Abstract
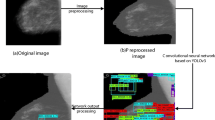
The diagnosis of breast cancer in early stage is essential for successful treatment. Detection can be performed in several ways, the most common being through mammograms. The projections acquired by this type of examination are directly affected by the composition of the breast, which density can be similar to the suspicious masses, being a challenge the identification of malignant lesions. In this article, we propose a computer-aided detection (CAD) system to aid in the diagnosis of masses in digitized mammograms using a model based in the U-Net, allowing specialists to monitor the lesion over time. Unlike most of the studies, we propose the use of an entire base of digitized mammograms using normal, benign, and malignant cases. Our research is divided into four stages: (1) pre-processing, with the removal of irrelevant information, enhancement of the contrast of 7989 images of the Digital Database for Screening Mammography (DDSM), and obtaining regions of interest. (2) Data augmentation, with horizontal mirroring, zooming, and resizing of images; (3) training, with tests of six-based U-Net models, with different characteristics; (4) testing, evaluating four metrics, accuracy, sensitivity, specificity, and Dice Index. The tested models obtained different results regarding the assessed parameters. The best model achieved a sensitivity of 92.32%, specificity of 80.47%, accuracy of 85.95% Dice Index of 79.39%, and AUC of 86.40%. Even using a full base without case selection bias, the results obtained demonstrate that the use of a complete database can provide knowledge to the CAD expert.










Similar content being viewed by others
Notes
http://www.eng.usf.edu/cvprg/Mammogra- phy/Database.html
http://medicalresearch.inescporto.pt/breastresearch/in- dex.php/Get_INbreast_Database
References
Al-antari MA et al.: A fully integrated computer-aided diagnosis system for digital x-ray mammograms via deep learning detection, segmentation, and classification. International Journal of Medical Informatics 117:44–54, 2018
Al-masni MA et al.: Simultaneous detection and classification of breast masses in digital mammograms via a deep learning yolo-based cad system. Computer Methods and Programs in Biomedicine 157:85–94, 2018
Bray F et al.: Global cancer statistics 2018: Globocan estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA: A Cancer Journal for Clinicians, 2018. https://doi.org/10.3322/caac.21492
Caldas FAA et al.: Controle de qualidade e artefatos em mamografia. Revista Radiologia Brasileira 38(4):295–300, 2005
Chakraborty J, Midya A, Rabidas R: Computer-aided detection and diagnosis of mammographic masses using multi-resolution analysis of oriented tissue patterns. Expert Systems with Applications 99:168–179, 2018. https://doi.org/10.1016/j.eswa.2018.01.010
Dhungel N, Carneiro G, Bradley AP: A deep learning approach for the analysis of masses in mammograms with minimal user intervention. Medical Image Analysis 37:114–128, 2017
Dhungel N et al.: Deep learning and structured prediction for the segmentation of mass in mammograms. In: Medical Image Computing and Computer-Assisted Intervention – MICCAI 2015. Springer International Pub- lishing, 2015, pp. 605–612
Diniz B et al.: Detection of mass regions in mammograms by bilateral analysis adapted to breast density using similarity indexes and convolutional neural networks. Computer Methods and Programs in Biomedicine 156:191–207, 2018. https://doi.org/10.1016/j.cmpb.2018.01.007
Dromain C et al.: Computed-aided diagnosis (CAD) in the detection of breast cancer. European Journal of Radi- ology 82(3):417–423, 2013
Felzenszwalb PF, Huttenlocher DP: Efficient graph- based image segmentation. International Journal of Computer Vision 59(2):167–181, 2004
Heath, M., et al.: Digital database for screening mammography (2001). URL http://marathon.csee.usf.edu/Mammography/Database.html
Li H, Zhuang S, Li D, Zhao J, Ma Y: Benign and malignant classification of mammogram images based on deep learning. Biomedical Signal Processing and Control 51:347–354, 2019. https://doi.org/10.1016/j.bspc.2019.02.017
de Nazaré Silva, J., de Carvalho Filho, A.O., Corrêa Silva, A., Cardoso de Paiva, A., Gattass, M.: Automatic detection of masses in mammograms using quality threshold clustering, correlogram function, and svm. Journal of Digital Imaging 28(3), 323–337 (2015). DOI https://doi.org/10.1007/s10278-014-9739-3
Pedro RWD, Machado-Lima A, Nunes FL: Is mass classification in mammograms a solved problem? - a critical review over the last 20 years. Expert Systems with Applications 119:90–103, 2019. https://doi.org/10.1016/j.eswa.2018.10.032
Ronneberger O, Fischer P, Brox T: U-net: Convolu- tional networks for biomedical image segmentation. In: Medical Image Computing and Computer-Assisted Intervention - MICCAI 2015. Cham: Springer International Publishing, 2015, pp. 234–241
de Sampaio WB, Silva AC, de Paiva AC, Gattass M: Detection of masses in mammograms with adaption to breast density using genetic algorithm, phylogenetic trees, lbp and svm. Expert Systems with Applications 42(22):8911–8928, 2015
Sharma S, Khanna P: Computer-aided diagnosis of malignant mammograms using zernike moments and svm. Journal of Digital Imaging 28(1):77–90, 2015
Zhang X et al.: Whole mammogram image classification with convolutional neural networks. In: 2017 IEEE International Conference on Bioinformatics and Biomedicine (BIBM), 2017, pp. 700–704
Zuiderveld, K.: Graphics gems iv. In: P.S. Heckbert (ed.) Graphics gems, chap. Contrast limited adaptive histogram equalization, pp. 474–485. Academic Press Professional, Inc., San Diego (1994)
Author information
Authors and Affiliations
Corresponding author
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Zeiser, F.A., da Costa, C.A., Zonta, T. et al. Segmentation of Masses on Mammograms Using Data Augmentation and Deep Learning. J Digit Imaging 33, 858–868 (2020). https://doi.org/10.1007/s10278-020-00330-4
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10278-020-00330-4