Abstract
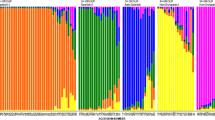
In order to determine how informative a set of microsatellites from tomato is across the genus Lycopersicon, 17 microsatellite loci, derived from regions in and around genes, were tested on 31 accessions comprising the nine species of the genus. The microsatellite polymorphisms were used to estimate the distribution of diversity throughout the genus and to evaluate the efficacy of microsatellites for establishing species relationships in comparison with existing phylogeny reconstructions. Gene diversity and genetic distances were calculated. A high level of polymorphism was found, as well as a large number of alleles unique for species. The level of polymorphism detected with the microsatellite loci within and among species was highly correlated with the respective mating systems, cross-pollinating species having a significantly higher gene diversity compared to self-pollinating species. In general, microsatellite-based trees were consistent with a published RFLP-based dendrogram as well as with a published classification based on morphology and the mating system. A tree constructed with low-polymorphic loci (gene diversity <0.245) was shown to represent a more-reliable topology than a tree constructed with more-highly polymorphic loci.
Similar content being viewed by others
Author information
Authors and Affiliations
Additional information
Received: 19 February 2001 / Accepted: 26 March 2001
Rights and permissions
About this article
Cite this article
Alvarez, A., van de Wiel, C., Smulders, M. et al. Use of microsatellites to evaluate genetic diversity and species relationships in the genus Lycopersicon. Theor Appl Genet 103, 1283–1292 (2001). https://doi.org/10.1007/s001220100662
Issue Date:
DOI: https://doi.org/10.1007/s001220100662