Abstract
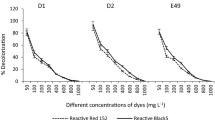
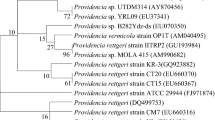
A broad-spectrum dye-decolorizing bacterium, strain DN322, was isolated from activated sludge of a textile printing wastewater treatment plant. The strain was characterized and identified as a member of Aeromonas hydrophila based on Gram staining, morphology characters, biochemical tests, and nearly complete sequence analysis of 16S rRNA gene and the gyrase subunit beta gene (gyrB). Strain DN322 decolorized a variety of synthetic dyes, including triphenylmethane, azo, and anthraquinone dyes. For color removal, the most suitable pH and temperature were pH 5.0–10.0 and 25–37°C, respectively. Triphenylmethane dye, e.g., Crystal Violet, Basic Fuchsin, Brilliant Green, and Malachite Green (50 mg l−1) were decolorized more than 90% within 10 h under aerobic culture condition and Crystal Violet could be used as sole carbon source and energy source for cell growth. The color removal of triphenylmethane dyes was due to a soluble cytosolic enzyme, and the enzyme was an NADH/NADPH-dependent oxygenase; For azo and anthraquinone dyes, e.g., Acid Amaranth, Great Red GR, Reactive Red KE-3B, and Reactive Brilliant Blue K-GR (50 mg l−1) could be decolorized more than 85% within 36 h under anoxic condition. This strain may be useful for bioremediation applications.







Similar content being viewed by others
References
AN SY, Min SK, Cha IH, Choi YL, Cho YS, Kim CH, Lee YC (2002) Decolorization of triphenylmethane and azo dyes by Citrobacter sp. Biotechnol Lett 24:1037–1040
Azmi W, Sani RK, Banerjee UC (1998) Biodegradation of triphenylmethane dyes. Enzyme Microb Technol 22:185–191
Banat IM, Nigam P, Singh D, Marchant R (1996) Microbial decolorization of textile- dye-containing effluents: a review. Biores Technol 58:217–227
Edwards U, Rogall T, Blocker H, Emde M, Bottger EC (1989) Isolation and direct complete nucleotide determination of entire genes: characterization of a gene coding for 16S ribosomal RNA. Nucleic Acids Res 17:7843–7853
Knapp JS, Newby PS (1995) The microbiological decolorization of an industrial effluent containing a diazo-linked chromophore. Water Res 29:1807–1809
Nobuki H, Kazuko K, Kazutoshi U (2000) Isolation and characterization of Aeromonas sp. B-5 capable of decolorizing various dyes. J Biosci Bioeng 90:570–573
Padamavathy S, Sandhya S, Swaminathan K, Subrahmanyam YV, Kaul SN (2003) Comparison of decolorization of reactive azo dyes by Microorganisms isolated from various source. J Environ Sci 15:628–632
Pearce CI, Lloyd JR, Guthrie JT (2003) The removal of color from textile wastewater using whole bacterial cells: a review. Dyes Pigm 58:179–196
Rajesh KS, Uttam CB (1999) Decolorization of triphenylmethane dyes and textile and dye-stuff effluent by Kurthia sp. Enzyme Microb Technol 24:433–437
Rafii F, Fraeankalin W, Cerniglia CE (1990) Azo reductase activity of anaerobic bacteria isolated from human intestinal micro flora. Appl Environ Microbiol 56:2146–2151
Sambrook J, Fritsch EF, Maniatis T (1989) Molecular cloning: a laboratory manual, 2nd edn. Cold Spring Harbor Laboratory Press, New York
Sani RK, Banerjee UC (1999) Decolorization of triphenylmethane dyes and textile and dye-stuff effluent by Kurthia sp. Enzyme Microb Technol 24:433–437
Verma P, Madamwar D (2003) Decolourization of synthetic dyes by a newly isolated strain of Serratia marcescens. World J Microbiol Biotechnol 19:615–618
Xu M, Guo J, Cen Y, Zhong X, Cao W, Sun G (2005) Shewanella decolorationis sp. nov., a dye-decolorizing bacterium isolated from activated sludge of a waste-water treatment plant. Int J Syst Evol Microbiol 55:363–368
Yáñez MA, Catalán V, Apráiz D, Figueras MJ, Martínez AJ (2003) Phylogenetic analysis of members of the genus Aeromonas based on gyrB gene sequences. Int J Syst Evol Microbiol 53:875–883
Yatome C, Ogawa T, Koga D, Idaka E (1981) Biodegradibility of azo and triphenylmethane dyes by Pseudomanas pseudomullei 13NA. J Soc Dyers Colour 97:166–168
Yatome C, Yamada S, Ogawa T, Matsui M (1993) Degradation of Crystal Violet by Nocardia Corallina. Appl Microbiol Biotechnol 38:565–569
Zhou JZ, Bruns MA, Tiedje JM (1996) DNA recovery from soils of diverse composition. Appl Environ Microbiol 62:316–322
Acknowledgements
This work was supported by the fund of Chinese National Programs for High Technology Research and Development (No. 2003AA214040), Guangdong Provincial Natural Science Fund (No.2004A30404002), Guangdong Provincial Natural Science Fund (No. 200115017) and Guangdong Provincial Natural Science Fund (No. 032319).
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Ren, S., Guo, J., Zeng, G. et al. Decolorization of triphenylmethane, azo, and anthraquinone dyes by a newly isolated Aeromonas hydrophila strain. Appl Microbiol Biotechnol 72, 1316–1321 (2006). https://doi.org/10.1007/s00253-006-0418-2
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00253-006-0418-2