Abstract
Metastasis is a complex series of sequential events involving several gene products and the regulated expression of several tumor cell genes. Using rat mammary adenocarcinoma cell lines of differing metastatic potentials and a differential complementary DNA (cDNA) hybridization method, our laboratory embarked in 1992 on a project to identify candidate metastasis-associated genes. Among the genes that were found to be abundantly overexpressed in highly metastatic rat cell lines compared to poorly metastatic cell lines, we identified a completely novel gene without any homologous or related genes in the database in 1994. The full-length cDNA of this gene was cloned, sequenced, and named mta1 (metastasis-associated gene 1), and eventually, its human cDNA counterpart, MTA1, was also cloned and sequenced by our group. MTA1 has now been identified as one of the members of a gene family (MTA gene family). The products of the MTA genes, the MTA proteins, are transcriptional co-regulators that function in histone deacetylation and nucleosome remodeling. In this review, we will briefly discuss the researches for the identification and characterization of the mta1 gene, its human counterpart MTA1, and their protein products.


Similar content being viewed by others
References
Liotta, L. A. (1986). Tumor invasion and metastases—role of the extracellular matrix: Rhoads memorial award lecture. Cancer Research, 46(1), 1–7.
Nicolson, G. L. (1988). Cancer metastasis: tumor cell and host organ properties important in metastasis to specific secondary sites. Biochimica et Biophysica Acta, 948(2), 175–224.
Chambers, A. F., & Tuck, A. B. (1993). Ras-responsive genes and tumor metastasis. Critical Review of Oncogenesis, 4(2), 95–114.
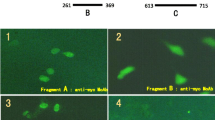
Toh, Y., Pencil, S. D., & Nicolson, G. L. (1994). A novel candidate metastasis-associated gene, mta1, differentially expressed in highly metastatic mammary adenocarcinoma cell lines. cDNA cloning, expression, and protein analyses. Journal of Biological Chemistry, 269(37), 22958–22963.
Nawa, A., Nishimori, K., Lin, P., Maki, Y., Moue, K., Sawada, H., et al. (2000). Tumor metastasis-associated human MTA1 gene: its deduced protein sequence, localization, and association with breast cancer cell proliferation using antisense phosphorothioate oligonucleotides. Journal of Cell Biochemistry, 79(2), 202–212.
Toh, Y., Oki, E., Oda, S., Tokunaga, E., Ohno, S., Maehara, Y., et al. (1997). Overexpression of the MTA1 gene in gastrointestinal carcinomas: correlation with invasion and metastasis. International Journal of Cancer, 74(4), 459–463.
Toh, Y., Kuwano, H., Mori, M., Nicolson, G. L., & Sugimachi, K. (1999). Overexpression of metastasis-associated MTA1 mRNA in invasive oesophageal carcinomas. British Journal of Cancer, 79(11–12), 1723–1726.
Neri, A., Welch, D., Kawaguchi, T., & Nicolson, G. L. (1982). Development and biologic properties of malignant cell sublines and clones of a spontaneously metastasizing rat mammary adenocarcinoma. Journal of National Cancer Institute, 68(3), 507–517.
Nicolson, G. L., Gallick, G. E., Spohn, W. H., Lembo, T. M., & Tainsky, M. A. (1992). Transfection of activated c-H-rasEJ/psv2neo or psv2neo genes into rat mammary cells: rapid stimulation of clonal diversification of spontaneous metastatic and cell-surface properties. Oncogene, 7(6), 1127–1135.
Pencil, S. D., Toh, Y., & Nicolson, G. L. (1993). Candidate metastasis-associated genes of the rat 13762NF mammary adenocarcinoma. Breast Cancer Research and Treatment, 25(2), 165–174.
Taniguchi, S., Miyamoto, S., Sadano, H., & Kobayashi, H. (1991). Rat elongation factor 1 alpha: sequence of cDNA from a highly metastatic fos-transferred cell line. Nucleic Acids Research, 19(24), 6949.
Thompson, E. W., Brunner, N., Torri, J., Johnson, M. D., Boulay, V., Wright, A., et al. (1993). The invasive and metastatic properties of hormone-independent but hormone-responsive variants of MCF-7 human breast cancer cells. Clinical and Experimental Metastasis, 11(1), 15–26.
Brunner, N., Frandsen, T. L., Holst-Hansen, C., Bei, M., Thompson, E. W., Wakeling, A. E., et al. (1993). MCF7/LCC2: a 4-hydroxytamoxifen resistant human breast cancer variant that retains sensitivity to the steroidal antiestrogen ICI 182,780. Cancer Research, 53(14), 3229–3232.
Zhang, R. D., Fidler, I. J., & Price, J. E. (1991). Relative malignant potential of human breast carcinoma cell lines established from pleural effusions and a brain metastasis. Invasion and Metastasis, 11(4), 204–215.
Toh, Y., Pencil, S. D., & Nicolson, G. L. (1995). Analysis of the complete sequence of the novel metastasis-associated candidate gene, mta1, differentially expressed in mammary adenocarcinoma and breast cancer cell lines. Gene, 159(1), 97–104.
Manavathi, B., Singh, K., & Kumar, R. (2007). MTA family of coregulators in nuclear receptor biology and pathology. Nuclear Receptor Signaling, 5, 1–8.
Toh, Y., & Nicolson, G. L. (2009). The role of the MTA family and their encoded proteins in human cancers: molecular functions and clinical implications. Clinical and Experimental Metastasis, 26(3), 215–227.
Toh, Y., & Nicolson, G. L. (2011). MTA1 of the MTA (metastasis-associated) gene family and its encoded proteins: molecular and regulatory functions and its role in human cancer progression. Atlas of Genetics and Cytogenetics in Oncology and Haematology, 15(3), 303–315.
Toh, Y., & Nicolson, G. L. (2013). Signaling pathways of MTA family proteins as regulators of cancer progression and metastasis. In R. R. Resende & H. Ulrich (Eds.), Trends in stem cell proliferation and cancer research (pp. 251–275). Dordrecht: Springer.
Singh, R. R., & Kumar, R. (2007). MTA family of transcriptional metaregulators in mammary gland morphogenesis and breast cancer. Journal of Mammary Gland Biology and Neoplasia, 12(2–3), 115–125.
Millard, C. J., Watson, P. J., Celardo, I., Gordiyenko, Y., Cowley, S. M., Robinson, C. V., et al. (2013). Class I HDACs share a common mechanism of regulation by inositol phosphates. Molecular Cell, 51(1), 57–67.
Alqarni, S. S., Murthy, A., Zhang, W., Przewloka, M. R., Silva, A. P., Watson, A. A., et al. (2014). Insight into the architecture of the NuRD complex: structure of the RbAp48-MTA1 subcomplex. Journal of Biological Chemistry, 289(32), 21844–21855.
Bowen, N. J., Fujita, N., Kajita, M., & Wade, P. A. (2004). Mi-2/NuRD: multiple complexes for many purposes. Biochimica et Biophysica Acta, 1677(1–3), 52–57.
Tong, J. K., Hassig, C. A., Schnitzler, G. R., Kingston, R. E., & Schreiber, S. L. (1998). Chromatin deacetylation by an ATP-dependent nucleosome remodelling complex. Nature, 395(6705), 917–921.
Xue, Y., Wong, J., Moreno, G. T., Young, M. K., Cote, J., & Wang, W. (1998). NuRD, a novel complex with both ATP-dependent chromatin-remodeling and histone deacetylase activities. Molecular Cell, 2(6), 851–861.
Zhang, Y., Ng, H. H., Erdjument-Bromage, H., Tempst, P., Bird, A., & Reinberg, D. (1999). Analysis of the NuRD subunits reveals a histone deacetylase core complex and a connection with DNA methylation. Genes and Development, 13(15), 1924–1935.
Wade, P. A., Gegonne, A., Jones, P. L., Ballesta, R. E., Aubry, F., & Wolffe, A. P. (1999). Mi-2 complex couples DNA methylation to chromatin remodelling and histone deacetylation. Nature Genetics, 23(1), 62–66.
Mazumdar, A., Wang, R. A., Mishra, S. K., Adam, L., Bagheri-Yarmand, R., Mandal, M., et al. (2001). Transcriptional repression of oestrogen receptor by metastasis-associated protein 1 corepressor. Nature Cell Biology, 3(1), 30–37.
Acknowledgments
The authors acknowledge support from members of Department of Gastroenterological Surgery, National Kyushu Cancer Center, Japan, and foundation and private donations to the Institute for Molecular Medicine.
Conflict of interest
The authors declare that they have no conflict of interest.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Toh, Y., Nicolson, G.L. Identification and characterization of metastasis-associated gene/protein 1 (MTA1). Cancer Metastasis Rev 33, 837–842 (2014). https://doi.org/10.1007/s10555-014-9510-8
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10555-014-9510-8