Abstract
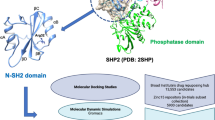
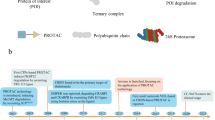
Over expression of T-lymphokine–activated killer cell–originated protein kinase (TOPK) has been associated with leukemia, myeloma tumors and various other cancers. The function and regulatory mechanism of TOPK in tumor cells remains unclear. Structural studies that could reveal the regulatory mechanism have been a challenge because of the unavailabity of TOPK’s crystal structure. Hence, in this study, the 3D structure of TOPK protein has been constructed by using multiple templates. The quality and reliability of the generated model was checked and the molecular dynamics method was utilized to refine the model. APBS method was employed to know the electrostatic potential surface of the modeled protein and it was found that the optimum pH for protein stability is 3.4 which will further help in mechanistic hypothesis of TOPK protein. Active site of TOPK was identified from available literature and HTVS was employed to identify the lead molecules. The expected binding modes of protein-ligand complexes were reproduced in the MD simulation which indicates that the complex is relatively stable. The pharmacokinetic properties of the lead molecules are also under acceptable range. TOPK act as a substrate for CDK1 and the protein-protein docking and dynamics studies were carried out to analyze the effect of Thr9Ala mutation of TOPK in the two protein complex formation. It shows that the wild type complex is more stable when compared with the mutant type. Such structural information at atomic level not only exhibits the action modes of TOPK inhibitors but also furnishes a novel starting point for structure based drug design of TOPK inhibitors.









Similar content being viewed by others
References
Abe Y, Matsumoto S, Kito K, Ueda N (2000) Cloning and expression of a novel MAPKK-like protein kinase, lymphokine-activated killer T-cell-originated protein kinase, specifically expressed in the testis and activated lymphoid cells. J Biol Chem 275:21525–21531
Gaudet S, Branton D, Lue RA (2000) Characterization of PDZ-binding kinase, a mitotic kinase. Proc Natl Acad Sci USA 97:5167–5172
Simons-Evelyn M, Bailey-Dell K, Toretsky JA, Ross DD, Fenton R, Kalvakolanu D, Rapoport AP (2001) PBK/TOPK is a novel mitotic kinase which is upregulated in Burkitt’s lymphoma and other highly proliferative malignant cells. Blood Cells Mol Dis 27:825–829
Côté S, Simard C, Lemieux R (2002) Regulation of growth-related genes by interleukin-6 in murine myeloma cells. Cytokine 20:113–120
Yuryev A, Wennogle LP (2003) Novel raf kinase protein-protein interactions found by an exhaustive yeast two-hybrid analysis. Genomics 81:112–125
Dougherty JD, Garcia ADR, Nakano I, Livingstone M, Norris B, Polakiewicz R, Wexler EM, Sofroniew MV, Kornblum HI, Geschwind DH (2005) PBK/TOPK, a proliferating neural progenitor-specific mitogen-activated protein kinase kinase. J Neurosci 25:10773–10785
Matsumoto S, Abe Y, Fujibuchi T, Takeuchi T, Kito K, Ueda N, Shigemoto K, Gyo K (2004) Characterization of a MAPKK-like protein kinase TOPK. Biochem Biophys Res Commun 325:997–1004
Nandi AK, Rapoport AP (2006) Expression of PDZ-binding kinase (PBK) is regulated by cell cycle-specific transcription factors E2F and CREB/ATF. Leuk Res 30:437–447
Park JH, Lin ML, Nishidate T, Nakamura Y, Katagiri T (2006) PDZ-binding kinase/T-LAK cell-originated protein kinase, a putative cancer/testis antigen with an oncogenic activity in breast cancer. Cancer Res 66:9186–9195
Zhu F, Zykova TA, Kang BS, Wang Z, Ebeling MC, Abe Y, Ma WY, Bode AM, Dong Z (2007) Bidirectional signals transduced by TOPK-ERK interaction increase tumorigenesis of HCT116 colorectal cancer cells. Gastroenterology 133:219–231
Oh SM, Zhu F, Cho YY, Lee KW, Kang BS, Kim HG, Zykova T, Bode AM, Dong Z (2007) T-lymphokine-activated killer cell-originated protein kinase functions as a positive regulator of c-Jun-NH2-kinase 1 signaling and H-Ras-induced cell transformation. Cancer Res 67:5186–5194
Bairoch A, Apweiler R (2000) The SWISS-PROT protein sequence database and its supplement TrEMBL in 2000. Nucleic Acids Res 28:45–48
Wang Z, Liu J, Sudom A, Ayres M, Li S, Wesche H, Powers JP, Walker NPC (2006) Crystal structures of IRAK-4 kinase in complex with inhibitors: a serine/threonine kinase with tyrosine as a gatekeeper. Structure 14:1835–1844
Brennan DF, Dar AC, Hertz NT, Chao WCH, Burlingame AL, Shokat KM, Barford D (2011) A Raf-induced allosteric transition of KSR stimulates phosphorylation of MEK. Nature 472:366–369
Chrencik JE, Patny A, Leung IK, Korniski B, Emmons TL, Hall T, Weinberg RA, Gormley JA, Williams JM, Day JE, Hirsch JL, Kiefer JR, Leone JW, Fischer HD, Sommers CD, Huang HC, Jacobsen EJ, Tenbrink RE, Tomasselli AG, Benson TE (2010) Structural and thermodynamic characterization of the TYK2 and JAK3 kinase domains in complex with CP-690550 and CMP-6. J Mol Biol 400:413–433
Rokas A, Williams BL, King N, Carroll SB (2003) Genome-scale approaches to resolving incongruence in molecular phylogenies. Nature 425:798–804
Šali A, Potterton L, Yuan F, van Vlijmen H, Karplus M (1995) Evaluation of comparative protein modeling by MODELLER. Proteins 23:318–326
Delano WL (2002) The PyMOL molecular graphics system. DeLano Scientific, Palo Alto, CA
Colovos C, Yeates TO (1993) Verification of protein structures: patterns of nonbonded atomic interactions. Protein Sci 2:1511–1519
Wiederstein M, Sippl MJ (2007) ProSA-web: interactive web service for the recognition of errors in three-dimensional structures of proteins. Nucleic Acids Res 35:407–410
Sippl MJ (1993) Recognition of errors in three-dimensional structures of proteins. Proteins 17:355–362
Wallner B, Elofsson A (2003) Can correct protein models be identified? Protein Sci 12:1073–1086
Jorgensen WL, Maxwell DS, Tirado-Rives J (1996) Development and testing of the OPLS all-atom force field on conformational energetics and properties of organic liquids. J Am Chem Soc 118:11225–11236
Kaminski GA, Friesner RA, Tirado-Rives J, Jorgensen WL (2001) Evaluation and reparametrization of the OPLS-AA force field for proteins via comparison with accurate quantum chemical calculations on peptides. J Phys Chem B 105:6474–6487
Jorgensen WL, Madura JD (1985) Temperature and size dependence for Monte Carlo simulations of TIP4P water. Mol Phys 56:1381–1392
Darden T, York D, Pedersen L (1993) Particle mesh Ewald: an N center-dot. log(N) method for Ewald sums in large systems. J Chem Phys 98:10089–10092
Tuckerman M, Berne BJ, Martyna GJ (1992) Reversible multiple time scale molecular dynamics. J Chem Phys 97:1990–2001
Oh SM, Zhu F, Cho YY, Lee KW, Kang BS, Kim H-G, Zykova T, Bode AM, Dong Z (2007) T-lymphokine activated killer cell originated protein kinase functions as a positive regulator of c-Jun-NH2-kinase 1 signaling and H-Ras-induced cell transformation. Cancer Res 67:5186–5194
Jørgensen AM, Topiol S (2008) Driving forces for ligand migration in the leucine transporter. Chem Biol Drug Des 72:265–272
Singh K, Kirubakaran P, Nagarajan S, Sakkiah S, Muthusamy K, Velmurgan D, Jeyakanthan J (2012) Homology modeling, molecular dynamics, e-pharmacophore mapping and docking study of Chikungunya virus nsP2 protease. J Mol Model 18:39–51
Baker NA, Sept D, Joseph S, Holst MJ, McCammon JA (2001) Electrostatics of nanosystems: application to microtubules and the ribosome. Proc Natl Acad Sci USA 98:10037–10041
Dolinsky TJ, Nielsen JE, McCammon JA, Baker NA (2004) PDB2PQR: an automated pipeline for the setup of Poisson-Boltzmann electrostatics calculations. Nucleic Acids Res 32:665–667
Sitkoff D, Sharp KA, Honig B (1994) Accurate calculation of hydration free energies using macroscopic solvent models. J Phys Chem 98:1978–1988
Glasser L, Herraez A, Hanson RM (2009) Interactive 3D phase diagrams using Jmol. J Chem Educ 86:566–null
Friesner RA, Murphy RB, Repasky MP, Frye LL, Greenwood JR, Halgren TA, Sanschagrin PC, Mainz DT (2006) Extra precision glide: docking and scoring incorporating a model of hydrophobic enclosure for protein - ligand complexes. J Med Chem 49:6177–6196
Kozakov D, Hall DR, Beglov D, Brenke R, Comeau SR, Shen Y, Li K, Zheng J, Vakili P, Paschalidis IC, Vajda S (2010) Achieving reliability and high accuracy in automated protein docking: Cluspro, PIPER, SDU, and stability analysis in CAPRI rounds 13–19. Proteins 78(15):3124–3130
Kozakov D, Brenke R, Comeau SR, Vajda S (2006) PIPER: an FFT-based protein docking program with pairwise potentials. Proteins 65(2):392–406
Comeau SR, Gatchell DW, Vajda S, Camacho CJ (2004) ClusPro: an automated docking and discrimination method for the prediction of protein complexes. Bioinformatics 20(1):45–50
Comeau SR, Gatchell DW, Vajda S, Camacho CJ (2004) ClusPro: a fully automated algorithm for protein-protein docking. Nucleic Acids Res 32(suppl 2):W96–W99
Schrödinger LLC (2011) QikProp, version 3.4. Schrödinger, LLC, New York, NY
Poroikov VV, Filimonov DA, Ihlenfeldt W-D, Gloriozova TA, Lagunin AA, Borodina YV, Stepanchikova AV, Nicklaus MC (2002) PASS biological activity spectrum predictions in the enhanced open NCI database browser. J Chem Inf Comput Sci 43(1):228–236
Larsson P, Wallner B, Lindahl E, Elofsson A (2008) Using multiple templates to improve quality of homology models in automated homology modeling. Protein Sci 17:990–1002
Venclovas C (2003) Comparative modeling in CASP5: progress is evident, but alignment errors remain a significant hindrance. Proteins 53:380–388
Mobarec JC, Sanchez R, Filizola M (2009) Modern homology modeling of G-protein coupled receptors: which structural template to use? J Med Chem 52:5207–5216
Kirubakaran P, Muthusamy K, Dhanachandra Singh K, Nagamani S (2012) Homology modeling, molecular dynamics, and molecular docking studies of Trichomonas vaginalis carbamate kinase. Med Chem Res 21:2105–2116
Curman D, Cinel B, Williams DE, Rundle N, Block WD, Goodarzi AA, Hutchins JR, Clarke PR, Zhou B-B, Lees-Miller SP, Andersen RJ, Roberge M (2001) Inhibition of the G2 DNA damage checkpoint and of protein kinases Chk1 and Chk2 by the marine sponge alkaloid debromohymenialdisine. J Biol Chem 276(21):17914–17919
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
ESM 1
Supplementary Fig. 1: Ramachandran plot for TOPK protein. Supplementary Fig. 2: Protein charge of folded nature and unfolded state and the stability of the protein. Supplementary Fig. 3: Active site region of TOPK protein predicted by Sitemap in Schrodinger 9.1. Supplementary Fig. 4: Root mean square fluctuation (RMSF) of the backbone atoms averaged over each residue for the active site regions are shown in red circle. (DOC 1757 kb)
Rights and permissions
About this article
Cite this article
Kirubakaran, P., Karthikeyan, M., Singh, K.D. et al. In silico structural and functional analysis of the human TOPK protein by structure modeling and molecular dynamics studies. J Mol Model 19, 407–419 (2013). https://doi.org/10.1007/s00894-012-1566-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00894-012-1566-1