Summary
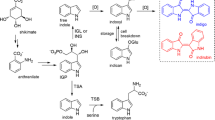
Although indole-3-acetic acid (IAA) is a well-known plant hormone, the main IAA biosynthetic pathway from l-tryptophan (Trp) via indole-3-pyruvic acid (IPyA) has yet to be elucidated. Previous studies have suggested that IAA is produced by Enterobacter cloacae isolated from the rhizosphere of cucumbers and its biosynthetic pathway may possibly be the same as that in plants. To elucidate this pathway, the IAA biosynthetic gene was isolated from a genomic library of E. cloacae by assaying for the ability to convert Trp to IAA. DNA sequence analysis showed that this gene codes for only one enzyme and its predicted protein sequence has extensive homology with pyruvate decarboxylase in yeast and Zymomonas mobilis. Cell-free extracts prepared from Escherichia coli harboring this gene could convert IPyA to indole-3-acetaldehyde (IAAld). These results clearly show that this pathway is mediated only by indolepyruvate decarboxylase, which catalyzes the conversion of IPyA to IAAld.
Similar content being viewed by others
References
Adelberg EA, Mandel M, Chen GCC (1965) Optimal conditions for mutagenesis by N-methyl-N′-nitro-N-nitrosoguanidine in Escherichia coli K12. Biochem Biophys Res Commun 18:788–795
Aragozzini F, Ferrari A, Pacini N, Gualandris R (1979) Indole-3-lactic acid as a tryptophan metabolite produced by Bifidobacterium spp. Appl Environ Microbiol 38:544–546
Atsumi S (1980) Studies on the production of indole-3-acetic acid with auxin-heterotrophic mutants derived from cultured crown gall cells. Plant Cell Physiol 21:1227–1232
Brenner DJ, Farmer III JJ, Frederiksen W, Le Minor L, Sakazaki R (1984) Facultatively anaerobic gram-negative rods. In: Krieg NR (ed) Bergey's manual of systematic bacteriology, vol 1. Williams & Wilkins, Baltimore, Md., pp 408–516
Comai L, Kosuge T (1980) Involvement of plasmid deoxyribonucleic acid in indoleacetic acid synthesis in Pseudomonas savastanoi. J Bacteriol 143:950–957
Comai L, Kosuge T (1982) Cloning and characterization of iaaM, a virulence determinant of Pseudomonas savastanoi. J Bacteriol 149:40–46
Ditta G, Stanfield S, Corbin D, Helinski DR (1980) Broad host range DNA cloning system for gram-negative bacteria: Construction of a gene bank Rhizobium meliloti. Proc Natl Acad Sci USA 77:7347–7351
Follin A, Inzé D, Budar F, Genetello C, Van Montagu M, Shell J (1985) Genetic evidence that the tryptophan 2-monooxygenase gene of Pseudomonas savastanoi is functionally equivalent to one of the T-DNA genes involved in plant tumor formation by Agrobacterium tumefaciens. Mol Gen Genet 201:178–185
Garcia-Tabares F, Herraiz-Tomico T, Amat-Guerri F, Garcia-Bilbao JL (1987) Production of 3-indoleacetic acid and 3-indolelactic acid in Azotobacter vinelandii cultures supplemented with tryptophan. Appl Microbiol Biotechnol 25:502–506
Gibson RA, Schneider EA, Wightman F (1972) Biosynthesis and metabolism of indol-3yl-acetic acid. II. In vivo experiments with 14C-labelled precursors of IAA in tomato and barley shoots. J Exp Bot 23:381–399
Gounaris AD, Turkenkopf I, Buckwald S, Young A (1971) Pyruvate decarboxylase. I. Protein dissociation into subunits under conditions in which thiamine pyrophosphate is released. J Biol Chem 246:1302–1309
Gunsalus IC, Stamer JR (1955) Transaminases in bacteria. Methods Enzymol 2:170–177
Jean M, De Moss RD (1968) Indolelactate dehydrogenase from Clostridium sporogenes. Can J Microbiol 14:429–435
Kaper JM, Verdstra H (1958) On the metabolism of tryptophan by Agrobacterium tumefaciens. Biochem Biophys Acta 30:401–420
Kellermann E, Seeboth PG, Hollenberg CP (1986) Analysis of the primary structure and promoter function of a pyruvate decarboxylase gene (PDCI) from Saccharomyces cerevisiae. Nucleic Acids Res 14:8963–8977
Kenten RH, Mann PJG (1952) The oxidation of amines by extracts of pea seedlings. Biochem J 50:360–369
Koga J, Adachi T, Hidaka H (1991) IAA biosynthetic pathway from tryptophan via indole-3-pyruvic acid in Enterobacter cloacae. Agric Biol Chem 55:701–706
Kosuge T, Heskett MG, Wilson EE (1966) Microbial synthesis and degradation of indole-3-acetic acid. J Biol Chem 241:3738–3744
Kuo DJ, Dikdan G, Jordan F (1986) Resolution of Brewers' yeast pyruvate decarboxylase into two isozymes. J Biol Chem 261:3316–3319
Laemmli UK (1970) Cleavage of structural proteins during the assembly of the head of bacteriophage T4. Nature 227:680–685
Maniatis T, Fritsch EF, Sambrook J (1982) Molecular cloning: a laboratory manual. Cold Spring Harbor Laboratory Press, Cold Spring Harbor, New York
Moore TC, Sanger CA (1968) Synthesis of indoleacetic acid from tryptophan via indolepyruvic acid in cell-free extracts of pea seedlings. Arch Biochem Biophys 127:613–621
Neale AD, Scopes RK, Wettenhall REH, Hoogenraad NJ (1987a) Pyruvate decarboxylase of Zymomonas mobilis: Isolation, properties, and genetic expression in Escherichia coli. J Bacteriol 169:1024–1028
Neale AD, Scopes RK, Wettenhall REH, Hoogenraad NJ (1987b) Nucleotide sequence of the pyruvate decarboxylase gene from Zymomonas mobilis. Nucleic Acids Res 15:1753–1761
Paris CG, Magasanik B (1981) Tryptophan metabolism in Klebsiella aerogenes: regulation of the utilization of aromatic amino acids as sources of nitrogen. J Bacteriol 145:257–265
Sanger F, Nicklen S, Coulson AR (1977) DNA sequencing with chain-terminating inhibitors. Proc Natl Acad Sci USA 74:5463–5467
Schröder G, Waffenschmidt S, Weiler EW, Schroder J (1984) The T-region of Ti plasmids codes for an enzyme synthesizing indole-3-acetic acid. Eur J Biochem 138:387–391
Sheldrake AR (1973) The production of hormones in higher plants. Biol Rev 48:509–559
Sone H, Fujii T, Kondo K, Tanaka J (1987) Molecular cloning of the gene encoding α-acetolactate decarboxylase from Enterobacter aerogenes. J Biotechnol 5:87–91
Thomashow LS, Reeves S, Thomashow MF (1984) Crown gall oncogenesis: Evidence that a T-DNA gene from the Agrobacterium Ti plasmid pTiA6 encodes an enzyme that catalyzes synthesis of indoleacetic acid. Proc Natl Acad Sci USA 81:5071–5075
Thomashow MF, Hugly S, Buchholz WG, Thomashow LS (1986) Molecular basis for the auxin-independent phenotype of crown gall tumor tissues. Science 231:616–618
Truelsen TA (1973) Indole-3-pyruvic acid as a intermediate in the conversion of tryptophan to indole-3-acetic acid. II. Distribution of tryptophan transaminase activity in plants. Physiol Plant 28:67–70
Yamada T, Palm CJ, Brooks B, Kosuge T (1985) Nucleotide sequences of the Pseudomonas savastanoi indoleacetic acid genes show homology with Agrobacterium tumefaciens T-DNA. Proc Natl Acad Sci USA 82:6522–6526
Author information
Authors and Affiliations
Additional information
Communicated by J. Schell
Rights and permissions
About this article
Cite this article
Koga, J., Adachi, T. & Hidaka, H. Molecular cloning of the gene for indolepyruvate decarboxylase from Enterobacter cloacae . Mol Gen Genet 226, 10–16 (1991). https://doi.org/10.1007/BF00273581
Received:
Issue Date:
DOI: https://doi.org/10.1007/BF00273581