Abstract
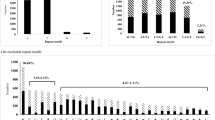
Expressed sequence tags (ESTs) are important resources for gene discovery and molecular marker development. From over 147,000 ESTs of Medicago truncatula, we have identified 4,384 ESTs containing perfect simple sequence repeats (EST-SSR) of di-, tri-, tetra- or pentanucleotides. Six hundred sixteen primer pairs (PPs) were designed and screened over a panel of eight genotypes representing six Medicago spp. and subspecies. Nearly, 74% (455) of the PPs produced characteristic SSR bands of expected size length in at least one Medicago species. Four hundred six (89%) of these 455 PPs produced SSR bands in all eight genotypes tested. Only 17 PPs were M. truncatula -specific. High levels of polymorphism (>70%) were detected for these markers in alfalfa, M. truncatula, and other annual medics. About 48% of the reported markers are part of gene transcripts linked to putative functions. Our results indicate that the SSR markers developed from M. truncatula ESTs are valuable genetic markers for the Medicago genus. These markers will be useful in establishing the genomic relationships of M. truncatula to important forage legume crops such as alfalfa and other annual medics.





Similar content being viewed by others
References
Anderson JA, Churchill GA, Sutrique JE, Tanksley SD, Sorrells ME (1993) Optimizing parental selection for genetic linkage maps. Genome 36:181–186
Ayers NM, McClung AM, Larkin PD, Bligh HFJ, Jones CA, Park WD (1997) Microsatellite and single nucleotide polymorphism differentiate apparent amylose classes in an extended pedigree of US rice germplasm. Theor Appl Genet 94:773–781
Baquerizo-Audiot E, Desplanque B, Prosperi JM, Santoni S (2001) Characterization of microsatellite loci in the diploid legume Medicago truncatula (barrel medic). Mol Ecol Notes 1:1–3
Barnes DK, Goplen BP, Baylor JE (1988) Highlights in the USA and Canada. In: Hanson AA, Barnes DK, Hill RR (eds) Alfalfa and alfalfa improvement. Agronomy Monograph No. 29, American Society of Agronomy – Crop Science Society of America– Soil Science Society of America, Madison, Wis., USA, pp1–24
Botstein D, White RL, Skolnick M, Davis RW (1980) Construction of genetic linkage map in man using restriction fragment length polymorphisms. Am J Hum Genet 32:314–331
Brown-Guedira GL, Thomson JA, Nelson RL, Warburton ML (2000) Evaluation of genetic diversity of soybean introductions and North American ancestors using RAPD and SSR markers. Crop Sci 40:815–823
Cho YG, Ishii T, Temnykh S, Chen X, Lipovich L, McCouch SR, Parl WD, Ayers N, Cartinhour S (2000) Diversity of microsatellites derived from genomic libraries and GenBank sequences in rice (Oryza sativa L.). Theor Appl Genet 100:713–722
Danesh D, Shekhawat NS, Cardinet G, Thoquet P, Mudge J, Penuela S, Kim D, Kiss G, Choi H, Limpens E, Zeyen R, Huguet T, Cook DR, Young ND (2002) Integrated microsatellite marker mapping and powdery mildew resistance in Medicago truncatula (abstract). In: Plant and Animal Genome X. The International Conference on the Status of Plant and Animal Genome Research, 12–16 January 2002, San Diego, Calif., pp 196
Diwan N, Bhaqwat AA, Bauchan GB, Cregan PB (1997) Simple sequence repeats DNA markers in alfalfa and perennial and annual Medicago species. Genome 40:887–895
Diwan N, Bouton JH, Kochert G, Cregan PB (2000) Mapping of simple sequence repeat (SSR) DNA markers in diploid and tetraploid alfalfa. Theor Appl Genet 101:165–172
Doyle JJ, Doyle JL, Ballenger JA, Palmer JD (1996) The distribution and phylogenetic significance of a 50-kb chloroplast DNA inversion in the flowering plant family Leguminosae. Mol Phylogenet Evol 5:429–438
Eujayl I, Sorrells ME, Wolters P, Baum M, Powell W (2002) Isolation of EST- derived microsatellite markers for genotyping the A and B genomes of wheat. Theor Appl Genet 104:399–407
Gaitán-Solís E, Duque MC, Edwards KJ, Tohme J (2002) Microsatellite repeats in common bean (Phaseolus vulgaris): isolation, characterization, and cross-species amplification in Phaseolus ssp. Crop Sci 42:2128–2136
Huguet T, Thoquet P, Gherardi M, Kereszt A, Ane JM, Vilotte L, Cardinet G, Baquerizo E, Santoni S, Prosperi JM (2001) The molecular linkage map of the model legume Medicago truncatula: a tool for legume genome comparison and gene mapping. In: 4th Workshop on Medicago truncatula. 7–10 July 2001, University of Wisconsin-Madison, Wis., pp 51
Kaló P, Endre G, Zimnyi L, Csanadi G, Kiss GB (2000) Construction of an improved linkage map of diploid alfalfa (Medicago sativa). Theor Appl Genet 100:641–657
Kantety RV, Rota ML, Matthews DE, Sorrells ME (2002) Data mining for simple-sequence repeats in expressed sequence tags from barley, maize, rice, sorghum, and wheat. Plant Mol Biol 48:501–510
Kilian A, Chen J, Han F, Steffenson B, Kleinhofs A (1997) Towards map-based cloning of the barley stem rust resistance genes Rpg1 and rpg4 using rice as an intergenomic cloning vehicle. Plant Mol Biol 35:187–195
Laurent P, Elduque C, Hayes H, Saunier K, Eggen A, Leveziel H (2000) Assignment of 60 human ESTs in cattle. Mammal Genome 11:748–754
Nei M, Li WH (1979) Mathematical model for studying genetic variation in terms of restriction endonucleases. Proc Natl Acad Sci 76:5269–5273
Paterson AH, Lin YR, Li S, Schertz KF, Doebley JF, Pinson SRM, Liu SC, Stansel JW, Irvine JE (1995) Convergent domestication of cereal crops by independent mutations at corresponding genetic loci. Science 269:1714–1717
Peakall R, Gilmore S, Keys W, Morgante M, Rafalski A (1998) Cross species amplification of soybean (Glycine max) simple sequence repeat (SSRs) within the genus and other legume genera: implication for transferability of SSRs in plants. Mol Biol Evol 15:1275–1287
Rebeiz M, Lewin HA (2000) COMPASS of 47,787 cattle ESTs. Anim Biotechnol 11:175–241
Roa AC, Chavarriaga-Aguirre P, Duque MC, Maya MM, Bonierbale MW, Iglesias C, Tohme J (2000) Cross-species amplification of cassava (Manihot esculenta) (Euphorbiaceae) microsatellites: allelic polymorphism and degree of relationship. Am J Bot 87:1647–1655
SAS Institute (1989) SAS/STAT user's guide, version 6, 4th edn, vol 1. SAS Institute, Cary, N.C.
Scott KD, Eggler P, Seaton G, Rossetto M, Ablett EM, Lee LS, Henry RJ (2000) Analysis of SSRs derived from grape ESTs. Theor Appl Genet 100:723–726
Temnykh S, Park WD, Ayers N, Cartinhour S, Hauck N, Lipovich L, Cho YG, Ishii T, McCouch SR (2000) Mapping and genome organization of microsatellite sequences in rice (Oryza sativa L.). Theor Appl Genet 100:697–712
Thoquet P, Ghérardi M, Journet E, Kereszt A, Ané JM, Prosperi JM, Huguet T (2002) The molecular genetic linkage map of the model legume Medicago truncatula: an essential tool for comparative legume genomics and the isolation of argonomically important genes. BMC Plant Biology 2:1–19
Tugendreich S, Boguski M, Seldini MS, Hieter P (1993) Linking yeast genetics to mammalian genomes: identification and mapping of the human homolog of CDC27 via the expressed sequence tag (EST) database. Proc Natl Acad Sci 90:10031–10035
Weber JL (1990) Informativeness of human (dC-dA)n (dG-dT)n polymorphisms. Genomics 7:524–530
White G, Powell W (1997) Isolation and characterization of microsatellite loci in Swietenia humilis (Meliaceae): an endangered tropical hardwood species. Mol Ecol 6:851–860
Wu KK, Burnquist W, Sorrells ME, Tew TL, Moore PH, Tanksley SD (1992) The detection and estimation of linkage in polyploids using single dose restriction fragments. Theor Appl Genet 83:294–300
Acknowledgements
We thank Drs. Douglas Cook and Ian Ray for the plant materials of their respective mapping populations used in this study. This research work was funded by The Samuel Roberts Noble Foundation.
Author information
Authors and Affiliations
Corresponding author
Additional information
Communicated by H.C. Becker
Electronic Supplementary Material
Rights and permissions
About this article
Cite this article
Eujayl, I., Sledge, M.K., Wang, L. et al. Medicago truncatula EST-SSRs reveal cross-species genetic markers for Medicago spp.. Theor Appl Genet 108, 414–422 (2004). https://doi.org/10.1007/s00122-003-1450-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00122-003-1450-6