Abstract
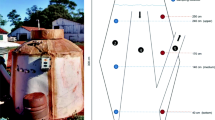
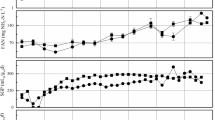
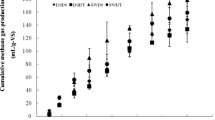
Anaerobic digestion is an alternative method for the treatment of animal manure and wastewater. The anaerobic bioconversion of biomass requires a multi-step biological process, including microorganisms with distinct roles. The diversity and composition of microbial structure in pilot-scale anaerobic digestion operating at ambient temperature in Brazil were studied. Influence of the seasonal and temporal patterns on bacterial and archaeal communities were assessed by studying the variations in density, dynamic and diversity and structure. The average daily biogas produced in the summer and winter months was 18.7 and 16 L day−1, respectively, and there was no difference in the average methane yield. Quantitative PCR analysis revealed that no differences in abundances and dynamics were found for bacterial communities and the total number of Archaea in different seasons. Analysis of bacterial clone libraries revealed a predominance of Firmicutes (54.5 %/summer and 46.7 %/winter) and Bacteroidetes (31.4 %/summer and 44.4 %/winter). Within the Archaea, the phylum Euryarchaeota was predominant in both digesters. Phylogenetic distribution showed changes in percentage between the phyla identified, but no alterations were recorded in the quality and amount of produced methane or community dynamics. The results may suggest that redundancy of microbial groups may have occurred, pointing to a more complex microbial community in the ecosystem related to this ambient temperature system.





Similar content being viewed by others
References
Guerci M, Knudsen MT, Bava L, Zucali M, Schönbach P, Kristensen T (2013) Parameters affecting the environmental impact of a range of dairy farming systems in Denmark, Germany and Italy. J Clean Prod 54:133–141
Abubaker J, Risberg K, Pell M (2012) Biogas residues as fertilisers—effects on wheat growth and soil microbial activities. Appl Energy 99:126–134
Holm-Nielsen JB, Seadi TA, Oleskowicz-Popiel P (2009) The future of anaerobic digestion and biogas utilization. Bioresour Technol 100:5478–5484
Kumar KV, Sridevi V, Rani K, Sakunthala M, Kumar CS (2013) A review on production of biogas, fundamentals, applications & its recent enhancing techniques. Elixir Chem Engg 57:14073–14079
Sundberg C, Al-Soud WA, Larsson M, Alm E, Yekta SS, Svensson BH, Sørensen SJ, Karlsson A (2013) 454 pyrosequencing analyses of bacterial and archaeal richness in 21 full-scale biogas digesters. FEMS Microbiol Ecol 85:612–626
Kunz A, Steinmetz RLR, Rammeb MA, Coldebella A (2009) Effect of storage time on swine manure solid separation efficiency by screening. Bioresour Technol 100:1815–1818
Levén L, Eriksson ARB, Schnurer A (2007) Effect of process temperature on bacterial and archaeal communities in two methanogenic bioreactors treating organic house holdwaste. FEMS Microbiol Ecol 59:683–693
Lee SH, Kang HJ, Lee YH, Lee TJ, Han K, Choic Y, Park HD (2012) Monitoring bacterial community structure and variability in time scale in full-scale anaerobic digesters. J Environ Monit 14:1893–1905
Ferrer I, Gamiz M, Almeida M, Ruiz A (2009) Pilot project of biogas production from pig manure and urine mixture at ambient temperature in Ventanilla (Lima, Peru). Waste Manag 29:168–173
Yusuf MOL, Debora A, Ogheneruona DE (2011) Ambient temperature kinetic assessment of biogas production from co-digestion of horse and cow dung. Res Agr Eng 57:97–104
Nikolaeva S, Sánchez E, Borja R (2013) Dairy wastewater treatment by anaerobic fixed bed reactors from laboratory to pilot-scale plant: a case study in Costa Rica operating at ambient temperature. Int J Environ Res 7:759–766
American Public Health Association (APHA) (2005) Standard methods for examination of water and wastewater, 21st edn. American Water Works Association, Washington, DC
Braun F, Hamelin J, Gévaudan G, Patureau D (2011) Development and application of an enzymatic and cell flotation treatment for the recovery of viable microbial cells from environmental matrices such as anaerobic sludge. Appl Environ Microbiol 77:8487–8493
Michelland RJ, Dejean S, Combes S, Fortun-Lamothe L, Cauquil L (2009) StatFingerprints: a friendly graphical interface program for processing and analysis of microbial fingerprint profiles. Mol Ecol Resour 9:1359–1363
Yu Y, Lee C, Kim J, Hwang S (2005) Group-specific primer and probe sets to detect methanogenic communities using quantitative real-time polymerase chain reaction. Biotechnol Bioeng 89:670–679
Tamaki H, Wright CL, Li X, Lin Q, Hwang C, Wang S, Thimmapuram J, Kamagata Y, Liu W (2011) Analysis of 16S rRNA amplicon sequencing options on the Roche/454 next-generation titanium sequencing platform. PLoS ONE 6:e25263
Alexander TW, Yanke JL, Reuter T, Topp E, Read RR, Selinger BL, McAllister TA (2011) Longitudinal characterization of antimicrobial resistance genes in feces shed from cattle fed different subtherapeutic antibiotics. BMC Microbiol 11:19
Klocke M, Mähnert P, Mundt K, Souidi K, Linke B (2007) Microbial community analysis of a biogas-producing completely stirred tank reactor fed continuously with fodder beet silage as mono-substrate. Syst Appl Microbiol 30:139–151
Moletta-Denat M, Bru-Adan V, Delgenès JP, Hamelin J, Wéry N, Godon JJ (2010) Selective microbial aerosolization in biogas demonstrated by quantitative PCR. Bioresour Technol 101:7252–7257
Hori T, Haruta S, Ueno Y, Ishii M, Igarashi Y (2006) Dynamic transition of a methanogenic population in response to the concentration of volatile fatty acids in a thermophilic anaerobic digester. Appl Environ Microbiol 72:1623–1630
Liu FH, Wang SB, Zhang JS, Zhang J, Yan X, Zhou HK, Zhao GP, Zhou ZH (2009) The structure of the bacterial and archaeal community in a biogas digester as revealed by denaturing gradient gel electrophoresis and 16S rDNA sequencing analysis. J Appl Microbiol 106:952–966
Peu P, Brugère H, Pourcher AM, Kérourédan M, Godon JJ, Delgenès JP, Dabert P (2006) Dynamics of a pig slurry microbial community during anaerobic storage and management. Appl Environ Microbiol 72:3578–3585
Bengelsdorf FR, Gerischer U, Langer S, Zak M, Kazda M (2013) Stability of a biogas-producing bacterial, archaeal and fungal community degrading food residues. FEMS Microbiol Ecol 84:201–212
Alvarez R, Villca S, Lidén G (2006) Biogas production from llama and cow manure at high altitude. Biomass Bioenerg 30:66–75
Hill DT, Taylor SE, Grift TE (2001) Simulation of low temperature anaerobic digestion of dairy and swine manure. Bioresour Technol 78:127–131
Rico JL, García H, Rico C, Tejero I (2007) Characterization of solid and liquid fractions of dairy manure with regard to their component distribution and methane production. Bioresour Technol 98:971–979
Chae KJ, Jang A, Yim SK, Kim IS (2008) The effects of digestion temperature and temperature shock on the biogas yields from the mesophilic anaerobic digestion of swine manure. Bioresour Technol 99:1–6
McGarvey JA, Miller WG, Zhang R, Ma Y, Mitloehner F (2007) Bacterial population dynamics in dairy waste during aerobic and anaerobic treatment and subsequent storage. Appl Environ Microbiol 73:193–202
Kampmann K, Ratering S, Kramer I, Schmidt M, Zerr W, Schnella S (2012) Unexpected stability of Bacteroidetes and Firmicutes communities in laboratory biogas reactors fed with different defined substrates. Appl Environ Microbiol 78:2106–2119
Krause L, Diaz NN, Edwards RA, Gartemann KH, Krömeke H, Neuweger H, Pühler A, Runte KJ, Schlüter A, Stoye J, Szczepanowski R, Tauch A, Goesmann A (2008) Taxonomic composition and gene content of a methane-producing microbial community isolated from a biogas reactor. J Biotechnol 136:91–101
Jaenicke S, Ander C, Bekel T, Bisdorf R, Dröge M (2011) Comparative and joint analysis of two metagenomic datasets from a biogas fermenter obtained by 454-pyrosequencing. PLoS ONE 6:e14519
Ritari J, Koskinen K, Hultman J, Kurola JM, Kymäläinen M, Romantschuk M, Paulin L, Auvinen P (2012) Molecular analysis of meso- and thermophilic microbiota associated with anaerobic biowaste degradation. BMC Microbiol 12:121
Simankova MV, Kotsyurbenko OR, Luedersb T, Nozhevnikova AN, Wagner B, Conrad R, Friedrich MW (2003) Isolation and characterization of new strains of methanogens from cold terrestrial habitats. Syst Appl Microbiol 26:312–318
Ma J, Zhao B, Frear C, Zhao Q, Yu L, Li X, Chen S (2013) Methanosarcina domination in anaerobic sequencing batch reactor at short hydraulic retention time. Bioresource Technol 137:41–50
Wang Y, Qian PY (2009) Conservative fragments in bacterial 16S rRNA genes and primer design for 16S ribosomal DNA amplicons in metagenomic studies. PloS ONE 4:e7401
Brosius J, Dull TJ, Sleeter DD, Noller HF (1981) Gene organization and primary structure of a ribosomal RNA operon from Escherichia coli. J Mol Biol 148:107–127
Acknowledgments
The authors are grateful to Kim Milferstedt, for support to the program database analysis. This study was supported by grants from EMBRAPA’s virtual Laboratory abroad (Labex Program) in Europe, Brazilian National Council for Scientific and Technological Development (CNPq), Research Support Foundation of Minas Gerais (FAPEMIG). The authors gratefully acknowledge Embrapa Dairy Cattle, Institut National de la Recherche Agronomique (INRA) and Federal University of Juiz de Fora (UFJF). We also thank Coordination of Improvement of the Personnel of Higher Education (CAPES) by the scholarship (Process 11853-12-3).
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
ESM 1
(DOCX 15 kb)
Rights and permissions
About this article
Cite this article
Resende, J.A., Godon, JJ., Bonnafous, A. et al. Seasonal Variation on Microbial Community and Methane Production during Anaerobic Digestion of Cattle Manure in Brazil. Microb Ecol 71, 735–746 (2016). https://doi.org/10.1007/s00248-015-0647-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00248-015-0647-y