Abstract
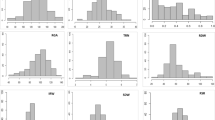
Two hundred one hexaploid wheat accessions, representing 200 years of selection and breeding history, were sampled from the National Small Grains Collection in Aberdeen, ID, and evaluated for five root traits at the seedling stage. A paper roll-supported hydroponic system was used for seedling growth. Replicated roots samples were analyzed by WinRHIZO. We observed accessions with nearly no branching and accessions with up to 132 cm of branching. Total seminal root length ranged from 70 to 248 cm, a 3.5-fold difference. Next-generation sequencing was used to produce single-nucleotide polymorphism (SNP) markers and genomic libraries that were aligned to the wheat reference genome IWGSCv1 and were called single-nucleotide polymorphism (SNP) markers. After filtering and imputation, a total of 20,881 polymorphic sites were used to perform association mapping in TASSEL. Gene annotations were conducted for identified marker-trait associations (MTAs) with − log10P > 3.5 (p value < 0.003). In total, we identified 63 MTAs with seven for seminal axis root length (SAR), 24 for branching (BR), four for total seminal root length (TSR), eight for root dry matter (RDM), and 20 for root diameter (RD). Putative proteins of interest that we identified include chalcone synthase, aquaporin, and chymotrypsin inhibitor for SAR, MYB transcription factor and peroxidase for BR, zinc fingers and amino acid transporters for RDM, and cinnamoyl-CoA reductase for RD. We evaluated the effects of height-reducing Rht alleles and the 1B/1R translocation event on root traits and found presence of the Rht-B1b allele decreased RDM, while presence of the Rht-D1b allele increased TSR and decreased RD.





Similar content being viewed by others
References
Abramovitch RB, Janjusevic R, Stebbins CE, Martin GB (2006) Type III effector AvrPtoB requires intrinsic E3 ubiquitin ligase activity to suppress plant cell death and immunity. Proc Natl Acad Sci 103:2851–2856. https://doi.org/10.1073/pnas.0507892103
Atkinson JA, Wingen LU, Griffiths M et al (2015) Phenotyping pipeline reveals major seedling root growth QTL in hexaploid wheat. J Exp Bot 66:2283–2292. https://doi.org/10.1093/jxb/erv006
Atta BM, Mahmood T, Trethowan RM (2013) Relationship between root morphology and grain yield of wheat in north-western NSW, Australia. Aust J Crop Sci 7:2108–2115
Bai C, Liang Y, Hawkesford MJ (2013) Identification of QTLs associated with seedling root traits and their correlation with plant height in wheat. J Exp Bot 64:1745–1753. https://doi.org/10.1093/jxb/ert041
Bates D, Maechler M, Bolker B, Walker S (2015) Fitting linear mixed-effects models using lme4. J Stat Softw 67:1–48. https://doi.org/10.18637/jss.v067.i01
Benfey PN, Linstead PJ, Roberts K et al (1993) Root development in Arabidopsis: four mutants with dramatically altered root morphogenesis. Development 119:57–70
Benson J, Brown-Guedira G, Paul Murphy J, Sneller C (2012) Population structure, linkage disequilibrium, and genetic diversity in soft winter wheat enriched for fusarium head blight resistance. Plant Genome 5:71–80. https://doi.org/10.3835/plantgenome2011.11.0027
Bradbury PJ, Zhang Z, Kroon DE et al (2007) TASSEL: software for association mapping of complex traits in diverse samples. Bioinformatics 23:2633–2635. https://doi.org/10.1093/bioinformatics/btm308
Cabrera A, Souza E, Guttieri M et al (2014) Genetic diversity, linkage disequilibrium, and genome evolution in soft winter wheat. Crop Sci 54:2433–2448. https://doi.org/10.2135/cropsci2013.09.0601
Csiszár J, Gallé Á, Horváth E et al (2012) Different peroxidase activities and expression of abiotic stress-related peroxidases in apical root segments of wheat genotypes with different drought stress tolerance under osmotic stress. Plant Physiol Biochem 52:119–129. https://doi.org/10.1016/j.plaphy.2011.12.006
Csiszár J, Lantos E, Tari I et al (2007) Antioxidant enzyme activities in Allium species and their cultivars under water stress. Plant Soil Environ 53:517–523
Dao TTH, Linthorst HJM, Verpoorte R (2011) Chalcone synthase and its functions in plant resistance. Phytochem Rev 10:397–412. https://doi.org/10.1007/s11101-011-9211-7
de Dorlodot S, Forster B, Pagès L et al (2007) Root system architecture: opportunities and constraints for genetic improvement of crops. Trends Plant Sci 12:474–481
Eastmond PJ (2006) SUGAR-DEPENDENT1 encodes a patatin domain triacylglycerol lipase that initiates storage oil breakdown in germinating Arabidopsis seeds. Plant Cell 18:665–675. https://doi.org/10.1105/tpc.105.040543
Hamada A, Nitta M, Nasuda S et al (2012) Novel QTLs for growth angle of seminal roots in wheat (Triticum aestivum L.). Plant Soil 354:395–405. https://doi.org/10.1007/s11104-011-1075-5
Han Y, Zhao W, Wang Z et al (2014) Molecular evolution and sequence divergence of plant chalcone synthase and chalcone synthase-like genes. Genetica 142:215–225. https://doi.org/10.1007/s10709-014-9768-3
Hill WG, Weir BS (1988) Variances and covariances of squared linkage disequilibria in finite populations. Theor Popul Biol 33:54–78. https://doi.org/10.1016/0040-5809(88)90004-4
Jeong R-D, Chandra-Shekara AC, Barman SR et al (2010) Cryptochrome 2 and phototropin 2 regulate resistance protein-mediated viral defense by negatively regulating an E3 ubiquitin ligase. Proc Natl Acad Sci 107:13538–13543. https://doi.org/10.1073/pnas.1004529107
Kabir M, Liu G, Guan P et al (2015) Mapping QTLs associated with root traits using two different populations in wheat (Triticum aestivum L.). Euphytica 206:175–191. https://doi.org/10.1007/s10681-015-1495-z
Kang HM, Zaitlen NA, Wade CM et al (2008) Efficient control of population structure in model organism association mapping. Genetics 178:1709–1723. https://doi.org/10.1534/genetics.107.080101
Lachowiec J, Shen X, Queitsch C, Carlborg Ö (2015) A genome-wide association analysis reveals epistatic cancellation of additive genetic variance for root length in Arabidopsis thaliana. PLoS Genet 11:1–22. https://doi.org/10.1371/journal.pgen.1005541
Laperche A, Devienne-Barret F, Maury O et al (2006) A simplified conceptual model of carbon/nitrogen functioning for QTL analysis of winter wheat adaptation to nitrogen deficiency. Theor Appl Genet 113:1131–1146. https://doi.org/10.1007/s00122-006-0373-4
Li H, Durbin R (2009) Fast and accurate short read alignment with Burrows-Wheeler transform. Bioinformatics 25:1754–1760. https://doi.org/10.1093/bioinformatics/btp324
Li P, Chen J, Wu P et al (2011) Quantitative trait loci analysis for the effect of Rht-B1 dwarfing gene on coleoptile length and seedling root length and number of bread wheat. Crop Sci 51:2561–2568. https://doi.org/10.2135/cropsci2011.03.0116
Li X, Guo Z, Lv Y et al (2017) Genetic control of the root system in rice under normal and drought stress conditions by genome-wide association study. PLoS Genet 13:e1006889. https://doi.org/10.1371/journal.pgen.1006889
Li S, Ma J, Liu P (2013) OPR3 is expressed in phloem cells and is vital for lateral root development in Arabidopsis. Can J Plant Sci 93:165–170. https://doi.org/10.4141/CJPS2012-149
Ling Q, Jarvis P (2015) Regulation of chloroplast protein import by the ubiquitin E3 ligase SP1 is important for stress tolerance in plants. Curr Biol 25:2527–2534. https://doi.org/10.1016/j.cub.2015.08.015
Liu M, Rathjen T, Weligama K et al (2017) Analysis of aneuploid lines of bread wheat to map chromosomal locations of genes controlling root hair length. Ann Bot 119:1333–1341. https://doi.org/10.1093/aob/mcx030
Lupton FGH, Oliver RH, Ellis FB et al (1974) Root and shoot growth of semi-dwarf and taller winter wheats. Ann Appl Biol 77:129–144. https://doi.org/10.1111/j.1744-7348.1974.tb06881.x
Maccaferri M, El-Feki W, Nazemi G et al (2016) Prioritizing quantitative trait loci for root system architecture in tetraploid wheat. J Exp Bot 67:1161–1178. https://doi.org/10.1093/jxb/erw039
Man J, Shi Y, Yu Z, Zhang Y (2015) Dry matter production, photosynthesis of flag leaves and water use in winter wheat are affected by supplemental irrigation in the Huang-Huai-Hai plain of China. PLoS One 10:1–19. https://doi.org/10.1371/journal.pone.0137274
Miller AJ, Fan X, Orsel M et al (2007) Nitrate transport and signalling. J Exp Bot 58:2297–2306. https://doi.org/10.1093/jxb/erm066
Money D, Gardner K, Migicovsky Z et al (2015) LinkImpute: fast and accurate genotype imputation for nonmodel organisms. G3 (Bethesda) 5:2383–2390. https://doi.org/10.1534/g3.115.021667
Nguyen MX, Moon S, Jung KH (2013) Genome-wide expression analysis of rice aquaporin genes and development of a functional gene network mediated by aquaporin expression in roots. Planta 238:669–681. https://doi.org/10.1007/s00425-013-1918-9
Norton GJ, Aitkenhead MJ, Khowaja FS et al (2008) A bioinformatic and transcriptomic approach to identifying positional candidate genes without fine mapping: an example using rice root-growth QTLs. Genomics 92:344–352. https://doi.org/10.1016/j.ygeno.2008.07.002
Pan R, Satkovich J, Hu J (2016) E3 ubiquitin ligase SP1 regulates peroxisome biogenesis in Arabidopsis. Proc Natl Acad Sci 113:E7307–E7316. https://doi.org/10.1073/pnas.1613530113
Pang J, Milroy SP, Rebetzke GJ, Palta JA (2015) The influence of shoot and root size on nitrogen uptake in wheat is affected by nitrate affinity in the roots during early growth. Funct Plant Biol 42:1179–1189. https://doi.org/10.1071/FP15215
Passardi F, Tognolli M, De Meyer M et al (2006) Two cell wall associated peroxidases from Arabidopsis influence root elongation. Planta 223:965–974. https://doi.org/10.1007/s00425-005-0153-4
Petrarulo M, Marone D, Ferragonio P et al (2015) Genetic analysis of root morphological traits in wheat. Mol Gen Genomics 290:785–806. https://doi.org/10.1007/s00438-014-0957-7
Poland JA, Brown PJ, Sorrells ME, Jannink JL (2012) Development of high-density genetic maps for barley and wheat using a novel two-enzyme genotyping-by-sequencing approach. PLoS One 7:e32253. https://doi.org/10.1371/journal.pone.0032253
Postma JA, Dathe A, Lynch JP (2014) The optimal lateral root branching density for maize depends on nitrogen and phosphorus availability. Plant Physiol 166:590–602. https://doi.org/10.1104/pp.113.233916
Pratelli R, Guerra DD, Yu S et al (2012) The ubiquitin E3 ligase LOSS OF GDU2 is required for GLUTAMINE DUMPER1-induced amino acid secretion in Arabidopsis. Plant Physiol 158:1628–1642. https://doi.org/10.1104/pp.111.191965
Pritchard JK, Stephens M, Donnelly P (2000) Inference of population structure using multilocus genotype data. Genetics 155:945–959. https://doi.org/10.1111/j.1471-8286.2007.01758.x
R Development Core Team (2013) R: A language and environment for statistical computing. R Found. Stat. Comput. Vienna Austria 0:{ISBN} 3-900051-07-0
Rahnama A, Munns R, Poustini K, Watt M (2011) A screening method to identify genetic variation in root growth response to a salinity gradient. J Exp Bot 62:69–77. https://doi.org/10.1093/jxb/erq359
Rasheed A, Wen W, Gao F et al (2016) Development and validation of KASP assays for genes underpinning key economic traits in bread wheat. Theor Appl Genet 129:1843–1860. https://doi.org/10.1007/s00122-016-2743-x
Ren Y, He Z, Li J et al (2012) QTL mapping of adult-plant resistance to stripe rust in a population derived from common wheat cultivars Naxos and Shanghai 3/Catbird. Theor Appl Genet 125:1211–1221. https://doi.org/10.1007/s00122-012-1907-6
Richards RA, Passioura JB (1981) Seminal root morphology and water use of wheat I. Environmental effects. Crop Sci 2:253–255. https://doi.org/10.2135/cropsci1981.0011183X002100020011x
Rosas U, Cibrian-Jaramillo A, Ristova D et al (2013) Integration of responses within and across Arabidopsis natural accessions uncovers loci controlling root systems architecture. Proc Natl Acad Sci 110:15133–15138. https://doi.org/10.1073/pnas.1305883110
Scheres B, Dilaurenzio L, Willemsen V et al (1995) Mutations affecting the radial organisation of the Arabidopsis root display specific defects throughout the embryonic axis. Development 121:53–62. https://doi.org/10.1101/gad.1426606.rates
Subira J, Ammar K, Álvaro F et al (2016) Changes in durum wheat root and aerial biomass caused by the introduction of the Rht-B1b dwarfing allele and their effects on yield formation. Plant Soil 403:291–304. https://doi.org/10.1007/s11104-015-2781-1
Uga Y, Sugimoto K, Ogawa S et al (2013) Control of root system architecture by DEEPER ROOTING 1 increases rice yield under drought conditions. Nat Genet 45:1097–1102. https://doi.org/10.1038/ng.2725
Waines JG, Ehdaie B (2007) Domestication and crop physiology: roots of green-revolution wheat. Ann Bot 100:991–998
Wang Y, Chen L, Du Y et al (2014) Genetic effect of dwarfing gene Rht13 compared with Rht-D1b on plant height and some agronomic traits in common wheat (Triticum aestivum L.). F Crop Res 162:39–47. https://doi.org/10.1016/j.fcr.2014.03.014
Wojciechowski T, Gooding MJ, Ramsay L, Gregory PJ (2009) The effects of dwarfing genes on seedling root growth of wheat. J Exp Bot 60:2565–2573. https://doi.org/10.1093/jxb/erp107
Xie Q, Kurukulasuriya F, Mayes S, Sparkes D (2017) Identifying seedling root architectural traits associated with yield and yield components in wheat. Ann Bot 119:1115–1130. https://doi.org/10.1093/aob/mcx001
You Q, Zhai K, Yang D et al (2016) An E3 ubiquitin ligase-BAG protein module controls plant innate immunity and broad-spectrum disease resistance. Cell Host Microbe 20:758–769. https://doi.org/10.1016/j.chom.2016.10.023
Zhan A, Schneider H, Lynch JP (2015) Reduced Lateral Root Branching Density Improves Drought Tolerance in Maize. Plant Physiol 168:1603–1615
Zheng H, Pan X, Deng Y et al (2016) AtOPR3 specifically inhibits primary root growth in Arabidopsis under phosphate deficiency. Sci Rep 6:24778. https://doi.org/10.1038/srep24778
Zheng BS, Yang L, Zhang WP et al (2003) Mapping QTLs and candidate genes for rice root traits under different water-supply conditions and comparative analysis across three populations. Theor Appl Genet 107:1505–1515. https://doi.org/10.1007/s00122-003-1390-1
Funding
This study received financial support from USDA Hatch grant 1013073 and Purdue College of Agriculture.
Author information
Authors and Affiliations
Consortia
Corresponding author
Rights and permissions
About this article
Cite this article
Beyer, S., Daba, S., Tyagi, P. et al. Loci and candidate genes controlling root traits in wheat seedlings—a wheat root GWAS. Funct Integr Genomics 19, 91–107 (2019). https://doi.org/10.1007/s10142-018-0630-z
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10142-018-0630-z