Abstract
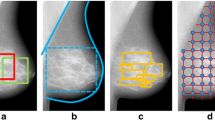
Purpose: The purpose of the study was to evaluate the usefulness of power law spectral analysis on mammographic parenchymal patterns in breast cancer risk assessment. Materials and Methods: Mammograms from 172 subjects (30 women with the BRCA1/BRCA2 gene mutation and 142 low-risk women) were retrospectively collected and digitized. Because age is a very important risk factor, 60 low-risk women were randomly selected from the 142 low-risk subjects and were age matched to the 30 gene mutation carriers. Regions of interest were manually selected from the central breast region behind the nipple of these digitized mammograms and subsequently used in power spectral analysis. The power law spectrum of the form \(P\left( f \right) = {B \mathord{\left/ {\vphantom {B {f^\beta }}} \right. \kern-\nulldelimiterspace} {f^\beta }}\) was evaluated for the mammographic patterns. The performance of exponent β as a decision variable for differentiating between gene mutation carriers and low-risk women was assessed using receiver operating characteristic analysis for both the entire database and the age-matched subset. Results: Power spectral analysis of mammograms demonstrated a statistically significant difference between the 30 BRCA1/BRCA2 gene mutation carriers and the 142 low risk women with an average β values of 2.92 (±0.28) and 2.47(±0.20), respectively. An A z value of 0.90 was achieved in distinguishing between gene mutation carriers and low-risk women in the entire database, with an A z value of 0.89 being achieved on the age-matched subset. Conclusions: The BRCA1/BRCA2 gene mutation carriers and low-risk women have different mammographic parenchymal patterns. It is expected that women identified as high risk by computerized feature analyses might potentially be more aggressively screened for breast cancer.







Similar content being viewed by others
References
Carter CL, Allen C, Henson DE: Relation of tumor size, lymph node status, and survival in 24,740 breast cancer cases. Cancer 63:181–189, 1989
Clay MG, Hishop G, Kan L, Olivotto IA, Burhenne LJ: Screening mammography in British Columbia 1988–1993. Am J Surg 167:490–492, 1994
Jemal A, Siegel R, Ward E, Murray T, Xu J, Thun MJ: Cancer statistics, 2007. CA Cancer J Clin 57:43–66, 2007
Singletary SE: Rating the risk factors for breast cancer. Ann Surg 237:474–482, 2003
Heine JJ, Malhotra P: Mammographic tissue, breast cancer risk, serial image analysis, and digital mammography. Acad Radiol 9:298–316, 2002
Stoutjesdijk MJ, Boetes C, Jager GJ, Beex L, Bult P, Hendriks J, Laheij R, Massuger L, van Die LE, Wobbes T, Barentsz JO: Magnetic resonance imaging and mammography in women with a hereditary risk of breast cancer. J Natl Cancer Inst 93:1095–1102, 2001
Warner E, Plewes DB, Shumak RS, Catzavelos GC, Di Prospero LS, Yaffe MJ, Goel V, Ramsay E, Chart PL, Cole DEC, Taylor GA, Cutrara M, Samuels TH, Murphy JP, Murphy JM, Narod SA: Comparison of breast magnetic resonance imaging, mammography, and ultrasound for surveillance of women at high risk for hereditary breast cancer. J Clin Oncol 19:3524–3531, 2001
Euhus DM, Smith KC, Robinson L, Stucky A, Olopade OI, Cummings S, Garber JE, Chittenden A, Mills GB, Rieger P, Esserman L, Crawford B, Hughes KS, Roche CA, Ganz PA, Seldon J, Fabian CJ, Klemp J, Tomlinson G: Pretest prediction of BRCA1 or BRCA2 mutation by risk counselors and the computer model BRCAPRO. J Natl Cancer Inst 94:844–851, 2002
Thompson D, Easton DF: Cancer incidence in BRCA1 mutation carriers. J Natl Cancer Inst 94:1358–1365, 2002
Wolfe JN: Breast patterns as an index of risk for developing breast cancer. Am J Roentgenol 126:1130–1139, 1976
Boyd NF, O’Sullivan B, Fishell E, Simor I, Cooke G: Mammographic patterns and breast cancer risk: methodologic standards and contradictory results. J Natl Cancer Inst 72:1253–1259, 1984
Boyd NF, Martin LJ, Stone J, Greenberg C, Minkin S, Yaffe MJ: Mammographic densities as a marker of human breast cancer risk and their use in chemoprevention. Curr Oncol Rep 3:314–321, 2001
Brisson J, Diorio C, Mâsse B: Wolfe’s parenchymal pattern and percentage of the breast with mammographic densities: redundant or complementary classifications? Caner Epidemiol Biomarkers Prev 12:728–732, 2003
Tahoces PG, Correa J, Souto M, Gomez L, Vidal JJ: Computer-assisted diagnosis: the classification of mammographic breast parenchymal patterns. Phys Med Biol 40:103–117, 1995
Magnin IE, Cluzeau F, Odet CL: Mammographic texture analysis: an evaluation of risk for developing breast cancer. Opt Eng 25:780–784, 1986
Taylor P, Hajnal S, Dilhuydy MH, Barreau B: Measuring image texture to separate “difficult” from “easy” mammograms. Br J Radiol 67:456–463, 1994
Byng JW, Yaffe MJ, Lockwood LE, Little LE, Tritchler DL, Boyd NF: Automated analysis of mammographic densities and breast carcinoma risk. Cancer 80:66–74, 1997
Boyd NF, Lockwood GA, Martin LJ, Knight JA, Byng JW, Yaffe MJ, Tritchler DL: Mammographic densities and breast cancer risk. Breast Dis 10:113–126, 1998
Boyd NF, Martin LJ, Stone J, Greenberg C, Minkin S, Yaffe MJ: Mammographic densities as a marker of human breast cancer risk and their use in chemoprevention. Cancer Prev 3:314–321, 2001
Atkinson C, Warren R, Bingham SA, Day NE: Mammographic patterns as a predictive biomarker of breast cancer risk: effect of tamoxifen. Cancer Epidemiol Biomarkers Prev 8:863–866, 1999
Huo Z, Giger ML, Wolverton DE, Zhong W, Cumming SA, Olopade OI: Computerized analysis of mammographic parenchymal patterns for breast cancer risk assessment: feature selection. Med Phys 27:4–12, 2000
Huo Z, Giger ML, Olopade OI, Wolveton DE, Weber BL, Metz CE, Zhong W, Cummings SA: Computerized analysis of digitized mammograms of BRCA1 and BRCA2 gene mutation carriers. Radiology 225:519–526, 2002
Li H, Giger ML, Huo Z, Olopade OI, Lan L, Weber BL, Bonta I: Computerized analysis of mammographic parenchymal patterns for assessing breast cancer risk: effect of ROI size and location. Med Phys 31:549–555, 2004
Li H, Giger ML, Olopade OI, Margolis A, Lan L, Chinander MR: Computerized texture analysis of mammographic parenchymal patterns of digitized mammograms. Acad Radiol 12:863–873, 2005
Li H, Giger ML, Olopade OI, Lan L: Fractal analysis of mammographic parenchymal patterns in breast cancer risk assessment. Acad Radiol 14:513–521, 2007
Burgess AE: Mammographic structure: data preparation and spatial statistics analysis. Medical Imaging 1998, Image Processing, San Diego, CA. In: Hanson K Ed. Proceedings of the Society of photo-optics Instrumentation Engineers, Bellingham, WA, vol. 3661, 1998, pp 642–653
Heine JJ, Velthuizen RP: Spectral analysis of full field digital mammography data. Med Phys 29:647–661, 2002
Metz CE: ROC methodology in radiologic imaging. Invest Radiol 21:720–733, 1986
Metz CE: Some practical issues of experimental design and data analysis in radiological ROC studies. Invest Radiol 24:234–245, 1989
Gail MH, Brinton LA, Byar DP, Corle DK, Green SB, Schairer C, Mulvihill JJ: Projecting individualized probabilities of developing breast cancer for white females who are being examined annually. J Natl Cancer Inst 81:1879–1886, 1989
Sonka M, Hlavac V, Boyle R: Image processing, analysis, and machine vision, Pacific Grove, CA: PWS, 1999
Burgess AE, Jacobson FL, Judy PF: Human observer detection experiments with mammograms and power-law noise. Med Phys 28:419–437, 2001
Bendat JS, Piersol AG: Random data: analysis and measurement procedures, New York: Wiley, 2000
Rice JA: Mathematical statistics and data analysis, Belmont, CA: Duxbury, 1995
Heine JJ, Velthuizen RP: Spectral analysis of full field digital mammography data. Med Phys 29:647–661, 2002
Eckstein MP, Whiting JS: Visual signal detection in structured backgrounds: I. Effect of number of possible spatial locations and signal contrast. J Opt Soc Am A 13:1777–1787, 1996
Soille P, Rivest JF: On the validity of fractal dimension measurements in image analysis. J Visual Commun Image Represent 7:217–229, 1996
Acknowledgments
We would like to thank Zhimin Huo, Ph.D., for initial studies on the database, Arthur E. Burgess, Ph.D., for useful discussions, Barbara L. Weber, M.D., for contributing cases to the database, and Dulcy E. Wolverton, M.D., for reviewing the mammograms. This work was supported in parts by USPHS grants R01-CA89452, R21-CA113800, and P50-CA125183 and by a grant from the US Army Medical Research and Materiel Command (DAMD 98-1209). M. L. Giger is a shareholder in R2/Hologic (Sunnyvale, CA). It is the policy of the University of Chicago that investigators disclose publicly actual or potential significant financial interests that may appear to be affected by the research activities.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Li, H., Giger, M.L., Olopade, O.I. et al. Power Spectral Analysis of Mammographic Parenchymal Patterns for Breast Cancer Risk Assessment. J Digit Imaging 21, 145–152 (2008). https://doi.org/10.1007/s10278-007-9093-9
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10278-007-9093-9