Abstract
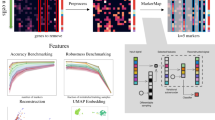
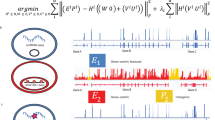
Recent experimental advances have enabled high-throughput single-cell measurement of gene expression, chromatin accessibility and DNA methylation. We previously employed integrative non-negative matrix factorization (iNMF) to jointly align multiple single-cell datasets (\(X_i\)) and learn interpretable low-dimensional representations using dataset-specific (\(V_i)\) and shared metagene factors (W) and cell factor loadings (\(H_i\)). We developed an alternating nonnegative least squares (ANLS) algorithm to solve the iNMF optimization problem [2]:
Access this chapter
Tax calculation will be finalised at checkout
Purchases are for personal use only
Similar content being viewed by others
References
Mairal, J., Bach, F., Ponce, J., Sapiro, G.: Online learning for matrix factorization and sparse coding. J. Mach. Learn. Res. 11(Jan), 19–60 (2010)
Welch, J.D., Kozareva, V., Ferreira, A., Vanderburg, C., Martin, C., Macosko, E.Z.: Single-cell multi-omic integration compares and contrasts features of brain cell identity. Cell 177(7), 1873–1887 (2019)
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2020 Springer Nature Switzerland AG
About this paper
Cite this paper
Gao, C., Welch, J.D. (2020). Iterative Refinement of Cellular Identity from Single-Cell Data Using Online Learning. In: Schwartz, R. (eds) Research in Computational Molecular Biology. RECOMB 2020. Lecture Notes in Computer Science(), vol 12074. Springer, Cham. https://doi.org/10.1007/978-3-030-45257-5_24
Download citation
DOI: https://doi.org/10.1007/978-3-030-45257-5_24
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-030-45256-8
Online ISBN: 978-3-030-45257-5
eBook Packages: Computer ScienceComputer Science (R0)