Abstract
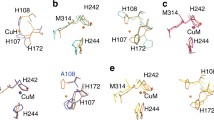
Many bioactive peptides, such as hormones and neuropeptides, require amidation at the C terminus for their full biological activity. Peptidylglycine α-hydroxylating monooxygenase (PHM) performs the first step of the amidation reaction—the hydroxylation of peptidylglycine substrates at the Cα position of the terminal glycine. The hydroxylation reaction is copper- and O2-dependent and requires 2 equiv of exogenous reductant. The proposed mechanism suggests that O2 is reduced by two electrons, each provided by one of two nonequivalent copper sites in PHM (CuH and CuM). The characteristics of the reduced oxygen species in the PHM reaction and the identity of the reactive intermediate remain uncertain. To further investigate the nature of the key intermediates in the PHM cycle, we determined the structure of the oxidized form of PHM complexed with hydrogen peroxide. In this 1.98-Å-resolution structure (hydro)peroxide binds solely to CuM in a slightly asymmetric side-on mode. The O–O interatomic distance of the copper-bound ligand is 1.5 Å, characteristic of peroxide/hydroperoxide species, and the Cu–O distances are 2.0 and 2.1 Å. Density functional theory calculations using the first coordination sphere of the CuM active site as a model system show that the computed energies of the side-on L3CuM(II)–O2 2− species and its isomeric, end-on structure L3CuM(I)–O2 ·− are similar, suggesting that both these intermediates are significantly populated within the protein environment. This observation has important mechanistic implications. The geometry of the observed side-on coordinated peroxide ligand in L3CuM(II)O2 2− is in good agreement with the results of a hybrid quantum mechanical–molecular mechanical optimization of this species.







Similar content being viewed by others
Abbreviations
- AIM:
-
Atoms in molecules
- DFT:
-
Density functional theory
- MM:
-
Molecular mechanical
- oxPHM:
-
Oxidized form of peptidylglycine α-hydroxylating monooxygenase
- oxPHMcc:
-
Oxidized catalytic core of peptidylglycine α-hydroxylating monooxygenase
- PAM:
-
Peptidylglycine α-amidating monooxygenase
- PAL:
-
Peptidyl-α-hydroxyglycine α-amidating lyase
- PDB:
-
Protein Data Bank
- PHM:
-
Peptidylglycine α-hydroxylating monooxygenase
- PHMcc:
-
Catalytic core of peptidylglycine α-hydroxylating monooxygenase
- QM:
-
Quantum mechanical
References
Merkler DJ, Kulathila R, Consalvo AP, Young SD, Ash DE (1992) Biochemistry 31:7282–7288
Noguchi M, Seino H, Kochi H, Okamoto H, Tanaka T, Hirama M (1992) Biochem J 283(Pt 3):883–888
Prigge ST, Mains RE, Eipper BA, Amzel LM (2000) Cell Mol Life Sci 57:1236–1259
Eipper BA, Milgram SL, Husten EJ, Yun HY, Mains RE (1993) Protein Sci 2:489–497
Katopodis AG, May SW (1990) Biochemistry 29:4541–4548
Suzuki K, Ohta M, Okamoto M, Nishikawa Y (1993) Eur J Biochem 213:93–98
Glauder J, Ragg H, Rauch J, Engels JW (1990) Biochem Biophys Res Commun 169:551–558
Ouafik L, May V, Saffen DW, Eipper BA (1990) Mol Endocrinol 4:1497–1505
Eipper BA, Stoffers DA, Mains RE (1992) Annu Rev Neurosci 15:57–85
Czyzyk TA, Ning Y, Hsu MS, Peng B, Mains RE, Eipper BA, Pintar JE (2005) Dev Biol 287:301–313
Jiang N, Kolhekar AS, Jacobs PS, Mains RE, Eipper BA, Taghert PH (2000) Dev Biol 226:118–136
Prigge ST, Kolhekar AS, Eipper BA, Mains RE, Amzel LM (1997) Science 278:1300–1305
Prigge ST, Eipper BA, Mains RE, Amzel LM (2004) Science 304:864–867
Prigge ST, Kolhekar AS, Eipper BA, Mains RE, Amzel LM (1999) Nat Struct Biol 6:976–983
Siebert X, Eipper BA, Mains RE, Prigge ST, Blackburn NJ, Amzel LM (2005) Biophys J 89:3312–3319
Jaron S, Blackburn NJ (1999) Biochemistry 38:15086–15096
Rhames FC, Murthy NN, Karlin KD, Blackburn NJ (2001) J Biol Inorg Chem 6:567–577
Jaron S, Mains RE, Eipper BA, Blackburn NJ (2002) Biochemistry 41:13274–13282
Chen P, Bell J, Eipper BA, Solomon EI (2004) Biochemistry 43:5735–5747
Eipper BA, Quon AS, Mains RE, Boswell JS, Blackburn NJ (1995) Biochemistry 34:2857–2865
Freeman JC, Nayar PG, Begley TP, Villafranca JJ (1993) Biochemistry 32:4826–4830
Blackburn NJ, Rhames FC, Ralle M, Jaron S (2000) J Biol Inorg Chem 5:341–353
Francisco WA, Blackburn NJ, Klinman JP (2003) Biochemistry 42:1813–1819
Francisco WA, Knapp MJ, Blackburn NJ, Klinman JP (2002) J Am Chem Soc 124:8194–8195
Francisco WA, Merkler DJ, Blackburn NJ, Klinman JP (1998) Biochemistry 37:8244–8252
Tian G, Berry JA, Klinman JP (1994) Biochemistry 33:226–234
Freeman JC, Villafranca JJ (1993) J Am Chem Soc 115:4923–4924
Crespo A, Marti MA, Roitberg AE, Amzel LM, Estrin DA (2006) J Am Chem Soc 128:12817–12828
Klinman JP (2006) J Biol Chem 281:3013–3016
Yoshizawa K, Kihara N, Kamachi T, Shiota Y (2006) Inorg Chem 45:3034–3041
Evans JP, Ahn K, Klinman JP (2003) J Biol Chem 278:49691–49698
Decker A, Solomon EI (2005) Curr Opin Chem Biol 9:152–163
Chen P, Solomon EI (2004) J Am Chem Soc 126:4991–5000
Kamachi T, Kihara N, Shiota Y, Yoshizawa K (2005) Inorg Chem 44:4226–4236
Messerschmidt A, Luecke H, Huber R (1993) J Mol Biol 230:997–1014
Bento I, Martins LO, Gato Lopes G, Armenia Carrondo M, Lindley PF (2005) Dalton Trans 3507–3513
Shin DS, Didonato M, Barondeau DP, Hura GL, Hitomi C, Berglund JA, Getzoff ED, Cary SC, Tainer JA (2009) J Mol Biol 385:1534–1555
Kolhekar AS, Keutmann HT, Mains RE, Quon AS, Eipper BA (1997) Biochemistry 36:10901–10909
Otwinowski Z, Minor W (1997) Methods Enzymol 276:307–326
Murshudov GN, Vagin AA, Dodson EJ (1997) Acta Crystallogr D Biol Crystallogr 53:240–255
Collaborative Computational Project N (1994) Acta Crystallogr D Biol Crystallogr 50: 760–763
Frisch MJ et al (2004) Gaussian 03. Gaussian, Wallingford
Henkelman G, Arnaldsson A, Jonsson A (2006) Comput Mater Sci 36:354–360
Crespo A, Scherlis DA, Marti MA, Ordejon P, Roitberg AE, Estrin DA (2003) J Phys Chem B 107:13728–13736
Soler JM, Artacho E, Gale JD, Garcia A, Junquera J, Ordejon P, Sanchez-Portal D (2002) J Phys Condens Matter 2745–2779
Marti MA, Scherlis DA, Doctorovich FA, Ordejon P, Estrin DA (2003) J Biol Inorg Chem 8:595–600
Perdew JP, Burke K, Ernzerhof M (1996) Phys Rev Lett 77:3865–3868
Wang J, Cieplak P, Kollman P (2000) J Comput Chem 21:1049–1074
Eichinger M, Tavan P, Hutter J, Parrinello M (1999) J Chem Phys 110:10452–10467
Gubelmann MH, Williams AF (1983) Struct Bonding (Berl) 55:1
Chufan EE, Prigge ST, Siebert X, Eipper BA, Mains RE, Amzel LM (2010) J Am Chem Soc 132:15565–15572
Bauman AT, Yukl ET, Alkevich K, McCormack AL, Blackburn NJ (2006) J Biol Chem 281:4190–4198
Hrycay EG, Gustafsson JA, Ingelman-Sundberg M, Ernster L (1976) Eur J Biochem 61:43–52
Davydov R, Makris TM, Kofman V, Werst DE, Sligar SG, Hoffman BM (2001) J Am Chem Soc 123:1403–1415
Froland WA, Andersson KK, Lee SK, Liu Y, Lipscomb JD (1992) J Biol Chem 267:17588–17597
Wolfe MD, Lipscomb JD (2003) J Biol Chem 278:829–835
Gherman BF, Tolman WB, Cramer CJ (2006) J Comput Chem 27:1950–1961
Miller SM, Klinman JP (1985) Biochemistry 24:2114–2127
Osako T, Nagatomo S, Tachi Y, Kitagawa T, Itoh S (2002) Angew Chem Int Ed 41:4325–4328
Mirica LM, Ottenwaelder X, Stack TD (2004) Chem Rev 104:1013–1045
Chen P, Fujisawa K, Solomon EI (2000) J Am Chem Soc 122:10177–10193
Wada A, Harata M, Hasegawa K, Jitsukawa H, Masuda M, Mukai M, Kitagawa T, Einaga H (1998) Angew Chem Int Ed 37:798–799
Maiti D, Lucas HR, Sarjeant AA, Karlin KD (2007) J Am Chem Soc 129:6998–6999
Maiti D, Sarjeant AA, Karlin KD (2007) J Am Chem Soc 129:6720–6721
Wilmot CM (2003) Biochem Soc Trans 31:493–496
Wilmot CM, Hajdu J, McPherson MJ, Knowles PF, Phillips SE (1999) Science 286:1724–1728
Matoba Y, Kumagai T, Yamamoto A, Yoshitsu H, Sugiyama M (2006) J Biol Chem 281:8981–8990
Acknowledgments
We acknowledge Jean Jakoncic and Vivian Stojanoff (beamline X6A of the National Synchrotron Light Source, Brookhaven National Laboratory) for assistance in the collection of X-ray diffraction data. We thank Sandra Gabelli and Mario Bianchet for assistance in the purification of the protein and crystallographic experiments. This work was supported by National Science Foundation grant MCB-920288 and National Institutes of Health grant DK-32949.
Author information
Authors and Affiliations
Corresponding author
Additional information
An interactive 3D complement page in Proteopedia is available at http://proteopedia.org/w/Journal:JBIC:18
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Rudzka, K., Moreno, D.M., Eipper, B. et al. Coordination of peroxide to the CuM center of peptidylglycine α-hydroxylating monooxygenase (PHM): structural and computational study. J Biol Inorg Chem 18, 223–232 (2013). https://doi.org/10.1007/s00775-012-0967-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00775-012-0967-z