidt, R. & Schatz, M. A new generation of the IMAGIC image processing system. J. Struct. Biol. 116, 17–24 (1996).
Fuss, I.J., Kanof, M.E., Smith, P.D. & Zola, H. Isolation of whole mononuclear cells from peripheral blood and cord blood. Curr. Protoc. Immunol. 85, 7.1.1–7.1.8 (2009).
Pouliot, P., Willart, M.A., Hammad, H. & Lambrecht, B.N. Studying the function of dendritic cells in mouse models of asthma. Methods Mol. Biol. 595, 331–349 (2010).
Salic, A. & Mitchison, T.J. A chemical method for fast and sensitive detection of DNA synthesis in vivo. Proc. Natl. Acad. Sci. USA 105, 2415–2420 (2008).
Acknowledgements
We thank the Swiss Light Source (SLS), the European Synchrotron Radiation Facility (ESRF) and the Deutsches Elektronen-Synchrotron/European Molecular Biology Laboratory (DESY/EMBL) for synchrotron-beam time allocation and the staff of beamlines X06SA and X06DA (SLS), ID-29 and ID14-3 (ESRF) and X33 (DESY/EMBL) for technical support. Access to these synchrotron facilities is supported by the European Commission under the Seventh Framework Programme: Research Infrastructures, grant agreement number 226716. This research project was supported by grants from the Research Foundation Flanders (FWO) (3G064307, G059710 and G0B7912N) and Ghent University (BOF instrument) to S.N.S.; a Bundesministerium für Bildung und Forschung (BMBF) research grant Sync-Life (Contract: 05K10YEA) to D.I.S.; an ANR-MIME-2006 grant to N.T. and W.P.B.; and an FWO-Odysseus grant to B.N.L. J.E. would like to acknowledge the joint 2011 APS/CCP4 School for assistance with crystallographic structure refinement. We thank the SWITCH Laboratory at VIB-Belgium for access to FFF-MALS facilities and assistance during data collection and analysis. We also thank F. Wang (Harvard University, Boston, Massachusetts, USA) for providing cDNA of rhBARF1 and Christian Gorba (EMBL Hamburg, Hamburg, Germany) for help with the NMA refinement. I.G. thanks the EM platform of the Partnership for Structural Biology (Grenoble) for access to EM equipment. J.E., K.V. and B.V. are supported as research fellows of the FWO. P.P. is a Marie Curie (FP7) post-doctoral fellow.
Author information
Authors and Affiliations
Contributions
J.E. and N.B. expressed and purified recombinant proteins, with contributions from A.B. J.E. carried out all crystallographic studies with contributions from N.B., A.B. and S.N.S. J.E., N.B. and B.V. carried out ITC with contributions from S.N.S. N.T. and W.P.B. carried out preliminary structural studies on independently produced material. J.E., N.B. and B.V. carried out SPR studies with contributions from N.T. and W.P.B. K.V. provided tools and assistance for tissue culture. P.P. and B.N.L. carried out cellular assays. J.E., A.V.S. and D.I.S. carried out SAXS studies. I.G. carried out EM imaging and data analysis. J.E., N.B., B.V. and S.N.S. established the experimental approach and designed experiments. S.N.S. directed the study. J.E. and S.N.S. wrote the manuscript with contributions from all authors.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–7 and Supplementary Tables 1–3. (PDF 7905 kb)
Rights and permissions
About this article
Cite this article
Elegheert, J., Bracke, N., Pouliot, P. et al. Allosteric competitive inactivation of hematopoietic CSF-1 signaling by the viral decoy receptor BARF1. Nat Struct Mol Biol 19, 938–947 (2012). https://doi.org/10.1038/nsmb.2367
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nsmb.2367
This article is cited by
-
Activating mutations in CSF1R and additional receptor tyrosine kinases in histiocytic neoplasms
Nature Medicine (2019)
-
Mechanisms of immunomodulation by mammalian and viral decoy receptors: insights from structures
Nature Reviews Immunology (2017)
-
Structural basis of GM-CSF and IL-2 sequestration by the viral decoy receptor GIF
Nature Communications (2016)
-
Extracellular assembly and activation principles of oncogenic class III receptor tyrosine kinases
Nature Reviews Cancer (2012)