Abstract
Aminopeptidases (EC 3.4.11.) belongs to exoprotease family, which can catalyze the cleavage of peptide bond which connects the N-terminal amino acid to the penultimate residue in a protein. Aminopeptidases catalyze the process of removal of the N-terminal amino acids of target substrates by sequential cleavage of one amino acid residue at a time. Microbial aminopeptidase are of great acceptance as industrial enzymes with varying applications in food and pharma industry since these enzymes possess unique characteristics than aminopeptidases from other sources. This review describes the various applications of microbial aminopeptidases in different industrial sectors. These enzymes are widely used in food industry as a debittering agent as well as in the preparation of protein hydrolysates. In baking, brewing, and cheese making aminopeptidases are extensively used for removing the bitterness of peptides. The inhibitors of these enzymes are found great clinical applications against various diseases such as cancer, diabetes, and viral infections. Aminopeptidases are widely used for the synthesis of biopeptides and amino acids, and found to be efficient than chemical synthesis. These enzymes are capable of hydrolyzing organophosphate compounds, thus having biological as well as environmental significance.
Key Points • Cleaves the amino-terminal amino acid residues from proteins and peptides. • Microbial aminopeptidase are of great acceptance as both therapeutic and industrial enzyme. • Review describes the potential applications of microbial aminopeptidases. |
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Microbial aminopeptidases—an overview
Enzymes that catalyze the hydrolysis of peptide bonds in protein are referred to as proteases or peptidases. The peptide bonds present in between amino acids of proteins are cleaved by specific proteases. Aminopeptidases (APs; EC 3.4.11) are a class of proteases that catalyze the cleavage of the amino-terminal amino acid residues from proteins and peptides. These are exopeptidases and are widely distributed in prokaryotic and eukaryotic organisms, found in subcellular organelles, in the cytoplasm, and as membrane components (Jankiewicz and Bielawski 2003). Aminopeptidases form a large enzyme family in microbial world (Gonzales and Robert-Baudouy 1996) and are important in many biological functions such as post translational modification of proteins, its breakdown and maturation; thus, they perform both regulatory and housekeeping functions (Chandu and Nandi 2003; Taylor 1993a)). Aminopeptidases are predominantly present in cells where they have important roles in the processing of newly synthesized proteins, breakdown of peptide hormones, and the processing of various enzymes. Microbial aminopeptidases have major roles in the utilization of external protein substrates present in the medium as a source of essential amino acids (Gonzales and Robert-Baudouy 1996). Removal of the amino-terminal methionine by methionine aminopeptidase during protein translation is considered as a typical aminopeptidase activity occurring in the cells (Bradshaw et al. 1998; Li and Chang 1995).
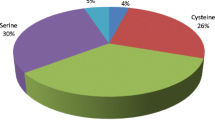
Aminopeptidases are classified into three major distinct classes based on the number of amino acids cleaved from polypeptide chains, substrate specificity, and sensitivity to various protease inhibitors (Fig. 1). Aminopeptidases which hydrolyze the first peptide bond in a polypeptide chain releasing a single amino acid residue are called aminoacylpeptidases (EC 3.4.11); some aminopeptidases remove dipeptides or tripeptides from polypeptide chains are named as dipeptidyl- and tripeptidyl peptidases (EC 3.4.14). Another class of enzymes which only act on di- or tripeptides are called dipeptidases (EC 3.4.15) and tripeptidases (EC 3.4.14.4). A novel prolyl tri/tetra-peptidyl aminopeptidase from Streptomyces mobaraensis that removes the tetra-peptide of pro-transglutaminase was reported (Umezawa et al. 2004). Aminopeptidases are further classified into two categories based on substrate specificity such as broad and narrow. Differences in the catalytic site of enzymes and enzyme binding pockets of substrates are majorly responsible for the substrate specificity of aminopeptidases, thus there are broad or narrow substrate specific aminopeptidases are present in microorganisms (Lowther and Matthews 2002; Holz et al. 2003). Based on the N-terminal amino acid specificities of aminopeptidases at the substrate site there are substrate specficities existing for prolyl aminopeptidase, X-prolyl dipeptidyl aminopeptidase, alanine aminopeptidase, methionyl aminopeptidase, arginine aminopeptidase, lysine aminopeptidase, leucine aminopeptidase, phenylalanine aminopeptidase, and so on. Based on catalytic mechanism and sensitivity to various protease inhibitors aminopeptidases are classified into metallo, cysteine, and serine peptidases. Metallo aminopeptidase are the largest group of aminopeptidases and are inhibited by metal-chelating agents such as EDTA, EGTA, and 1,10-phenanthroline. Cysteine aminopeptidase are inhibited by Hg2+, iodoacetamide, N-ethylmaleimide, and p-chloromercuribenzoate. Serine aminopeptidase are sensitive to phenyl methyl-sulfonyl fluoride and diisopropyl flurophosphates. There are no ionic co-factors seen associated with cysteine and serine aminopeptidases, and its catalysis requires a highly reactive cysteine or serine residue. Among organisms, this class of aminopeptidases is found less diversified than metalloaminopeptidases. This includes cysteine peptidases of relatively broad specificity such as PepC and bleomycin hydrolase, and narrow specificity serine peptidases such as proline-specific peptidases (prolyl aminopeptidase or PepI, PepX, and dipeptidyl peptidase IV) (Gonzales and Robert-Baudouy 1996).
It has been observed that various cellular compartments of microbial cells possess aminopeptidase activity. A large fraction of these enzymes are found in soluble forms in the cytoplasm or seen associated with in the cell wall or secreted into the exterior environment (Goldberg et al. 1997). A signal sequence which is a characteristic of exported proteins is present at the N-terminal end of an extracellular leucine aminoptidase of A. proteolytica has been identified (Guenet et al. 1992). The microorganisms which are reported to be producing substrate specific aminopeptidase are mainly belong to genus Aspergillus (Matsushita-Morita et al. 2010), Streptomyces (Rahulan et al. 2009; Nandan et al. 2011; Wu et al. 2010), Pseudomonas (Wu et al. 2014), lactic acid bacteria (Tchorbanov et al. 2011), and Bacillus (Shen et al. 2011). Different classes of substrate specific extracellular aminopeptidases are produced from Streptomyces griseus (Hershcovitz et al. 2004), Streptomyces gedenensis (Rahulan et al. 2009), Streptomyces lavendulae (Nandan et al. 2011), Aeromonas proteolytica (Holz 2002), the filamentous fungi Aspergillus oryzae (Huang et al. 2015), and Aspergillus sojae (Chien et al. 2002). Cytoplasmic and soluble aminopeptidases are reported from various Lactobacillus strains (Nandan et al. 2010; Vesanto et al. 1994; Klein et al. 1994). Aminopeptidases from lactic acid bacteria are of industrially important since they are widely used in food industry (Pan and Tanokura 2004; Choi et al. 1996).
Current research on aminopeptidases mainly focused on gene cloning and expression, protein purification and characterization, catalytic mechanisms, and in silico evaluations (Arima et al. 2004; Arima et al. 2006a; Sonoda et al. 2009; Nandan and Nampoothiri 2014; Arif et al. 2018; Labrie et al. 2019). The gene sequences of aminopeptidases and its biochemical functions are two determining factors of the substrate specificity of aminopeptidases (Rawlings et al. 2004). Numerous substrate-based library screening methods have been developed for the fast and unfailing determination of specificity of enzymes (Backes et al. 2000; Harris et al. 2000; Choe et al. 2006). A fast and reliable evaluation of the substrate specificity of individual aminopeptidase was developed by Drag et al. (2008) using solid phase chemistry with the substrate 7-amino 4-carbamoylmethylcoumarin fluorophore. Aminopeptidases are still an ongoing topic of research with its role already connected in explaining various vital processes such as protein processing and turnover, tissue invasion, regulation of peptide hormone synthesis, viral infections, and plant defense responses with reasonable confidence. Various attempts have been undertaken to study and determine the substrate specificity of aminopeptidases. One such attempt uses sequence similarity, which has its primary deficiency due to the lack of availability of sequence signatures (Petrovic et al. 2007). The mechanism of catalysis in aminopeptidase needs to be carefully studied for the successful utilization of the enzymes in the industry. Innovative technologies using recombinant DNA and site-directed mutagenesis have been raised for the development of stable aminopeptidases with improved substrate specificity (Nandan and Nampoothiri 2017a).
Applications of microbial aminopeptidases
Aminopeptidases have considerable applications in various fields because of their wide range substrate specificity, inflexible enantioselectivity, and high thermal stability (Arima et al. 2006b). As illustrated in Fig. 2, food industry is the primary sector which recognized and utilized the substrate-specific microbial aminopeptidases.
Therapeutic application
Aminopeptidases play important roles in diverse cellular processes such as protein modification, protein degradation, cell-cycle control, and hormone level regulation. Therefore, these enzymes play a significant role in many pathophysiological conditions from infections to cancer (Taylor 1993a; Taylor 1993b). Microorganisms are the major sources of high yield production of medically important aminopeptidases with economic feasibility. Pharmaceutical applications of aminopeptidases are directed to control the pathophysiological effects, in a way helps in the development of diagnostic tools such as biomarkers of these physiological pathways. Microbial aminopeptidases are potential targets for structure based drug design since these enzymes have high economic feasibility, high yields, regular availability, ease of modification, and high catalytic efficiency. Microbial aminopeptidases and their human counterparts share structural and functional similarity. The studies on the mechanism of co-catalytic active site of aminopeptidases are helpful in designing the new inhibitors of aminopeptidases acting as pharmaceuticals (Table 1) (Holz 2002; Lowther and Matthews 2002).
Bacterial methionine aminopeptidases have been implicated as potential targets for developing broad spectrum antibacterial drugs (Helgren et al. 2016; Schiffmann et al. 2006). Inhibitors of methionine aminopeptidase are having therapeutic function since they are reported to have regulatory roles in angiogenesis and tumor progression (Selvakumar et al. 2005; Zhong and Bowen 2006). The inhibitors of glutamyl aminopeptidase (aminopeptidase A) act as potential antihypertensive compounds, thus mediating the conversion of angiotensin II into angiotensin III, which ultimately regulating the arterial blood pressure (Cogolludo et al. 2005; Inguimbert et al. 2005). Lactobacillus delbrueckii is an efficient producer of aspartyl/glutamyl aminopeptidase (Stressler et al. 2016). Inhibitors of alanyl aminopeptidase (aminopeptidase N) are being used in the treatment and regulation of inflammatory diseases, cancer, leukemia, diabetic nephropathy, and rheumatoid arthritis (Bauvois and Dauzonne 2006; Ansorge et al. 2006). Aminopeptidase N acts as a membrane receptor for many types of corona viruses in human host (Fehr and Perlman 2015). The inhibitors of these membrane-bound aminopeptidases can function as antiviral agents, and thus can be considered as a treatment strategy for the viral diseases. It has been reported that E. coli is a potent microbial producer of aminopeptidase N, and its crystal structure revealed that methionine 260 serves as a substrate recognition residue (Ito et al. 2006). The design of inhibitors of dipeptidyl peptidase IV (DPP IV) and related proline-specific aminopeptidases are used in the treatment of type 2 diabetes and immunological disorders (Augustyns et al. 2005; Mest 2006). The inhibition of X-prolyl dipeptidyl aminopeptidase (PepX) is helping in treating against infection by Streprococcus gordonii (Goldstein et al. 2001). Rigolet et al. (2005) conducted a comparative study between the structures of Lactococcus lactis PepX and its human counterpart. They have identified the key residues and projected them as drug targets since PepX is involved in many bacterial infections. Dipeptidyl peptidase and aminopeptidases from periodontopathic rods inhabiting human oral microflora are useful diagnostic tool by acting as markers of periodontopathic bacteria (Nemoto et al. 2018). An antibiotic of microbial origin called bestatin act a potent inhibitor of most of the aminopeptidases including alanine aminopeptidase, cystinyl aminopeptidase, aminopeptidase B, and aminopeptidase N (Scornik and Botbol 2001). Apstatin, an inhibitor of a membrane-bound aminopeptidase P function as a vasodilator by potentiating the effect of bradykinin. Bradykinin is a neuropeptide having role in the blood vessel dilation and is cleaved by aminopeptidase P (Maggiora et al. 1999). Bacterial aminopeptidases are also perfect enzymes for structure–function analysis, since they can efficiently produce and purified as recombinant proteins. A novel immune regulatory function for Streptococcus pneumonia aminopeptidase in generating CD8+T cell population with reduced effector function gave rise to the possibility of direct regulation by pneumococcal components. This study elucidates a novel mechanism by which these enzymes can directly modulate host T cell effector function and may play a vital role in pneumococcal disease (Blevins et al. 2017).
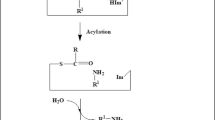
Bioactive peptide synthesis
Enzymatic peptide synthesis presents a useful and helpful strategy because it can perform specified reactions under milder conditions than those used in chemical synthesis. The peptide synthesis applications of microbial aminopeptidases are already been reported in literature (Table 2). From the view point of biotechnology, proline aminopeptidases might be a perfect tool for synthesizing peptides containing proline by catalyzing aminolysis reaction. The application of proline aminopeptidase (PAP) from Streptomyces thermoluteus in synthesizing proline containing peptides was demonstrated using variants of PAP (Yamamoto et al. 2010). Proline containing peptides are reported to be having nutraceutical properties. Studies on prolyl hydroxyproline (Pro-Hyp) proved that they can stimulate the growth of fibroblasts from mouse skin (Shigemura et al. 2009). The peptide Pro-Arg is well-known for its defensive activity against hydrogen peroxide induced cell death and other oxidative stresses. In another study, the milk peptides Val-Pro-Pro and Ile-Pro-Pro showed activity against the transmembrane aminopeptidase angiotensin-converting enzyme (ACE) (Meyer et al. 2009). Proline rich diketopiperazine such as cyclo (Pro-Pro) showed antibacterial activity against Micrococcus luteus and Pseudomonas aeruginosa (Huberman et al. 2007). An executioner caspase-3 was activated by cyclo (Pro-Phe) in HT-29 cells was studied (Brauns et al. 2005). Whey protein hydrolysate treated with exopeptidases showed ACE inhibitory activity, thus reducing systolic pressure in spontaneously hypertensive rats (Cheung et al. 2015). ACE inhibitory peptide generation from skimmed milk hydrolysate using aminopeptidase was explained by Pan et al. (2005). Since majority of bioactive peptides are produced by exopeptidases such as aminopeptidases, the impact of these enzymes in determining the properties of these bioactives are of much concern. The bioactive peptide profiles of three European dry fermented sausages were analyzed and explained the role of microbial peptidases in producing bioactive peptides (Gallego et al. 2018). Many intracellular peptidases have been reported from the starter cultures such as L. sakei, L. helveticus, L. delbrueckii, L. curvatus, L. plantarum, L. brevis, L. casei, and L. paracasei. These cultures have been reported to exert high exopeptidase activity by producing enzymes such as dipeptidase, tripeptidase, aminopeptidase, arginine aminopeptidase, PepA, PepX, PepP, and X-prolyl dipeptidyl peptidase etc, thus showed an increased amount of bioactive peptides in the medium (Toldra et al. 2018).
Degradation of organophosphorus compounds
Organophosphorus poisoning is considered as a major health problem across the world. These compounds are known for their toxicological effects such as the inactivation of the enzyme acetylcholinesterase and the subsequent loss of nerve function. Thus, the safety and protection from the detoxification process of these compounds is of much concern to the general public. Scientists are looking for an efficient ecofriendly biodegradation method for the efficient removal of these compounds from the environment. Various microbial sources are explored for the biological degradation of organophosphate pesticides (Sidhu et al. 2019). Organophosphorous acid anhydrolase isolated from Alteromonas undina was purified which can detoxify organophosphorous compounds has been reported (Cheng et al. 1993). Aminopeptidase P was found to catalyze the hydrolysis of a wide range of organophosphate triesters. The activity of aminopeptidase P was promoted in the presence of Mn2+. Mutant proteins of aminopeptidase P was made and tested for the hydrolytic activity against organophosphorous compounds. The study concluded the structural details of aminopeptidase P to facilitate the hydrolysis of organophosphate tri esters (Jao et al. 2004). A proline-specific aminopeptidase P was found to be sharing 31% sequence similarity with the acid anhydrolase capable of degrading organophosphorus compounds was identified from E. coli (Singh and Walker 2006). The catalytic activities of E. coli aminopeptidase P showed substrate preference towards organophosphorus compounds and methylphosphonate derivatives (Fig. 3) (Hsu et al. 2008).
Food processing applications
Protein hydrolysate and debittering
Food protein hydrolysate preparation is considered as one of the leading industrial applications of aminopeptidases. Protein hydrolysates derived from milk, soy, meat, and cereals are essentially prepared by the combined action of endo and exopeptidases (Fig. 4) (Meyer-Barton et al. 1994; Chevalet et al. 2001; Scharf et al. 2006). These food protein hydrolysates are widely used for the generation of bioactive peptides with nutritional and pharmacological importance. These hydrolysates are the rich source of pre-digested ingredients which can be easily utilized during intestinal absorption (FitzGerald and O’Cuinn 2006). As previously described, the substrate specificity of aminopeptidases contributes to the hydrolysis efficiency which determines the properties of the hydrolysates which is being prepared. The substrate specificity of an aminopeptidase from Bacillus licheniformis was tested by checking its efficiency in hydrolyzing proteins such as peanut protein isolate and zein both with different percentages of leucine content. Peptide profiling results suggested that the enzyme is a leucine specific aminopeptidase (Lei et al. 2018). A comparative study was conducted to estimate the hydrolysis efficiency of leucine aminopeptidase from Streptomyces gedenensis and other two commercially available enzymes such as pepsin and trypsin. This result reflects the substrate specificity of S. gedanensis aminopeptidase to hydrolyse peptides with amino-terminal hydrophobic and aromatic residues is more when compared with commercial enzymes (Rahulan et al. 2012). Moreover, the studies confirmed that the bioactivity of peptide hydrolysates is determined by the amount of free fatty acids (FAAs) present (Rahulan et al. 2012). Researchers emphasized that bitterness is a major limitation in food industry which limits the consumption and utilization of food protein hydrolysates. Aminopeptidases are capable of reducing the bitterness by increasing the degree of hydrolysis thus releasing free amino acids (Giesler et al. 2013). The debittering property of aminopeptidases was demonstrated in various studies. The degree of hydrolysis and free fatty acid contents were increased when treated with recombinant aminopeptidase (alcalase) from A. sojae GIM3.30. Thus, it reduced the bitterness of casein and soy protein hydrolysates (Huang et al. 2015). An aminopeptidase from B. licheniformis improves the hydrolysis and debittering efficiency of soy protein isolate (Lei et al. 2017). Bacterial strains such as Bacillus spp., Streptomyces spp., Aeromonas spp., and various fungal strains are producers of leucine aminopeptidases whose debittering activity was extensively studied (Stressler et al. 2013; Nampoothiri et al. 2005; Lin et al. 2004). Furthermore, aminopeptidases from A. oryzae and A. sojae are regarded as food safe enzymes and have a long history of use as debittering enzymes. Hydrolysis with A. sojae recombinant leucine aminopeptidase increased the degree of hydrolysis thereby decreased the bitterness of casein and soy protein hydrolysates (Huang et al. 2015). Extracellular proteome analysis of A. oryzae during soy sauce fermentation using iTRAQ method resulted in the identification of dipeptidase, dipeptidyl aminopeptidase, and leucine aminopeptidase. These enzymes have important implications for soy sauce fermentation (Zhao et al. 2018). The debittering effect of Aeromonas cavieT-64 is reflected in the hydrolysis of milk casein and soy protein. The bitterness of the solutions were remarkably reduced by the release of free amino acids (Izawa et al. 1997). The release of free amino acids such as tyrosine, phenyl alanine, leucine, isoleucine, and valine had reduced the bitterness of milk protein and soy protein hydrolysate when treated with aminopeptidases from the fungal strain Grifola frondosa (Nishiwaki et al. 2002). Aspartyl aminopeptidases have specificity towards N-terminal aspartic and glutamic acids had been isolated from A. oryzae, yeast, E.coli, and L. delbrueckii. Food proteins rich with the glutamyl or aspartyl group when treated with aspartyl aminopeptidases from A. oryzae leads to the production of flavored hydrolysates by the release of free aromatic amino acids (Kusumoto et al. 2008; Yokoyama et al. 2006; Watanabe et al. 2007). Salt tolerant aspartyl aminopeptidase was isolated from A. oryzae (Gao et al. 2018). Aspartyl aminopeptidase can also contribute to the umami taste production of hydrolysate from wheat gluten, casein, and fermented soy products (Stressler et al. 2016).
Cured meat products
Dry-cured meat products are prepared by various processes involving drying and ripening of meat. Aminopeptidase appears to be responsible for the increase in free amino acids during dry-curing. In an analysis of proteases in the fresh pork muscle during ripening, the release of free amino acids from bitter peptides were found to be greater in number (Virgili et al. 1998). The overall process of meat protein processing involved the use of both endopeptidases and exopeptidases. During processing, meat proteins are extensively hydrolyzed by muscle endopeptidases followed by exopeptidases. Endopeptidases such as calpains and cathepsins hydrolyze the proteins into smaller peptides and oligopeptides. Exopeptidases such as aminopeptidases and carboxypeptidases are responsible for the release of free amino acids capable of contributing flavor in dry-cured products (Mora et al. 2018). Microbial starter cultures such as lactic acid bacteria, Staphylococci, yeasts are rich source of several peptidases like aminopeptidases, dipeptidases, and tripeptidases (Bintsis et al. 2003; Macedo et al. 2003; Bolumar et al. 2003). The prolyl aminopeptidase, arginyl aminopeptidase, and aminopeptidase I, II, and IV of cell-free extracts from L. sakei were proved to enhance the sensory qualities of dry-fermented sausages (Bolumar et al. 2006). Leuconostoc mesenteroides and L. curvatus strains have been reported to show elevated activity of X-prolyl dipeptidyl aminopeptidase (Zotta et al. 2007). Most of the exopeptidases are reported to be involved in the release of small peptides, free amino acids, and some bioactive peptides. These free amino acids are contributing to the development of aroma. The proteomic profile of small peptides in dry-cured meat resulted in the identification of amino acids such as alanine, lysine, serine, tyrosine, arginine, and valine released as a result of combined action of microbial and muscle aminopeptidases (Mora et al. 2015). Free amino acids are non-volatile compounds that contribute to the improvement of both meat taste and flavor. Micrococcus roseus produced two intracellular aminopeptidases with affinity for nonpolar amino acids, L-Pro, and L-Arg. M. roseus aminopeptidase appears to release amino acids and thereby contribute to flavor development in cured bacon (Hinrichsen et al. 1994). Lactic acid bacteria proteases have also been studied in sausage products. Three L. plantarum strains were screened for endopeptidase and aminopeptidase activities to evaluate them as starters in sausage. Whole cells of L. plantarum CRL 681 generated hydrophilic peptides while whole cells when used with cell-free extract produced hydrophobic peptides from both sarcoplasmic and myofibrillar proteins. L. curvatus, L. sake, and L. casei are the most common microorganisms in dry-fermented sausages, and their use as starter cultures is also widespread (Sanz et al. 1999). Fadda et al. (1999) had studied the aminopeptidase activities of L. curvatus and L. sake in dry-fermented sausages. The activity of L. curvatus resulted mainly in the generation of large amounts of glutamic acid, b-alanine, arginine, histidine, and lysine while the activity of L. sake increased the levels of glutamic acid, b-alanine, g-aminobutyric acid, alanine, threonine, phenylalanine, leucine, and ornithine. In the preparation of sausages, microbial aminopeptidases such as aminopeptidase 1 and 2 from L. sake can contribute to the release of free amino acids. Sodium chloride was used as a curing agent which could activate arginine aminopeptidase and aminopeptidase 2 from L. sake (Flores et al. 1998).
Ripened cheeses
The conversion of fresh cheese curd into mature cheese is largely determined by the progress of proteolysis. For the development of an admissible cheese flavor, a well-balanced breakdown of milk protein casein into small peptides and amino acids is necessary. Casein is rich with proline residues, the peptide bond-containing proline are resistant towards most of the proteases; thus, the peptidases specific for proline has a crucial role in the degradation of casein proteins. A great variety of proline-specific aminopeptidases such as proline iminopeptidase, aminopeptidase P, and prolyl dipeptidase have been found in starter organisms commonly used in cheese manufacture. Caseins are also rich in glutamine residues which are hydrolyzed by microbial aminopeptidase A (Visser 1993). The bitterness in cheese is contributed by the medium-sized peptides resulting from the enzymatic digestion of casein. Various studies observed that the most important role in bitterness is played by the starter cultures (Lemieux and Simard 1991). During cheese ripening aminopeptidases from the thermophilic lactic acid bacteria remains to be active throughout the process (Gatti et al. 1999). Prolyl aminopeptidases from Penicillium camemberti plays an important role in the ripening of Camembert-type cheese (Fuke and Matsuoka 1993). The activities of PepX and lysine aminopeptidase of Lactococcus lactis were found to be increased during the ripening of Saint Paulin cheese (Chapot-Chartier et al. 1994). El-Kholy et al. (1998) studied the strong aminopeptidase and dipeptidyl aminopeptidase activities of thermophilic lactobacilli in Ras-type cheese. Lactic acid bacterial proteolytic enzymes are particularly important in the formation of flavors during cheese ripening (Visser 1993). The meta-omics analysis data of surface ripened cheese community revealed the differential expression of proline-specific aminopeptidase gene from Hafnia alvei and cysteine aminopeptidase gene from Geotrichum candidum (Dugat-Bony et al. 2015). The optimum time for ripening of a traditional Brazilian cheese named as artisanal Minas cheese was studied and the data indicated that higher temperatures accelerate the process of cheese ripening due to the presence of high amount of aminopeptidases (Martins et al. 2015).
Fermented fish products
Fish fermentation is the conversion of organic substances into simple compounds such as peptides, amino acids, and other nitrogenous compounds either by the action of microorganisms or their endogenously producing enzymes. Fermented fish products are very popular particularly in Southeast Asian countries contributing significantly to the protein intake of population. These products have distinctive characteristics, mostly in terms of aroma, and texture during fermentation process. A bacterial dynamic study had isolated a total of 210 bacterial species from a dry fermented fish product Ngari. The dominant bacteria inhabited were Staphylococcus carnosus, L. pobuzihii, Bacillus indicus, Enterococcus faecium, and Tetragenococcus halophilus (Devi et al. 2015). Fish fermentation is a natural process mainly depends both on naturally occurring bacteria (in the muscle or the intestinal tract) as well as its enzymes. The activities of three endogenous aminopeptidases such as arginine aminopeptidase, alanyl aminopeptidase, and leucyl aminopeptidases were reported in the processing of dry-salted fish. The levels were significantly increased during the final stage of fish processing (Wu and Cao 2018).There are many studies on the quality improvement of fish sauce by optimizing the starter inocula strains. Zheng et al. (2017) have found that inoculation of Psychrobacter sp. SP-1 during meat processing significantly increased the protease activity and also promoted the umami taste and meaty aroma. Fish sauce is a prime seasoning for Asian cuisines and contains salt concentrations around 25% to 30%. Therefore, studies are focusing on designing of halophilic starter cultures for preparing quality fish sauce. A novel Staphylococcus sp. has been isolated from fermented fish sauce and suggested to be applied as a starter culture to increase the free amino acid content, and thus improves the umami and aroma of the fish sauce (Udomsil et al. 2015).The recovery of flavor molecules from fermented fish products mostly relies on the use of commercial protease preparations such as flavourzyme which is a leucyl aminopeptidase and protamex (Suresh and Prabhu 2013). These commercial enzymes are widely used during the processing of various fermented fish products which helps in quality improvement and process acceleration (Giyatmi and Irianto 2017). The endogenous aminopeptidase activity in fish sauce was studied by Vo-Van et al. (1984). These studies described the identification and purification of the aminopeptidase appearing during sardine fish sauce fermentation and changes of this enzyme activity during the entire process. Thus, aminopeptidases plays an important role in contributing to free amino acids in fish sauce.
Cocoa processing
Cocoa is cultivated in tropical regions around the world, and its fruit is the main component in chocolate production. Cocoa has astringent bitterness by the content of tannin and polyphenols. An important phase of cocoa processing is fermentation that reduces bitterness and fermentation. During the fermentation process, the pulp of fruit is degraded by the action of various microbes such as lactic acid bacteria, acetic-acid bacteria, and yeasts naturally occurring in the environment. Enzymatic reactions play important roles in protein hydrolysis in cocoa almonds to produce flavoring precursor compounds. Proteolysis is very important for cocoa flavor development (Voigt et al. 1994a, b).Therefore, in the production of a standardized chocolate mass, the addition of enzyme can be used in the treatment of cocoa (Gray 2011). The study by Oliveira et al. (2011) showed that the better quality of the chocolate were produced by the action of protease and carboxypeptidase (flavor protease) used in the processing of cocoa almonds. Merz et al. (2015) identified the eight enzymes in the commercial preparation of Flavourzyme from Aspergillus oryzae. Flavourzymes is a widely used enzyme cocktail in cocoa fermentation. Among the eight enzymes, two were reported as leucine aminopeptidase A and leucine aminopeptidase 2. The protease activity in the pulp and seed of cocoa during the fermentation of two cocoa cultivars were compared by Sousa et al. 2016. The result showed same isoenzyme behavior for both the cultivars with regard to the aminopeptidase and carboxypeptidase. The use of a proteolytic enzyme with an activity of 1000 leucine aminopeptidase units (LAPU) g−1 provided by Novo Nordisk A/S (Bagsvaerd, Denmark) was useful in improving cocoa flavor precursors and affected the flavor perception in their products (De Brito et al. 2004). Among cocoa proteins, proline content (0.72–1.97 g/100 g of cocoa) showed maximum, and because of its specific structure, it possesses many limitations on the conformational aspects of peptides and proteins, and thus proline-specific peptidases takes the role for hydrolyzing such proteins (Kratzer et al. 2009). It is stated that both endogenous enzymes present in cocoa seeds as well as exogenous enzyme derived from microorganisms have importance in the processing of cocoa and thus developing cocoa flavor precursors.
Commercial aminopeptidases and market
The global enzymes market value was $7082 million in 2017, and is predicted to reach $10,519 million in 2024, registering a compound annual growth rate (CAGR) of 5.7% from 2018 to 2024. The crucial factors driving the enzyme market are the growing array in enzyme applications, its products and strict environmental norms suppressing the use of chemicals (Research and Markets 2019). Proteases are the most valuable commercial enzyme covering 60% of total enzyme market. The flavourzyme (Novo Nordisk), debitrase (Imperial Biotechnology Ltd), corolase (Rohm GmbH), and pronase (Calbiochem) are some of the various trade names of industrially available aminopeptidases (Nandan and Nampoothiri 2017b). Microorganisms are preferred sources of industrial enzymes as they are economic, effective, and have a controllable enzyme reaction mechanism. Table 3 shows some of the patents claims on aminopeptidases either to enhance the enzymatic activity or to improve various industrial applications. Most of the accepted patents on aminopeptidases are from Aspergillus stain as shown in table. Patented enzymes are widely used for various industrial applications. The global market size of industrial enzymes has been segmented on the basis of type, source, application of enzymes and its sources. According to the report published by Markets and Markets (2019), the top level manufacturers in the food enzymes market include Associated British Foods (UK), DowDuPont (US), Novozymes (Denmark), DSM (Netherlands), and Chr. Hansen (Denmark). These companies have a wide range of product portfolios and advanced technologies for food enzymes at major strategic locations. The increasing interest in using microbial enzymes especially proteases for the production of various industrially important products in the market is of huge importance. In order to find a cost-effective way, scientists are in search of suitable source for enzyme production. Microbial proteases are and will very likely remain the most utilized enzymes at both academic and applied levels.
Conclusion
This review focuses on the application of microbial aminopeptidases in different sectors. Microbial aminopeptidases provide a greater amount of catalysis with a wide range of applications across many industries such as food and pharma. Biocatalysis and structural studies of these enzymes will provide a greater platform for better screening of novel inhibitors. An aminopeptidase-mediated bioactive peptide synthesis is more preferred because the process is ecofriendly and economic. The substrate specific aminopeptidases from various strains of microorganisms and their applications are yet to be studied. Various molecular and biochemical approaches are being carried out to improve the characteristics of the substrate specific aminopeptidases. Aminopeptidase profiles of several bacterial and fungal strains that are widely used as starter cultures are yet to elaborate and find application in novel bioprocesses in food processing. The substrate specificity of microbial aminopeptidase has to be exploited further for their commercial interest and industrial applications.
References
Ansorge S, Bank U, Nordhoff K, Taeger M, Striggow F (2006) Dual alanyl aminopeptidase and dipeptidyl peptidase IV inhibitors for functionally influencing different cells and for treating immunological, inflammatory, neuronal and other diseases. Patent WO2005/034940, 21 April 2005
Arif A, Mohammed K, Nadeem MS (2018) Biochemical and in silico evaluation of recombinant E. coli aminopeptidase and in vitro processed human interferon α-2b. Turk J Biol 42(3):240–249. https://doi.org/10.3906/biy-1801-83
Arima J, Iwabuchi M, Hatanaka T (2004) Gene cloning and overproduction of an aminopeptidase from Streptomyces septatus TH-2 and comparison with a calcium activated enzyme from Streptomyces griseus. Biochem Biophys Res Commun 317(2):531–538. https://doi.org/10.1016/j.bbrc.2004.03.082
Arima J, Uesugi Y, Iwabuchi M, Hatanaka T (2006a) Study on peptide hydrolysis by aminopeptidases from Streptomyces griseus, Streptomyces septatus and Aeromonas proteolytica. Appl Microbiol Biot 70(5):541–547. https://doi.org/10.1007/s00253-005-0105-8
Arima J, Uesugi Y, Uraji M, Yatsushiro S, Tsuboi S, Iwabuchi M (2006b) Modulation of Streptomyces leucine aminopeptidase by calcium: identification and functional analysis of key residues in activation and stabilization by calcium. J Biol Chem 281(9):5885–5894. https://doi.org/10.1074/jbc.M509025200
Augustyns K, Van der Veken P, Senten K, Haemers A (2005) The therapeutic potential of inhibitors of dipeptidyl peptidase IV (DPP IV) and related proline-specific dipeptidyl aminopeptidases. Curr Med Chem 12(8):971–998. https://doi.org/10.2174/0929867053507298
Backes BJ, Harris JL, Leonetti F, Craik CS, Ellman JA (2000) Synthesis of positional –scanning libraries of fluorogenic peptide substrate libraries to define the extended substrate specificity of plasmin and thrombin. Nat Biotechnol 18:187–193. https://doi.org/10.1038/72642
Bauvois B, Dauzonne D (2006) Aminopeptidase-N/CD13 (EC 3.4.11.2) inhibitors: chemistry, biological evaluations, and therapeutic prospects. Med Res Rev 26(1):88–130. https://doi.org/10.1002/med.20044
Ben-Bassat A, Bauer KA, Chang S, Chang SY (1991) Bacterial methionine N-terminal peptidase, US Patent 5,013,662, 7 May 1991
Bintsis T, Vafopoulou-Mastrojiannaki A, Litopoulou-Tzanetaki E, Robinson RK (2003) Protease, peptidase and esterase activities by lactobacilli and yeast isolates from Feta cheese brine. J Appl Microbiol 95(1):68–77. https://doi.org/10.1046/j.1365-2672.2003.01980.x
Blevins LK, Parsonage D, Oliver MB, Domzalski E, Swords WE, Alexander-Miller MA (2017) A novel function for the Streptococcus pneumonia aminopepptidase N: inhibition of T cell effector function through regulation of TCR signaling. Front Immunol 8:1–17. https://doi.org/10.3389/fimmu.2017.01610
Bolumar T, Sanz Y, Aristoy MC, Toldra F (2003) Purification and characterization of a prolyl aminopeptidase from Debaryomyces hansenii. Appl Environ Microbiol 69(1):227-232. https://doi.org/10.1128/aem.69.1.227-232.2003
Bolumar T, Sanz Y, Flores M, Aristoy MC, Toldra F, Flores J (2006) Sensory improvement of dry-fermented sausages by the addition of cell-free extracts from Debaryomyces hansenii and Lactobacillus sakei. Meat Sci 72(3):457–466. https://doi.org/10.1016/j.meatsci.2005.08.010
Bradshaw RA, Brickey WW, Walker KW (1998) N-terminal processing: the methionine aminopeptidase and N alpha-acetyl transferase families. Trends Biochem Sci 23(7):263–267. https://doi.org/10.1016/s0968-0004(98)01227-4
Brauns SC, Dealtry G, Milne P, Naude R, de Venter VM (2005) Caspase-3 activation and induction of PARP cleavage by cyclic dipeptide cyclo (Phe-Pro) in HT-29 cells. Anticancer Res 25(6B):4197–4202
Chandu D, Nandi D (2003) PepN is the major aminopeptidase in Escherichia coli: insights on substrate specificity and role during sodium–salicylate induced stress. Microbiology 149(12):3437–3447. https://doi.org/10.1099/mic.0.26518-0
Chapot-Chartier MP, Deniel C, Rousseau M, Vassal L, Gripon JC (1994) Autolysis of two strains of Lactococcus lactis during cheese ripening. Int Dairy J 4(3):251–269. https://doi.org/10.1016/0958-6946(94)90016-7
Cheng TC, Harvey SP, Stroup AN (1993) Purification and properties of a highly active organophosphorus acid anhydrolase from Alteromonas undina. Appl Environ Microbiol 59(9):3138–3140
Cheung LKY, Aluko R, Cliff MA, Li-Chan ECY (2015) Effects of exdopeptidase treatment on antihypertensive activity and taste attributes of enzymatic whey protein hydrolysates. J Funct Foods 13:262–275. https://doi.org/10.1016/j.jff.2014.12.036
Chevalet L, Souppe J, De Leseleuc J, Burnet J, Warmerdam MJ (2001) Aspergillus niger aminopeptidase compositions for making bread doughs and cheese. US Patent 6,271,013, 7 August 2001
Chien HCR, Lin LL, Chao SH, Chen CC, Wang WC, Shaw CY, Tsai YC, Hu HY, Hsu WH (2002) Purification, characterization, and genetic analysis of a leucine aminopeptidase from Aspergillus sojae. Biochim Biophys Acta 1576(1–2):119–126. https://doi.org/10.1016/s0167-4781(02)00307-x
Choe Y, Leonetti F, Greenbaum DC, Lecaille F, Bogyo M, Bromme D, Ellman JA, Craik CS (2006) Substrate profiling of cysteine proteases using a combinatorial peptide library identifies functionally unique specificities. J Biol Chem 281(18):12824–12832. https://doi.org/10.1074/jbc.M513331200
Choi H, Laleye L, Amamtea GF, Simard RE (1996) Production of Aminopeptidase from skim milk whey permeate medium by Lactobacillus casei ssp. casei. J Dairy Sci 79(6):956–963. https://doi.org/10.3168/jds.S0022-0302(96)76446-9
Cogolludo A, Perez-Vizcaino F, Tamargo J (2005) New insights in the pharmacological therapy of arterial hypertension. Curr Opin Nephrol Hypertens 14(5):423–427. https://doi.org/10.1097/01.mnh.0000168334.09454.1c
De Brito ES, García NHP, Amancio AC (2004) Use of a proteolytic enzyme in cocoa (Theobroma cacao L.) processing. Braz Arch Biol Technol 47(4):553–558. https://doi.org/10.1590/S1516-89132004000400008
Devi KR, Deka M, Jeyaram K (2015) Bacterial dynamics during year-long spontaneous fermentation for production of Ngari, a dry fermented fish product of Northeast India. Int J Food Microbiol 199:62–71. https://doi.org/10.1016/j.ijfoodmicro.2015.01.004
Drag M, Mikolajczyk J, Bekes M, Reyes-Turcu FE, Ellman JA, Wilkinson KD, Salvesen GS (2008) Positional –scanning fluorigenic substrate libraries reveal unexpected specificity determinants of DUBs deubiquitinating enzymes. Biochem J 415(3):367–375. https://doi.org/10.1042/BJ20080779
Dugat-Bony E, Straub C, Teissandier A, Onesime D, Loux V, Monnet C, Irlinger F, Landau S, Leclercq-Perlat M, Bento P, Fraud S, Gibrat J, Aubert J, Fer F, Guedon E, Pons N, Kennedy S, Beckerich J, Swennen D, Bonnarme P (2015) Overview of a surface-ripened cheese community functioning by meta-omics analyses. PLoS One 10(4):1–25. https://doi.org/10.1371/journal.pone.0124360
El-Kholy W, El-Soda M, Ezzat N, El-Shafei H (1998) Autolysis and intracellular release from cheese related dairy lactobacilli. Lait 78(4):439–452. https://doi.org/10.1051/lait:1998441
Fadda S, Sanz Y, Vignolo G, Aristoy MC, Oliver G, Toldra F (1999) Characterization of muscle sarcoplasmic and myofibrillar protein hydrolysis caused by Lactobacillus plantarum. Appl Environ Microbiol 65(8):3540–3546
Fehr AR, Perlman S (2015) Coronaviruses: an overview of their replication and pathogenesis. Coronaviruses 1282:1–23. https://doi.org/10.1007/978-1-4939-2438_1
FitzGerald RJ, O’Cuinn G (2006) Enzymatic debittering of food protein hydrolysates. Biotechnol Adv 24(2):234–237. https://doi.org/10.1016/j.biotechadv.2005.11.002
Flores M, Sanz Y, Spanier AM (1998) Contribution of muscle and microbial aminoeptidases to flavor development in dry-cured meat products. In: Contis ET, Ho CT, Mussinan CJ, Parliment TH, Shahidi F, Spanier AM (eds) Developments in food science, volume 40. Elsevier, pp 547–557. https://doi.org/10.1016/S0167-4501(98)80076-9
Fuke Y, Matsuoka H (1993) The purification and characterization of prolyl aminopeptidase from Penicillium camemberti. J Dairy Sci 76(9):2478–2484. https://doi.org/10.3168/jds.S0022-0302(93)77582-7
Gallego M, Mora L, Escudero E, Toldra F (2018) Bioactive peptides and free amino acids profiles in different types of European dry-fermented sausages. Int J Food Microb 276:71–78. https://doi.org/10.1016/j.ijfoodmicro.2018.04.009
Gao X, Yin Y, Zhou C (2018) Purification, characterisation and salt-tolerance molecular mechanisms of aspartyl aminopeptidase from Aspergillus oryzae 3.042. Food Chem 240:377–385. https://doi.org/10.1016/j.foodchem.2017.07.081
Gatti M, Fornasari ME, Mucchetti G, Addeo F, Neviani E (1999) Presence of peptidase activities in different varieties of cheese. Lett Appl Microbiol 28(5):368–372. https://doi.org/10.1046/j.1365-2672.1999.00541.x
Giesler L, Linke D, Rabe S, Appel D, Berger RG (2013) Hydrolysis of wheat gluten by combining peptidases of Flammulina velutipes and electrodialysis. J Agric Food Chem 61(36):8641–8649. https://doi.org/10.1021/jf401716m
Giyatmi, Irianto HE (2017) Enzymes in fermented fish. Adv Food Nutr Res 80:199–216. https://doi.org/10.1016/bs.afnr.2016.10.004
Goldberg AL, Akopian TN, Kisselev AF, Lee DH (1997) Protein degradation by the proteosome and dissection of its in vivo importance with synthetic inhibitors. Mol Biol Rep 24(1–2):69–75. https://doi.org/10.1023/A:1006860828265
Goldstein JM, Banbula A, Kordula T, Mayo JA, Travis J (2001) Novel extracellular x-Prolyl Dipeptidyl-Peptidase (DPP) from Streptococcus gordonii FSS2: an emerging subfamily of viridans streptococcal x-Prolyl DPPs. Infect Immun 69(9):5494–5501
Gonzales T, Robert-Baudouy J (1996) Bacterial aminopeptidases: properties and functions. FEMS Microbiol Rev 18(4):319–344. https://doi.org/10.1111/j.1574-6976.1996.tb00247.x
Gray N (2011) Enzymes may boost chocolate flavor: food navigator. https://www.foodnavigator.com/Article/2011/05/17/Enzymes-may-boost-chocolate-flavour-Study. Accessed 1July 2019
Guenet C, Lepage P, Harris BA (1992) Isolation of the leucine aminopeptidase gene from Aeromonas proteolytica. Evidence for an enzyme precursor. J Biol Chem 267 (12):8390–8395
Harris JL, Backes BJ, Leonetti F, Mahrus S, Ellman JA, Craik CS (2000) Rapid and general profiling of protease specificity by using combinatorial fluorogenic substrate libraries. Proc Natl Acad SciU S A97(14):7754–7759. https://doi.org/10.1073/pnas.140132697
Helgren TR, Wangtrakuldee P, Staker BL, Hagen TJ (2016) Advances in bacterial aminopeptidase inhibition. Curr Top Med Chem 16(4):397–414. https://doi.org/10.2174/1568026615666150813145410
Hershcovitz YF, Rabinovitch L, Langut Y, Reiland V, Shoham G, Shoham Y (2004) Identification of the catalytic residues in the double-zinc aminopeptidase from Streptomyces griseus. FEBS Lett 571(1–3):192–196. https://doi.org/10.1016/j.febslet.2004.07.001
Hinrichsen LL, Montel MC, Talon R (1994) Proteolytic and lipolytic activities of Micrococcus roseus (65), Halomonas elongate (16) and Vibrio sp (168) isolated from Danish bacon curing brines. Int J Food Microbiol 22(2–3):115–126. https://doi.org/10.1016/0168-1605(94)90136-8
Holz RC (2002) The aminopeptidase from Aeromonas proteolytica: structure and mechanism of co-catalytic metal centers involved in peptide hydrolysis. Coord Chem Rev 232(1–2):5–26. https://doi.org/10.1016/S0010-8545(01)00470-2
Holz RC, Bzymek KP, Swierczek SI (2003) Co-catalytic metallopeptidases as pharmaceutical targets. Curr Opin Chem Biol 7(2):197–206. https://doi.org/10.1016/S1367-5931(03)00033-4
Hsu YT, Su CY, Du HC, Jao SC, Li WS (2008) Evaluation of organophosphorous chemicals degrading enzymes: a comparison of Escherichia coli and human cytosolic aminopeptidase P. Chem Biodivers 5(7):1401–1411. https://doi.org/10.1002/cbdv.200890128
Huang WQ, Zhong LF, Meng ZZ, You ZJ, Li JZ, Luo XC (2015) The structure and enzyme characteristics of a recombinant leucine aminopeptidase rLap1 from Aspergillus sojae and its application in debittering. Appl Biochem Biotechnol 177(1):190–206. https://doi.org/10.1007/s12010-015-1737-5
Huberman L, Gollop N, Mumcuoglu KY, Breuer E, Bhusare SR, Shai Y, Galun R (2007) Antibacterial substances of low molecular weight isolated from the blowfly, Lucilia sericata. Med Vet Entomol 21(2):127–131. https://doi.org/10.1111/j.1365-2915.2007.00668.x
Inguimbert N, Coric P, Dhotel H, Bonnard E, Llorens-Cortes C, Mota N, Fournie-Zaluski MC, Roques BP (2005) Synthesis and in vitro activities of new non-peptidic APA inhibitors. J Pept Res 65(2):175–188. https://doi.org/10.1111/j.1399-3011.2004.00211.x
Ito K, Nakajima Y, Onohara Y, Takeo M, Nakashima K, Matsubara F, Ito T, Yoshimoto T (2006) Aminopeptidase N (proteobacteria alanyl aminopeptidase) from Escherichia coli: crystal structure and conformational change of the methionine 260 residue involved in substrate recognition. J Biol Chem 281:33664–33676. https://doi.org/10.1074/jbc.M605203200
Izawa N, Tokuyasu K, Hayashi K (1997) Debittering of protein hydrolysates using Aeromonas caviae aminopeptidase. J Agric Food Chem 45(3):543–545. https://doi.org/10.1021/jf960784t
Jankiewicz U, Bielawski W (2003) The properties and functions of bacterial aminoepeptidases. Acta Microbiol Pol 52(3):217–231
Jao SC, Huang LF, Tao YS, Li WS (2004) Hydrolysis of organophosphate triesters by Escherichia coli aminopeptidase P. J Mol Catal 27(1):7–12. https://doi.org/10.1016/j.molcatb.2003.09002
Kauppinen S (1996) An enzyme with aminopeptidase activity. Patent WO 96/28542
Klein JR, Henrich B, Plapp R (1994) Cloning and nucleotide sequence analysis of the Lactobacillus delbriieckii ssp. lactis DSM7290 cysteine aminopeptidase gene pepC. FEMS Microbiol Lett 124(3):291–300. https://doi.org/10.1111/j.1574-6968.1994.tb07299.x
Kratzer U, Frank R, Kalbacher H, Biehl B, Wostemeyer J, Voigt J (2009) Subunit structure of the vicilin-like globular storage protein of cocoa seeds and the origin of cocoa and chocolate specific aroma precursors. Food Chem 113(4):903–913. https://doi.org/10.1016/j.foodchem.2008.08.017
Kusumoto KI, Matsushita-Morita M, Furukawa I, Suzuki S, YamagataY KY (2008) Efficient production and partial characterization of aspartyl aminopeptidase from Aspergillus oryzae. J Appl Microbiol 105(5):1711–1719. https://doi.org/10.1111/j.1365-2672.2008.03889.x
Labrie SJ, Mosterd C, Loignon S, Dupuis ME, Desjardins P, Rousseau GM, Tremblay DM, Romero DA, Horvath P, Fremaux C, Moineau S (2019) A mutation in the methionine aminopeptidase gene provides phage resistance in Streptococcus thermophiles. Sci Rep 9(1):13816. https://doi.org/10.1038/s41598-019-49975-4
Lee Y, Lee S, Jjung C, Kim H, Choi S, Kim J, Kim H, Seo J (2006) Aminopeptidase derived from Bacillus licheniformis, gene Encoding the aminopeptidase, expression vector containing the gene, transformant and method for preparation thereof, US Patent 7,098,018 B2, 29 Aug 2006
Lei F, Zhao Q, Sun-Waterhouse D, Zhao M (2017) Characterization of a salt-tolerant aminopeptidase from marine Bacillus licheniformis SWJS33 that improves hydrolysis and debittering efficiency for soy protein isolate. Food Chem 214:347–353. https://doi.org/10.1016/j.foodchem.2016.07.028
Lei F, Hu C, Zhang N, He D (2018) The specificity of an aminopeptidase affects its performance in hydrolyzing peanut protein isolate and zein. LWT Food Sci Technol 102:37–44. https://doi.org/10.1016/j.lwt.2018.10.041
Lemieux L, Simard RE (1991) Bitter flavour in dairy products. I. A review of the factors likely to influence its development, mainly in cheese manufacture. Lait 71(6):599–636. https://doi.org/10.1051/lait:1991647
Li X, Chang YH (1995) Amino-terminal protein processing in Saccharomyces cerevisiae is an essential function that requires two distinct methionine aminopeptidases. Proc Natl Acad SciU S A92(26):12357–12361. https://doi.org/10.1073/pnas.92.26.12357
Lin LL, Hsu WH, Wu CP, Chi MC, Chou WM, Hu HY (2004) A thermostable leucine aminopeptidase from Bacillus kaustophilus CCRC 11223. Extremophiles 8(1):79–87. https://doi.org/10.1007/s00792-003-0364-1
Lowther WT, Matthews BW (2002) Metalloaminopeptidases: common functional themes in disparate structural surroundings. Chem Rev 102(12):4581–4608. https://doi.org/10.1021/cr0101757
Macedo AC, Tˆania G, Tavares F, Malcata X (2003) Purification and characterization of an intracellular aminopeptidase from a wild strain of Lactobacillus plantarum isolated from traditional Serra da Estrela cheese. Enzym Microb Technol 32(1):41–48. https://doi.org/10.1016/S0141-0229(02)00234-X
Maggiora LL, Orawski AT, Simmons WH (1999) Apstatin analogue inhibitors of Aminopeptidase P, a bradykinin degrading enzyme. J Med Chem 42(13):2394–2402. https://doi.org/10.1021/jm9805642
Markets and Markets (2019) Food enzymes market by type (carbohydrase, protease, lipase), application (beverages, bakery products, dairy products, confectionery products, processed foods), formulation (lyophilized powder and liquid), source, and region - global forecast to 2023. https://www.marketsandmarkets.com/Market-Reports/food-enzymes-market-800.html. Accessed 30 June 2019
Martins JM, Galinari E, Pimentel-Filho NJ, Ribeiro JI Jr, Furtado MM, Ferreira CLLF (2015) Determining the minimum ripening time of artisanal Minas cheese, a traditional Brazilian cheese. Braz J Microbiol 46(1):219–230. https://doi.org/10.1590/S1517-838246120131003
Matsushita-Morita M, Furukawa I, Suzuki S, Yamagata Y, Koide Y, Ishida H, Kusumoto KI (2010) Characterization of recombinant prolyl aminopeptidase from Aspergillus oryzae. J Appl Microbiol 109(1):156–165. https://doi.org/10.1111/j.1365-2672.2009.04641.x
Merz M, Eisele T, Berends P, Appel D, Rabe S, Blankk I, Stressler T, Fischer L (2015) Flavourzyme, an enzyme preparation with industrial relevance: automated nine-step purification and partial characterization of eight enzymes. Agric Food Chem 63(23):5682–5693. https://doi.org/10.1021/acs.jafc.5b01665
Mest HJ (2006) Dipeptidyl peptidase-IV inhibitors can restore glucose homeostasis in type 2 diabetics via incretin enhancement. Curr Opin Investig Drugs 7(4):338–343
Meyer J, Butikofer U, Walther B, Wechsler D, Sieber R (2009) Hot topic: changes in angiotensin-converting enzyme inhibition and concentrations of the tripeptides Val-Pro-Pro and Ile-Pro-Pro during ripening of different Swiss cheese variations. J Dairy Sci 92(3):826–836. https://doi.org/10.3168/jds.2008-1531
Meyer-Barton E, Klein JR, Henrich B, Plapp R (1994) X-prolyl-dipeptidyl peptidase from Lactobacillus delbrueckii ssp. lactis, nucleic acids coding for the same and its use in fermented foodstuff preparation process. Patent WO94/16082, 21 July 1994
Mora L, Gallego M, Escudero E, Reig M, Aristoy M, Toldra F (2015) Small peptides hydrolysis in dry-cured meats. Int J Food Microbiol 212:9–15. https://doi.org/10.1016/j.ijfoodmicro.2015.04.018
Mora L, Gallego M, Toldra F (2018) ACEI-inhibitory peptides naturally generated in meat and meat products and their health relevance. Nutrients 10(9):1259–1271. https://doi.org/10.3390/nu10091259
Nampoothiri KM, Nagy V, Kovacs K, Szakacs G, Pandey A (2005) L-leucine aminopeptidase production by filamentous Aspergillus fungi. Lett Appl Microbiol 41(6):498–504. https://doi.org/10.1111/j.1472-765X.2005.01789.x
Nandan AS, Nampoothiri KM (2014) Unveiling aminopeptidase P from Streptomyces lavendulae: molecular cloning, expression and biochemical characterization. Enzym Microb Technol 55:7–13. https://doi.org/10.1016/j.enzmictec.2013.11.003
Nandan A, Nampoothiri KM (2017a) Molecular advances in microbial aminopeptidases. Bioresour Technol 245(Part B):1757–1765. https://doi.org/10.1016/j.biortech.2017.05.103
Nandan A, Nampoothiri KM (2017b) Chapter 21 microbial aminopeptidases. In: Pandey A, Negi S, Soccol CR (eds) Current developments in biotechnology and bioengineering. Production, Isolation and Purification of Industrial Products. Elsevier, Cambridge, pp 491–507.
Nandan A, Gaurav A, Pandey A, Nampoothiri KM (2010) Arginine specific aminopeptidase from Lactobacillus brevis. Braz Arch Biol Technol 53(6):1443–1450. https://doi.org/10.1590/S1516-89132010000600021
Nandan A, Pandey A, Nampoothiri KM (2011) Proline-specific extracellular aminopeptidase purified from Streptomyces lavendulae. Appl Biochem Biotechnol 163(8):994–1001. https://doi.org/10.1007/s12010-010-9103-0
Nemoto YO, Shimoyama Y, Nakasato M, Nishimata H, Ishikawa T, Sasaki M, Kimura S, Nemoto TK (2018) Distribution of dippeptidyl peptidase (DPP)4, DPP5, DPP5, DPP7 and DPP11 in human oral microbiota-potent biomarkers indicating presence of periodontopathic bacteria. FEMS Microbiol Lett 365(22):221. https://doi.org/10.1093/femsle/fny221
Nishiwaki T, Yoshimizu S, Furuta M, Hayashi K (2002) Debittering of enzymatic hydrolysates using an aminopeptidase from the edible Basidiomycete Grifola frondosa. J Biosci Bioeng 93(1):60–63. https://doi.org/10.1016/S1389-1723(02)80055-X
Oliveira HS, Mamede ME, Góes-Neto A, Koblitz MG (2011) Improving chocolate flavor in poor-quality cocoa almonds by enzymatic treatment. J Food Sci 76(5):755–759. https://doi.org/10.1111/j.1750-3841.2011.02168.x
Pan D, Tanokura M (2004) Purification and characterization of an aminopeptidase from Lactobacillus helveticus JCM 1004. Food Chem 88(4):511–516. https://doi.org/10.1016/j.foodchem.2004.01.082
Pan D, Luo Y, Tanokura M (2005) Antihypertensive peptides from skimmed milk hydrolysate digested by cell-free extract of Lactobacillus helveticus JCM1004. Food Chem 91(1):123–129. https://doi.org/10.1016/j.foodchem.2004.05.055
Petrovic N, Schacke W, Gahagan JR, O'Conor CA, Winnicka B, Conway RE, Mina-Osorio P, Shapiro LH (2007) CD13/APN regulates endothelial invasion and filopodia formation. Blood 110(1):142–150. https://doi.org/10.1182/blood-2006-02-002931
Rahulan R, Nampoothiri KM, Szakacs G, Nagy V, Pandey A (2009) Statistical optimization of L-leucine aminopeptidase production from Streptomyces gedenensis IFO 13427 under submerged fermentation using response surface methodology. Biochem Eng J43(1):64–71. https://doi.org/10.1016/j.bej.2008.08.011
Rahulan R, Dhar K, Nampoothiri K, Pandey A (2012) Aminopeptidase from Streptomyces gedanensis as a useful tool for protein hydrolysate preparations with improved functional properties. J Food Sci 77(7):791–797. https://doi.org/10.1111/j.1750-3841.2012.02773.x
Rawlings ND, Tolle DP, Barrett AJ (2004) MEROPS: the peptidase database. Nucleic Acids Res 32(database issue):160–164. https://doi.org/10.1093/nar/gkh071
Research and Markets, Global Industrial Enzymes Market Growth, Trends, and Forecast 2019-2024 Competition for Raw Materials with Other Industries and Price Volatility Restraining Market Growth (2019). https://www.globenewswire.com/news-release/2019/03/29/1788385/0/en/Global-Industrial-EnzymesMarket-Growth-Trends-and-Forecast-2019-2024-Competition-for-Raw-Materials-with-Other-Industries-and-Price-Volatility-Restraining-Market-Growth.html. Accessed 30 June 2019
Rigolet P, Xi XG, Rety S, Chich JF (2005) The structural comparison of the bacterial PepX and human DPP-IV reveals sites for the design of inhibitors of PepX activity. FEBS J 272(8):2050–2059. https://doi.org/10.1111/j.1742-4658.2005.04631.x
Sanz Y, Fadda S, Vignolo G, Aristoy MC, Oliver G, Toldra F (1999) Hydrolytic action of Lactobacillus casei CRL 705 on pork muscle sarcoplasmic and myofibrillar proteins. J Agric Food Chem 47(8):3441–3448. https://doi.org/10.1021/jf981255n
Scharf U, Stolz P, Huscroft SC, Schmidt-Hahn K (2006) Use of aminopeptidase in dough, doughs and bread improvers comprising aminopeptidase. Patent Application. WO2006/009447, 26 Jan 2006
Schiffmann R, Neugebauer A, Klein CD (2006) Metal-mediated inhibition of Escherichia coli methionine aminopeptidase: structure-activity relationships and development of a novel scoring function for metal-ligand interactions. J Med Chem 49(2):511–522. https://doi.org/10.1021/jm050476z
Schuster E, Sprossler B, Titze K, Gottschalk M, Khanh NQ, Wolf S, Plainer H (1997) Leucine aminopeptidases produced recombinantly from Aspergillus soyae, Patent no: WO 97/04108, 6 Feb 1997
Scornik OA, Botbol V (2001) Bestatin as an experimental tool in mammals. Curr Drug Metab 2(1):67–85. https://doi.org/10.2174/1389200013338748
Selvakumar P, Lakshmikuttyamma A, Dimmock JR, Sharma RK (2005) Methionine aminopeptidase 2 and cancer. Biochim Biophys Acta 1765(2):148–154. https://doi.org/10.1016/j.bbcan.2005.11.001
Shen Y, Wang F, Lan D, Liu Y, Yang B, Wang Y (2011) Biochemical properties and potential applications of recombinant leucine aminopeptidase from Bacillus kaustophilus CCRC 11223. Int J Mol Sci 12(11):7609–7625. https://doi.org/10.3390/ijms12117609
Shigemura Y, Iwai K, Morimatsu F, Iwamoto T, Mori T, Oda C, Taira T, Park EY, Nakamura Y, Sato K (2009) Effect of prolyl-hydroxyproline(Pro-Hyp), a food-derived collagen peptide in human blood, on growth of fibroblasts from mouse skin. J Agric Food Chem 57(2):444–449. https://doi.org/10.1021/jf802785h
Sidhu GK, Singh S, Kumar V, Dhanjal DS, Datta S, Singh J (2019) Toxicity, monitoring and biodegradation of organophosphate pesticides: a review. Crit Rev Env Sci Tec 49(13):1135–1187. https://doi.org/10.1080/10643389.2019.1565554
Singh BK, Walker A (2006) Microbial degradation of organophosphorus compounds. FEMS Microbiol Rev 30(3):428–471. https://doi.org/10.1111/j.1574-6976.2006.00018.x
Sonoda H, Daimon K, Yamaji H, Sugimura A (2009) Efficient production of active Vibrio proteolyticus aminopeptidase in Escherichia coli by co-expression with engineered vibriolysin. Appl Environ Microbiol 84(1):191–198. https://doi.org/10.1007/s00253-009-2089-2
Sonoda H, Daimon K, Sugimura A, (2014) Method for producing aminopeptidase. EP 2 206777B1, 15 Jan 2014
Sousa LS, Rocha FS, Silveira PTD, Bispo ED, Soares SE (2016) Enzymatic activity of proteases and its isoenzymes in fermentation process in cultivars of cocoa (Theobroma cacao L.) produced in southern Bahia, Brazil. Food Sci Technol Campinas 36(4):656–663. https://doi.org/10.1590/1678-457x.10916
Stressler T, Eisele T, Schlayer M, Lutz-Wahl S, Fischer L (2013) Characterization of the recombinant exopeptidases pepx and pepn from Lactobacillus helveticus ATCC 12046 important for food protein hydrolysis. PLoS One 8(7):1–12. https://doi.org/10.1371/journal.pone.0070055
Stressler T, Ewert J, Merz M, Funk J, Claaßen W, Lutz-Wahl S (2016) A novel glutamyl (aspartyl)-specific aminopeptidase a from Lactobacillus delbrueckii with promising properties for application. PLoS One 11(3):1521–1139. https://doi.org/10.1371/journal.pone.0152139
Suresh PV, Prabhu GN (2013) Seafood. In: Chandrasekaran M (ed) valorization of food processing byproducts. CRC Press, Boca Raton, pp 685–736
Taylor A (1993a) Aminopeptidases: structure and function. FASEB J 7(2):290–298. https://doi.org/10.1096/fasebj.7.2.8440407
Taylor A (1993b) Aminopeptidase towards a mechanism of action. Trends Biochem Sci 18(5):167–171
Tchorbanov B, Marinova MD, Grozeva L (2011) Debittering of protein hydrolysates by Lactobacillus LBL-4 aminopeptidase. Enzyme Res 10:1–7. https://doi.org/10.4061/2011/538676
Toldra F, Reig M, Aristoy M, Mora L (2018) Generation of bioactive peptides during food processing. Food Chem 267:395–404. https://doi.org/10.1016/j.foodchem.2017.06.119
Tomohiro S, Nami N, Tomohiro K, Noriki N, Noriaki Y, Hidehiko W (2006) Method of improving taste and/or favor of food or drink. Patent application EP 1629719 A1, 1 Mar 2006
Udomsil N, Rodtong S, Tanasupawat S, Yongsawatdigul J (2015) Improvement of fish sauce quality by strain CMC5-3-1: a novel species of Staphylococcus sp. J Food Sci 80(9):2015–2022. https://doi.org/10.1111/1750-3841.12986
Umezawa Y, Yokoyama K, Kikuchi Y, Date M, Ito K, Yoshimoto T, Matsui H (2004) Novel prolyl tri/tetra-peptidyl aminopeptidase from Streptomyces mobaraensis: substrate specificity and enzyme gene cloning. J Biochem 136(3):293–300. https://doi.org/10.1093/jb/mvh129
Umitsuki G, Abe K (2000) Leucine aminopeptidase gene, recombinant DNA, and process for producing leucine aminopeptidase, US 6,127,161, 3 Oct 2000. https://doi.org/10.1371/journal.ppat.1006310
Vesanto E, Varmanen P, Steele JL, Palva A (1994) Characterization and expression of the Lactobacillus helveticus pepC gene encoding a general aminopeptidase. Eur J Biochem 224(3):991–997. https://doi.org/10.1111/j.1432-1033.1994.00991.x
Virgili R, Schivazappa C, Parolari G, Bordini CS, Degni M (1998) Proteases in fresh pork muscle and their influence on bitter taste formation in dry-cured ham. J Food Biochem 22(1):53–63. https://doi.org/10.1111/j.1745-4514.1998.tb00230.x
Visser S (1993) Proteolytic enzymes and their relation to cheese ripening and flavor; an overview. J Dairy Sci 76(1):329–350. https://doi.org/10.3168/jds.S0022-0302(93)77354-3
Voigt J, Biehl B, Heinrichs H, Kamaruddin S, Marsoner GC, Hugi A (1994a) In-vitro formation of cocoa-specific aroma precursors: aroma related peptides generated from cocoa seed protein by co-operation of an aspartic endoprotease and a carboxipeptidase. Food Chem 49(2):173–180. https://doi.org/10.1016/0308-8146(94)90155-4
Voigt J, Heinrichs H, Voigt G, Biehl B (1994b) Cocoa specific aroma precursors are generated by proteolytic digestion of the vicilin-like globulin of cocoa seeds. Food Chem 50(2):177–184. https://doi.org/10.1016/0308-8146(94)90117-1
Vo-Van T, Kusakabe I, Murakami K (1984) The aminopeptidase activity in fish sauce. Agric Biol Chem 48(2):525–527. https://doi.org/10.1080/00021369.1984.10866173
Watanabe J, Tanaka H, Akagawa T, Mogi Y, Yamazaki T (2007) Characterization of Aspergillus oryzae aspartyl aminopeptidase expressed in Escherichia coli. Biosci Biotechnol Biochem 71(10):2557–2560. https://doi.org/10.1271/bbb.70107
Wenzel F, Uhlig H, Lehmann K (1968) Aminopeptidase cleaving L-leucinamide, hypertension and oxytocin. US 3,405,034, 8 Oct 1968
Wu YY, Cao SM (2018) Study on endogenous protease and protein degradation of dry-salted Decapterus maruadsi. CyTA-J Food 16(1):350–356. https://doi.org/10.1080/19476337.2017.1406006
Wu B, Shi P, Li J, Wang Y, Meng K, Bai Y, Yao B (2010) A new aminopeptidase from the keratin-degrading strain Streptomyces fradiae var. k11. Appl Biochem Biotechnol 160(3):730–739. https://doi.org/10.1007/s12010-009-8537-8
Wu YT, Zhou ND, Zhou ZM, Gao XX, Tian YP (2014) A thermo-stable lysine aminopeptidase from Pseudomonas aeruginosa: isolation, purification, characterization, and sequence analysis. J Basic Microbiol 54(10):1110–1119. https://doi.org/10.1002/jobm.201300752
Yamamoto Y, Usuki H, Iwabuchi M, Hatanaka T (2010) Prolyl aminopeptidase from Streptomyces thermoluteus sub sp. Fuscus strain NBRC14270 and synthesis of proline-containing peptides by its S144C variant. Appl Environ Microbiol 76(18):6180–6185. https://doi.org/10.1128/AEM.01242-10
Yokoyama R, Kawasaki H, Hirano H (2006) Identification of yeast aspartyl aminopeptidase gene by purifying and characterizing its product from yeast cells. FEBS J 273(1):192–198. https://doi.org/10.1111/j.1742-4658.2005.05057.x
Zhao G, Ding LL, Yao Y, Yanping C, Pan ZH, Kong DH (2018) Extracellular proteome analysis and flavor formation during soy sauce fermentation. Front Microbiol 9:1–7. https://doi.org/10.3389/fmicb.2018.01872
Zheng B, Liu Y, He X, Hu S, Li S, Chen M, Jiang W (2017) Quality improvement on half –fin anchovy (Setipinna taty) fish sauce by Psychrobacter sp. SP-1 fermentation. J Sci Food Agric 97(13):4484–4493. https://doi.org/10.1002/jsfa.8313
Zhong H, Bowen JP (2006) Antiangiogenesis drug design: multiple pathways targeting tumor vasculature. Curr Med Chem 13(8):849–862. https://doi.org/10.2174/092986706776361085
Zotta T, Ricciardi A, Parente E (2007) Enzymatic activities of lactic acid bacteria isolated from Cornetto di Matera sourdough. Int J Food Microbiol 115(2):165–172. https://doi.org/10.1016/j.ijfoodmicro.2006.10.026
Acknowledgments
AN acknowledges the research fellowship from Department of Science and Technology (DST)-INSPIRE, New Delhi and KMN grateful to CSIR, New Delhi for various fundings.
Author information
Authors and Affiliations
Contributions
AN, the first author collected all the data and wrote the article. KMN, the corresponding author conceived the topic, drafted it, and did critical reading to prepare the manuscript. Both authors read and approved the manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors declared that they have no conflict of interest regarding this manuscript.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Nandan, A., Nampoothiri, K.M. Therapeutic and biotechnological applications of substrate specific microbial aminopeptidases. Appl Microbiol Biotechnol 104, 5243–5257 (2020). https://doi.org/10.1007/s00253-020-10641-9
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00253-020-10641-9