Abstract
Rapeseed (Brassica napus L.) is one of the major oilseed crops and an important source for tocopherols known as vitamin E in human nutrition. Increasing the tocopherol content and altering the tocopherol composition is a major goal in rapeseed breeding. The genes encoding enzymes from the tocopherol pathway have been cloned from model species. However, only scant data about tocopherol genes from crop species have been available. We have cloned four sequences of a gene family from B. napus with homology to the Arabidopsis thaliana VTE4 gene. The sequences were amplified by PCR with primers derived from the A. thaliana gene. BAC-clones were isolated to analyze the genomic structure of the BnaX.VTE4-loci. In contrast to the A. thaliana gene all B. napus sequences have two additional introns. For functional analysis, the BnaA.VTE4.a1 sequence was transformed into A. thaliana. Seeds from transgenic offspring showed a 50-fold increase of the α-tocopherol fraction which is in accordance with the predicted function of the gene. A marker assay was established and the BnaA.VTE4.a1 sequence was mapped to the end of chromosome A02 of the Tapidor × Ningyou7 genetic map, where also two QTL for α-tocopherol content had been mapped. Thus, the BnaA.VTE4.a1 gene is a promising candidate for these QTL and can be used for marker assisted selection for α-tocopherol content in rapeseed.





Similar content being viewed by others
References
Andersen JR, Lübberstedt T (2003) Functional markers in plants. Trends Plant Sci 8:554–560
Bergmüller E, Porfirova S, Dörmann P (2003) Characterization of an Arabidopsis mutant deficient in γ-tocopherolmethyltransferase. Plant Mol Biol 52:1181–1190
Brunel D, Froger N, Pelletier G (1999) Development of amplified consensus genetic markers (ACGM) in Brassica napus from Arabidopsis thaliana sequences of known biological function. Genome 42:387–402
Cheng Z, Sattler S, Maeda H, Sakuragi Y, Bryant DA, DellaPenna D (2003) Highly divergent methyltransferases catalyze a conserved reaction in tocopherol and plastoquinone synthesis in cyanobacteria and photosynthetic eukaryotes. Plant Cell 15:2343–2356
Collakova E, DellaPenna D (2001) Isolation and functional analysis of homogentisate phytyltransferase from Synechocystis sp. PCC 6803 and Arabidopsis. Plant Physiol 127:1113–1124
Dähnhardt D, Falk J, Appel J, van der Kooij TA, Schulz-Friedrich R, Krupinska K (2002) The hydroxyphenylpyruvate dioxygenase from Synechocystis sp. PCC 6803 is not required for plastoquinone biosynthesis. FEBS Lett 523:177–181
Dolde D, Vlahakis C, Hazebroek J (1999) Tocopherols in breeding lines and effects of planting location, fatty acid composition and temperature during development. JAOCS 76:349–355
Emanuelsson O, Nielsen H, Brunak S, von Heijne G (2000) Predicting subcellular localization of proteins based on their N-terminal amino acid sequence. J Mol Biol 300:1005–1016
Falk J, Andersen G, Kernebeck B, Krupinska K (2003) Constitutive overexpression of barley 4-hydroxyphenylpyruvate dioxygenase in tobacco results in elevation of the vitamin E content in seeds but not in leaves. FEBS Lett 540:35–40
Feinberg AP, Vogelstein B (1983) A technique for radiolabeling DNA restriction endonuclease fragments to high specific activity. Anal Biochem 132:6–13
Fourmann M, Barret P, Froger N, Baron C, Charlot F, Delourme R, Brunel D (2002) From Arabidopsis thaliana to Brassica napus: development of amplified consensus genetic markers (ACGM) for construction of a gene map. Theor Appl Genet 105:1196–1206
Gilliland LU, Magallanes-Lundback M, Hemming C, Supplee A, Koornneef M, Bentsink L, DellaPenna D (2006) Genetic basis for natural variation in seed vitamin E levels in Arabidopsis thaliana. PNAS. doi:10.1073/pnas.0606221103
Goffman FD, Becker HC (1999) Inheritance of tocopherol contents in seeds of rapeseed (Brassica napus L.). Proceedings of the 10th rapeseed congress, Canberra, Australia
Goffman FD, Becker HC (2002) Genetic variation of tocopherol content in a germplasm collection of Brassica napus L. Euphytica 125:189–196
Kifle S, Shao M, Jung C, Cai D (1999) An improved transformation protocol for studying gene expression in hairy roots of sugar beet (Beta vulgaris L.). Plant Cell Rep 18:514–519
Koncz C, Schell J (1986) The promoter of TL-DNA gene 5 controls the tissue-specific expression of chimaeric genes carried by a novel type of Agrobacterium binary vector. Mol Gen Genet MGG 204:383–396
Kumar R, Raclaru M, Schusseler T, Gruber J, Sadre R, Lühs W, Zarhloul KM, Friedt W, Enders D, Frentzen M, Weier D (2005) Characterisation of plant tocopherol cyclases and their overexpression in transgenic Brassica napus seeds. FEBS Lett 579:1357–1364
Lagercrantz U, Lydiate DJ (1996) Comparative genome mapping in Brassica. Genetics 144:1903–1910
Long Y, Shi J, Qiu D, Li R, Zhang C, Wang J, Hou J, Zhao J, Shi L, Park BS, Choi SR, Lim YP, Meng J (2007) Flowering time quantitative trait loci analysis of oilseed Brassica in multiple environments and genomewide alignment with Arabidopsis. Genetics 177:2433–2444
Marwede V, Schierholt A, Mollers C, Becker HC (2004) Genotype × environment interactions and heritability of tocopherol contents in canola. Crop Sci 44:728–731
Marwede V, Gul MK, Becker HC, Ecke W (2005) Mapping of QTL controlling tocopherol content in winter oilseed rape. Plant Breed 124:20–26
Motohashi R, Ito T, Kobayashi M, Taji T, Nagata N, Asami T, Yoshida S, Yamaguchi-Shinozaki K, Shinozaki K (2003) Functional analysis of the 37 kDa inner envelope membrane polypeptide in chloroplast biogenesis using a Ds-tagged Arabidopsis pale-green mutant. Plant J 34:719–731
Nielsen H, Engelbrecht J, Brunak S, von Heijne V (1997) Identification of prokaryotic and eukaryotic signal peptides and prediction of their cleavage sites. Protein Eng 10:1–6
Norris SR, Shen X, DellaPenna D (1998) Complementation of the Arabidopsis pds1 mutation with the gene encoding p-hydroxyphenylpyruvate dioxygenase. Plant Physiol 117:1317–1323
Parkin IA, Lydiate DJ, Trick M (2002) Assessing the level of collinearity between Arabidopsis thaliana and Brassica napus for A. thaliana chromosome 5. Genome 45:356–366
Parkin IA, Sharpe AG, Lydiate DJ (2003) Patterns of genome duplication within the Brassica napus genome. Genome 46:291–303
Parkin IA, Gulden SM, Sharpe AG, Lukens L, Trick M, Osborn TC, Lydiate DJ (2005) Segmental structure of the Brassica napus genome based on comparative analysis with Arabidopsis thaliana. Genetics 171:765–781
Paterson AH, Lan TH, Amasino R, Osborn TC, Quiros C (2001) Brassica genomics: a complement to, and early beneficiary of, the Arabidopsis sequence. Genome Biol 2: reviews1011.1–reviews1011.4
Pongracz G, Weiser H, Matziger D (1995) Tocopherole-Antioxidantien der Natur. Fat Sci Technol 3:90–104
Porfirova S, Bergmüller E, Tropf S, Lemke R, Dörmann P (2002) Isolation of an Arabidopsis mutant lacking vitamin E and identification of a cyclase essential for all tocopherol biosynthesis. Proc Natl Acad Sci USA 99:12495–12500
Raclaru M, Gruber J, Kumar R, Sadre R, Lühs W, Zarhloul M, Friedt W, Frentzen M, Weier D (2006) Increase of the tocochromanol content in transgenic Brassica napus seeds by overexpression of key enzymes involved in prenylquinone biosynthesis. Mol Breed 18:93–107
Rana D, Boogaart T, O’Neill CM, Hynes L, Bent E, Macpherson L, Park JY, Lim YP, Bancroft I (2004) Conservation of the microstructure of genome segments in Brassica napus and its diploid relatives. Plant J 40:725–733
Saghai-Maroof MA, Soliman KM, Jorgensen RA, Allard RW (1984) Ribosomal DNA spacer-length polymorphisms in barley: Mendelian inheritance, chromosomal location, and population dynamics. PNAS 81:8014–8018
Schledz M, Seidler A, Beyer P, Neuhaus G (2001) A novel phytyltransferase from Synechocystis sp. PCC 6803 involved in tocopherol biosynthesis. FEBS Lett 499:15–20
Shintani D, DellaPenna D (1998) Elevating the vitamin E content of plants through metabolic engineering. Science 282:2098–2100
Soll J, Kemmerling M, Schultz G (1980) Tocopherol and plastoquinone synthesis in spinach chloroplasts subfractions. Arch Biochem Biophys 204:544–550
Valvekens D, Montagu MV, Lijsebettens MV (1988) Agrobacterium tumefaciens-mediated transformation of Arabidopsis thaliana root explants by using kanamycin selection. PNAS 85:5536–5540
Van Eenennaam AL, Lincoln K, Durrett TP, Valentin HE, Shewmaker CK, Thorne GM, Jiang J, Baszis SR, Levering CK, Aasen ED, Hao M, Stein JC, Norris SR, Last RL (2003) Engineering vitamin E content: from Arabidopsis mutant to soy oil. Plant Cell 15:3007–3019
Van Ooijen JW, Voorrips RE (2001) JoinMap® 3.0. Software for the calculation of genetic linkage maps. Plant Research International, Wageningen, The Netherlands
Acknowledgments
We wish to thank Martina Bach and Jens Hermann for their excellent technical assistance. This work was funded by the Stiftung Schleswig-Holsteinische Landschaft and the German Research Foundation (DFG) as part of the Research Training Group GRK820.
Author information
Authors and Affiliations
Corresponding author
Additional information
Communicated by H. Becker.
The nucleotide sequences data reported in this paper have been submitted to GenBank nucleotide sequence database with the accession numbers EU637012 to EU637015 and FJ435091 to FJ435093.
Electronic supplementary material
Below is the link to the electronic supplementary material.
122_2009_1066_MOESM1_ESM.ppt
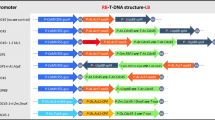
S.1: Sequence alignment of the amplified regions using the primer combination vte4-4fw/vte4-4rv from three different BAC-derived BnaX.VTE4 sequences. Primer orientation is indicated by blue arrows. In red: deletion used to distinguish the BnaA.VTE4.a2 sequence from Tapidor from the other sequence variants by length polymorphism. Supplementary material 1 (PPT 105 kb)
Rights and permissions
About this article
Cite this article
Endrigkeit, J., Wang, X., Cai, D. et al. Genetic mapping, cloning, and functional characterization of the BnaX.VTE4 gene encoding a γ-tocopherol methyltransferase from oilseed rape. Theor Appl Genet 119, 567–575 (2009). https://doi.org/10.1007/s00122-009-1066-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00122-009-1066-6