Abstract
The black marsh turtle (Siebenrockiella crassicollis) has morphologically differentiated X and Y sex chromosomes. To elucidate the origin and evolutionary process of S. crassicollis X and Y chromosomes, we performed cross-species chromosome painting with chromosome-specific DNA from Chinese soft-shelled turtle (Pelodiscus sinensis) and chromosome mapping of the sex-linked genes of S. crassicollis using FISH. The X and Y chromosomes of S. crassicollis were hybridized with DNA probe of P. sinensis chromosome 5, which is homologous to chicken chromosome 5. S. crassicollis homologues of 14 chicken chromosome 5-linked genes were all localized to the X long arm, whereas two genes were mapped to the Y short arm and the other 12 genes were localized to the Y long arm in the same order as the X chromosome. This result suggests that extensive linkage homology has been retained between chicken chromosome 5 and S. crassicollis X and Y chromosomes and that S. crassicollis X and Y chromosomes are at an early stage of sex chromosome differentiation. Comparison of the locations of two site-specific repetitive DNA sequences on the X and Y chromosomes demonstrated that the centromere shift was the result of centromere repositioning, not of pericentric inversion.
Similar content being viewed by others
Introduction
Testudines exhibits two different types of sex-determining systems: temperature-dependent sex determination (TSD) in which sex ratio is determined by the incubation temperature of eggs and genotypic sex determination (GSD) with both male heterogamety (XY/XX) and female heterogamety (ZZ/ZW) (Head et al. 1987; Sarre et al. 2004; Valenzuela and Lance 2004; Olmo and Signorino 2005). Most of testudinian species exhibit TSD, and heteromorphic sex chromosomes have been known in only eight GSD species: the XY/XX system in six species, Brazilian radiolated swamp turtle (Acanthochelys radiolata, formerly Platemys radiolata, Chelidae) (McBee et al. 1985), the eastern long-necked turtle (Chelodina longicollis, Chelidae) (Ezaz et al. 2006) and Murray river turtle (Emydura macquarii, Chelidae) (Martinez et al. 2008), Chiapas giant musk turtle (Staurotypus salvinii, Kinosternidae), Mexican giant musk turtle (Staurotypus triporcatus, Kinosternidae) (Bull et al. 1974), the black marsh turtle (Siebenrockiella crassicollis, Geoemydidae) (Carr and Bickham 1981); and the ZZ/ZW system in two species, the brown-roofed turtle (Pangshura (Kachuga) smithii, Geoemydidae) (Sharma et al. 1975) and Chinese soft-shelled turtle (Pelodiscus sinensis, Trionychidae) (Kawai et al. 2007). Chromosome numbers of turtles are highly variable, ranging from 2n = 22 to 66 (Ayres et al. 1969; Bickham and Baker 1976; Bickham et al. 1983). Their karyotypes are generally composed of two major chromosomal components, macrochromosomes and microchromosomes, according to their physical sizes; the most common diploid chromosome number is around 50–52, consisting of 12–14 pairs of macrochromosomes and 12–14 pairs of microchromosomes. The sizes of sex chromosomes are varied between species. C. longicollis and P. sinensis have the XY- and ZW-types of micro-sex chromosomes, respectively, and the other six species have large- or middle-sized sex chromosomes. The sex chromosomes of E. macquarii are considered to have been established by a translocation of an ancestral Y microchromosome as found in the species of a sister clade, C. longicollis, onto the tip of the fourth largest autosome (Martinez et al. 2008); however, the origins of the sex chromosomes of these species and their homologies have been still unknown except for the ZW micro-sex chromosomes of P. sinensis. Comparative gene mapping between P. sinensis and chicken revealed that the six largest chromosomes were almost equivalent between two species (Matsuda et al. 2005); however, P. sinensis Z chromosome was homologous to chicken chromosome 15 but not to the chicken Z chromosome (Kawagoshi et al. 2009). This result suggests that the ZW-type sex chromosomes of birds and turtles differentiated independently in each lineage from different autosome pairs of the common ancestor of Arcosauromorpha, which diverged around 250 million years ago (MYA) (Kumazawa and Nishida 1999; Janke et al. 2001; Rest et al. 2003; Iwabe et al. 2005).
The black marsh turtle (S. crassicollis, Geoemydidae) is abundant throughout much of the southeast Asian rainforest (Thailand, Sumatra, Borneo, Java and peninsular Malaysia) and has heteromorphic sex chromosomes of the XY/XX-type (Carr and Bickham 1981, 1986). We revealed that the ZW sex chromosomes of P. sinensis correspond to chicken chromosome 15 (Kawagoshi et al. 2009); however, the origins of the testudinian XY sex chromosomes are still unknown. In this study, to elucidate the origin and differentiation process of S. crassicollis XY chromosomes, we first conducted chromosome painting with chromosome-specific DNA probes from P. sinensis, and then performed chromosome mapping of S. crassicollis homologues of chicken genes located on the chromosome that is homologous to S. crassicollis XY chromosomes. We also cloned site-specific repetitive DNA sequences from S. crassicollis and used them as cytogenetic markers to determine the process of chromosomal rearrangements that occurred between the X and Y chromosomes. This is the first report on the origin and differentiation process of the XY-type sex chromosomes in Testudines, which was examined by comparative gene mapping.
Material and methods
Cell culture and chromosome preparation
One captive bred male of S. crassicollis was purchased and used for this study. After intraperitoneal injection of pentobarbital, the heart, lung, and the mesentery were removed and used for cell culture. All experimental procedures with the animal conformed to the guideline established by the Animal Care Committee, Nagoya University, Japan. The tissues were minced and cultured in Dulbecco's modified Eagle's medium (Invitrogen-GIBCO, Carlsbad, CA, USA) supplemented with 15% fetal bovine serum (Invitrogen-GIBCO), 100 μg/ml kanamycin, and 1% antibiotic-antimycotic (PSA) (Invitrogen-GIBCO). The cultures were incubated at 26°C in a humidified atmosphere of 5% CO2 in air. Primary cultured fibroblast cells were harvested using trypsin and then subcultured. For chromosome preparation, the fibroblast cells at log phase were harvested after colcemid treatment (120 ng/ml) for 45 min, suspended in 0.075 M KCl at room temperature for 20 min and fixed with methanol/acetic acid (3:1) three times. The cell suspension was dropped on cleaned glass slides and air-dried. For gene mapping by FISH, replication R-banding was performed to identify each chromosome as described previously (Matsuda and Chapman 1995). The fibroblast cell cultures were treated with BrdU (12 μg/ml) (Sigma-Aldrich, St. Louis, MO, USA) at late replication stage for 12 h including 45 min colcemid treatment, and chromosome preparations were made as mentioned above. The slides were dried at room temperature for 2–3 days. After staining the slides with Hoechst 33258 (1 μg/ml) for 10 min, R bands were obtained by heating them at 65°C for 3 min and exposing them to UV light at 65°C for an additional 6.5 min. The slides were kept at −80°C until use.
C banding
To examine the chromosomal distribution of constitutive heterochromatin in S. crassicollis chromosomes, C banding was performed by the standard BSG method (Sumner 1972) with slight modification; chromosome slides were treated with 0.2 N HCl at room temperature for 5 min and then 5% Ba(OH)2 at 50°C for 2 min.
Chromosome painting
Chromosome-specific DNA probes of P. sinensis were used for comparative chromosome painting of S. crassicollis chromosomes. The P. sinensis chromosome paints were prepared and provided by Fengtang Yang and Patricia O'Brien, Department of Veterinary Medicine, Cambridge University, UK. Chromosome painting was performed as described in Matsubara et al. (2003) with slight modification. DNA probe (1 μg) was labeled with biotin-16-dUTP (Roche Diagnostics, Basel, Switzerland) using a nick translation kit (Roche Diagnostics). After pre-hybridization for 15 min at 37°C, hybridization was carried out at 37°C for 5 days. After hybridization, the slide was washed, incubated with fluorescein-conjugated avidin (FITC-avidin) (Roche Diagnostics), and stained with 0.75 μg/ml propidium iodide (PI).
Molecular cloning of repetitive DNA sequences
High molecular weight genomic DNA was extracted from liver tissue. Genomic DNA was digested with 18 restriction endonucleases, ApaI, AluI, BamHI, BglI, BglII, Csp45I, DraI, EcoRI, EcoRV, HaeIII, HindIII, HinfI, PvuII, SacI, SalI, SpeI, XbaI, and XhoI, size-fractionated with 2% agarose gel electrophoresis, and then stained with ethidium bromide. The prominent DNA bands of repetitive sequences were eluted from the gel using a QIAquick Gel Extraction Kit (Qiagen, Hilden, Germany), ligated into pGEM®-7Zf(+) vector (Promega, Tokyo, Japan), and transformed into competent cells of Escherichia coli strain DH5α (TOYOBO, Osaka, Japan). The sizes of DNA fragments inserted into the vector were confirmed by electrophoresis of PCR products that were amplified using T7 and Sp6 primers, and the clones were used for FISH.
Molecular cloning of S. crassicollis homologues of chicken genes
Testes of S. crassicollis were homogenized and lysed with TRIzol Reagent (Invitrogen, Carlsbad, CA, USA), and total RNA was extracted following the manufacturer's instruction. Reverse transcription-PCR and molecular cloning of cDNA fragments were performed as described previously (Kawai et al. 2009). Nucleotide sequences were determined using an ABI PRISM 3100 DNA Analyzer (Applied Biosystems, Foster, CA, USA) after sequencing reactions with a Big Dye Terminator v1.1 Cycle Sequencing Kit (Applied Biosystems).
FISH mapping of genes
FISH was performed for chromosomal localization of repetitive DNA sequences and cDNA fragments of functional genes following Matsuda and Chapman (1995). Repetitive DNA fragment (250 ng) was labeled with biotin-16-dUTP by nick translation and hybridized to chromosome slides. After hybridization, the slides were incubated with FITC-avidin and stained with 0.75 μg/ml PI. For chromosome mapping of cDNA fragments, 250 ng cDNA probe was labeled with biotin-16-dUTP (Roche Diagnostics) by nick translation. After hybridization, the probe DNA was reacted with goat anti-biotin antibody (Vector Laboratories), and then stained with Alexa Fluor 488 rabbit anti-goat IgG (H + L) conjugate (Invitrogen-Molecular Probes) and subsequently counter-stained with 0.75 μg/ml PI.
Southern blot hybridization
Genomic DNA of S. crassicollis was digested with nine restriction endonucleases (AluI, ApaI, BamHI, HaeIII, HindIII, HinfI, RsaI, HpaII, MspI). The digested DNA was fractionated in 2% agarose gel and transferred onto nylon membranes (Roche Diagnostics). DNA fragments of repetitive sequences were labeled with digoxigenin (DIG)-11-dUTP using PCR DIG Labeling Mix (Roche Diagnostics) and hybridized to the membranes. Hybridization was carried out at 45°C overnight in DIG Easy Hyb solution (Roche Diagnostics). After hybridization, the membranes were washed at 45°C in 0.1% SDS/2× SSC, 0.1% SDS/1× SSC, 0.1% SDS/0.5× SSC and 0.1% SDS/0.1× SSC for 15 min each. Chemiluminescent signals were detected using anti-digoxigenin-AP Fab fragments and CDP-Star (Roche Diagnostics) and exposed to BioMax MS autoradiography film (Kodak, Rochester, NY, USA).
Slot-blot hybridization
To examine the conservation of repetitive sequences among different species, slot-blot hybridization was conducted with DIG-11-dUTP-labeled repetitive DNA fragments. Genomic DNA was collected from three species of Testudines, the black marsh turtle (S. crassicollis, Geoemydidae), Mexican giant musk turtle (S. triporcatus, Kinosternidae), and Chinese soft-shelled turtle (P. sinensis, Trionychidae); seven species of Crocodilia, Siamese crocodile (Crocodylus siamensis, Crocodylidae), the false gharial (Tomistoma schlegelii, Crocodylidae), Indian gharial (Gavialis gangeticus, Gavialidae), American alligator (Alligator mississippiensis, Alligatoridae), Chinese alligator (Alligator sinensis, Alligatoridae), the common caiman (Caiman crocodilus, Alligatoridae), and the broad-nosed caiman (Caiman latirostris, Alligatoridae) (Kawagoshi et al. 2008); two species of Aves, chicken (Gallus gallus, Phasianidae, Galliformes) and ostrich (Struthio camelus, Struthionidae, Struthioniformes); and four species of Squamata, Burmese python (Python molurus bivittatus, Pythonidae), Japanese four-striped rat snake (Elaphe quadrivirgata, Colubridae), habu (Protobothrops flavoviridis, formerly Trimeresurus flavoviridis, Viperidae), and Hokou gecko (Gekko hokouensis, Gekkonidae). Genomic DNA (200 ng) was denatured with NaOH and transferred onto nylon membranes using BIO-DOT SF blotting equipment (Bio-Rad, Hercules, CA, USA). Hybridization and detection of signals were performed as described in the section of Southern blot hybridization.
Results
Karyotype of S. crassicollis
Chromosome numbers of 2n = 50 and 2n = 52 have been reported for S. crassicollis (Killebrew 1977; Carr and Bickham 1981, 1986). In the present study, we examined 17 Giemsa-stained metaphase spreads of one male S. crassicollis. Chromosome number was 2n = 50 for all metaphase spreads, and the karyotype consisted of four pairs of large chromosomes, five and two pairs of middle- and small-sized chromosomes, respectively, and 14 pairs of indistinguishable microchromosomes (Fig. 1). Sex chromosomes were the third or fourth largest heteromorphic chromosomes; the X and Y were submetacentric and metacentric, respectively.
C bands of S. crassicollis chromosomes
C-positive heterochromatin blocks were observed in the centromeric regions of almost all autosomes, and the centromeric heterochromatin was large in several pairs of chromosomes (Fig. 2). The short arm of the X chromosome and a distal half of the Y short arm were heterochromatic and an intense heterochromatin block was located in the pericentromeric region of the X long arm.
Chromosome painting probed with chromosome-specific DNA of P. sinensis
FITC-labeled chromosome 5-specific DNA probe of P. sinensis painted the X and Y chromosomes of S. crassicollis but not any other chromosomes (Fig. 3), indicating that S. crassicollis X and Y sex chromosomes are a counterpart of P. sinensis chromosome 5, which is homologous to chicken chromosome 5 (Matsuda et al. 2005). The painted signal was not detected in the pericentromeric region of the X long arm, which corresponded to C-positive heterochromatin.
Chromosome painting with chromosome 5-specific DNA probe of P. sinensis to PI-stained metaphase spreads of female Pelodiscus sinensis (a) and male S. crassicollis (b). (c) Hoechst-stained pattern of the same metaphase spread (b). The probe painted chromosome 5 of P. sinensis (a) and the X and Y chromosomes of S. crassicollis (b). Arrows indicate the X and Y chromosomes. Scale bar represents 10 μm
Molecular cloning of S. crassicollis homologues of chicken chromosome 5-linked genes
S. crassicollis homologues of the following 14 chicken chromosome 5-linked genes were cloned using a total cDNA pool derived from S. crassicloiis testes: BUB1B, CPT1A, CRY2, CTR9, DDB1, DEAF1, DYNC1H1, EIF4G2, GARNL1, PAPOLA, PSMC1, PYGL, ST5, and WT1. PCR primers of 14 genes are shown in Table 1. Nucleotide sequences of the cDNA fragments of 14 genes were deposited in DDBJ (http://www.ddbj.nig.ac.jp/). Nucleotide sequence identities of the equivalent regions of the cDNA fragments between chicken and S. crassicollis ranged from 65.4% to 96.4% (Table 2). The CPT1A cDNA fragment showed the lowest identity (65.4%) with the chicken homologue.
FISH mapping of S. crassicollis homologues of chicken chromosome 5-linked genes
Hoechst-stained bands obtained by the replication banding method, which correspond to G bands, made it possible to identify each chromosome precisely and to determine the subchromosomal locations of the genes. Fourteen genes were all localized to the X long arm, whereas two of them (DDB1 and DEAF1) were mapped to the Y short arm, and the other 12 genes were all localized at the same positions on the Y long arm as those of the X chromosome (Figs. 4, 5). For FISH mapping, more than 25–30 metaphase spreads were observed for each gene, and the hybridization efficiency on the X and/or Y chromosomes ranged from 20% to 32% of metaphase spreads.
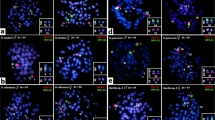
Chromosomal localization of S. crassicollis homologues of 14 chicken chromosome 5-linked genes, DDB1, DEAF1, CTR9, ST5, WT1, EIF4G2, CRY2, BUB1B, CPT1A, GARNL1, PAPOLA, DYNC1H1, PSMC1, and PYGL, to chromosomes of male S. crassicollis. a, b FISH pattern of BUB1B on PI-stained metaphase spread (a) and Hoechst-stained pattern of the same metaphase spread (b). c–z, a′, b′ FISH patterns of DDB1 (c), DEAF1 (e), CTR9 (g), ST5 (i), WT1 (k), EIF4G2 (m),CRY2(o), CPT1A(q), GARNL1(s), PAPOLA(u), DYNC1H1(w), PSMC1(y) and PYGL(a′) on PI-stained X chromosomes, and FISH patterns of DDB1 (d), DEAF1 (f), CTR9 (h), ST5 (j), WT1 (l), EIF4G2 (n), CRY2 (p), CPT1A (r), GARNL1 (t), PAPOLA (v), DYNC1H1 (x), PSMC1 (z) and PYGL (b′) on PI-stained Y chromosomes. Arrows indicate the hybridization signals of the genes. Scale bars represent 10 μm (a, b) and 2.5 μm (c–z, a′, b′)
Comparative maps of 14 functional genes, DDB1, DEAF1, CTR9, ST5, WT1, EIF4G2, CRY2, BUB1B, CPT1A, GARNL1, PAPOLA, DYNC1H1, PSMC1, and PYGL on chicken chromosome 5 (GGA5) and S. crassicollis X (SCRX) and Y (SCRY) chromosomes. The gene order of 14 genes on GGA5 was taken from the Ensemble Chicken Web Server (http://uswest.ensembl.org/Gallus_gallus/Info/Index) and Chicken Genome Resources (http://www.ncbi.nlm.nih.gov/genome/guide/chicken/)
Molecular cloning of repetitive sequences and their chromosomal distribution
Genomic DNA of S. crassicollis was digested with 18 restriction endonucleases, and the prominent DNA bands at approximately 300 bp in AluI digest were isolated (data not shown). Thirteen clones were obtained and used for FISH analysis. FISH signals were detected for two of 13 DNA clones. The SCR-AluI-01 sequence was distributed predominantly in the centromeric heterochromatin regions of three pairs of macrochromosomes, six pairs of middle- and/or small-sized chromosomes and nine pairs of microchromosomes (Fig. 6). Small signals were also detected in the telomeric regions of several pairs of large- and middle-sized chromosomes. In the sex chromosomes, the signals were observed in the telomeric regions of both the X and Y chromosomes, the pericentromeric region of the X long arm, and an interstitial region of the Y short arm. The SCR-AluI-09 sequence was localized to the centromeric heterochromatin of chromosome 1 and 2 and two pairs of middle-sized chromosomes and to the pericentromeric region of the X long arm and an interstitial region of the Y short arm (Fig. 6). The SCR-AluI-01 and SCR-AluI-09 sequences were not overlapped with the intense C-positive heterochromatin of the X long arm.
Chromosomal distribution of the SCR-AluI-01 and SCR-AluI-09 sequences on male S. crassicollis chromosomes. a, b Distribution pattern of SCR-AluI-01 (a) and SCR-AluI-09 (b) on PI-stained metaphase spreads. Arrows indicate the X and Y chromosomes. c–h Enlarged photographs of the X and Y chromosomes. (c) FISH pattern of SCR-AluI-01 on PI-stained X and Y chromosomes. d, e PI-stained (d) and Hoechst-stained (e) X and Y chromosomes. (f) FISH pattern of SCR-AluI-09 on PI-stained X and Y chromosomes. g, h PI-stained (g) and Hoechst-stained (h) X and Y chromosomes. Scale bars represent 10 μm in (a, b) and 2.5 μm in (c–h)
Nucleotide sequences of repetitive sequences
The length and G + C content were 308 bp and 70.8% for SCR-AluI-01 (AB674502) (Fig. 7a) and 287 bp and 69.3% for SCR-AluI-09 (AB674503) (Fig. 7b). SCR-AluI-09 was composed of seven 36 bp-unit and one 34 bp-unit elements (Fig. 7c). Nucleotide sequence identities and G + C content among the internal units of SCR-AluI-09 ranged from 66.7% between unit 5 and 6 to 97.2% between unit 1 and 3, and their G + C content was 68.8% on average, ranging from 61.1% (unit 6) to 75.0% (unit 1 and 3). There was no sequence homology between SCR-AluI-01 and SCR-AluI-09. These nucleotide sequences were compared with all the DNA sequences in the nonredundant database using the BLAST program (http://blast.ncbi.nlm.nih.gov/Blast.cgi). The nucleotide sequence of SCR-AluI-01 had the homology with partial sequences of two BAC clones (CHY3-80H12 and CHY3-29H12) derived from the painted turtle (Chrysemys picta) genome. Homologies of SCR-AluI-01 with CHY3-29H12 and CHY3-80H12 were both 87% in the equivalent regions of 302 bp. No sequences with significant homology were detected for SCR-AluI-09.
Nucleotide sequences of the SCR-AluI-01 and SCR-AluI-09 fragments. a Nucleotide sequence of the SCR-AluI-01 fragment. Internal restriction sites of four restriction endonucleases are represented as follows: HaeIII (straight line), RsaI (crooked line), HinfI (dotted line), and MspI (double line). b Nucleotide sequence of the SCR-AluI-09 fragment. Each internal unit of SCR-AluI-09 is shown by different color box and number. Internal restriction sites of four restriction endonucleases are represented as follows: HaeIII (straight line), RsaI (crooked line), HinfI (dotted line), and MspI (double line). c Nucleotide sequences of eight internal units of SCR-AluI-09 and comparison of their nucleotide sequences. Hyphens indicate the same nucleotides as those of the consensus sequence of eight internal units shown at the top
Genomic organization of repetitive sequences
In hybridization probed with SCR-AluI-01, polymeric hybridization bands were found at 308, 616, and 924 bp in AluI digest, indicating that this sequence can be categorized as satellite DNA; however, one another repeated unit of about 400 bp with AluI site, which was not molecularly cloned in the present study, was contained in this family (Fig. 8a). In ApaI digest, hybridization bands comprising two units of the SCR-AluI-01 element and their dimeric bands were found at about 600 and 1,200 bp, respectively. The same sizes of hybridization bands were also observed in AluI digest. These results suggest that AluI and ApaI sites of SCR-AluI-01 are highly conserved in this sequence family and ApaI site is located in every other unit. HinfI and RsaI sites were also highly conserved; however, the unit sizes of the SCR-AluI-01 elements with HinfI and RsaI sites were different from those with AluI and ApaI sites, suggesting that the SCR-AluI-01 sequences consist of various unit sizes of elements with different restriction sites. The hybridization bands detected at less than 308 bp in HaeIII and MspI digests were derived from multiple internal restriction sites contained in the elements. CCGG sequence is cleaved by HpaII and MspI, and HpaII does not cleave when the CG sequence is methylated. In digests cleaved with two isoschizomers, all hybridization bands were observed at less than 600 bp in MspI digest, while no low molecular weight bands were found in HpaII digest. This remarkable difference indicates that the SCR-AluI-01 sequences are hypermethylated in the genome.
Smears of DNA in the range from 300 bp to 2 kb were observed when digested with AluI and hybridized with SCR-AluI-09 probe (Fig. 8b). Hybridization patterns also appeared as broad smear in BamHI, HindIII, HinfI, RsaI and HpaII digests, suggesting that these restriction sites are less conserved in the SCR-AluI-09 sequences. Intense single bands were observed in HaeIII and MspI digests, which might be derived from conserved internal restriction sites in the SCR-AluI-09 sequence elements. The HpaII sites were highly methylated in the SCR-AluI-09 sequences as well as the SCR-AluI-01 sequences.
Nucleotide sequence conservation of repetitive sequences
Nucleotide sequence conservation of the SCR-AluI-01 and SCR-AluI-09 sequences was examined by slot-blot hybridization for 16 species from three reptilian orders and two avian orders (Fig. 9). In SCR-AluI-01, intense hybridization signals were observed in three testudinian species, whereas weak signals were observed for the other all species (Fig. 9a). In contrast, the hybridization signal for SCR-AluI-09 was observed for only S. crassicollis, and there were no signals in the other two testudinian species and all species of Crocodilia, Aves, and Squamata (Fig. 9b).
Slot-blot hybridization probed with the SCR-AluI-01 (a) and SCR-AluI-09 (b) fragments. Genomic DNA used for this experiment were as follows: SCR (Siebenrockiella crassicollis), STR (Staurotypus triporcatus), and PSI (Pelodiscus sinensis) of Testudines written in black; CSI (Crocodylus siamensis), TSC (Tomistoma schlegelii), GGA (Gavialis gangeticus), AMI (Alligator mississippiensis), ASI (Alligator sinensis), CCR (Caiman crocodilus), and CLA (Caiman latirostris) of Crocodilia written in red; SCA (Struthio camelus) and GGA (Gallus gallus) of Aves written in green; PMB (Python molurus bivittatus), EQU (Elaphe quadrivirgata), PFL (Protobothrops flavoviridis), and GHO (Gekko hokouensis) of Squamata written in blue (PMB, EQU, and PFL) and purple (GHO)
Discussion
In this study, we investigated the origin and evolutionary process of the X and Y sex chromosomes of S. crassicollis using cross-species chromosome painting with chromosome-specific DNA from Chinese soft-shelled turtle (P. sinensis) and chromosome mapping of the sex-linked genes of S. crassicollis. The X and Y chromosomes of S. crassicollis was homologous to chicken chromosome 5, and S. crassicollis homologues of 14 chicken chromosome 5-linked genes were all localized to the X and Y chromosomes in the same order, although the centromeric position was different between two chromosomes. Comparison of the locations of two site-specific repetitive DNA sequences on the X and Y chromosomes suggested that the centromere shift on the Y chromosome was the result of centromere repositioning that occurred on the proto-X chromosome, not of pericentric inversion.
Homology of sex chromosomes: chromosome painting and candidate gene mapping
Cross-species chromosome painting with P. sinensis chromosome-specific DNA revealed the homology of S. crassicollis X and Y sex chromosomes with P. sinensis chromosome 5, which correspond to chicken chromosome 5 (Matsuda et al. 2005). S. crassicollis homologues of 14 chicken chromosome 5-linked genes were all localized to the X and Y chromosomes of S. crassicollis, indicating that this genetic linkage has been retained in Testudines and Aves since Archosauromorpha first appeared about 250 MYA (Kumazawa and Nishida 1999; Janke et al. 2001; Rest et al. 2003; Iwabe et al. 2005). This linkage has also been extensively conserved in human chromosome 11 and 14 for eight genes (CPT1A, CRY2, CTR9, DDB1, DEAF1, EIF4G2, ST5, WT1) and five genes (DYNC1H1, GARNL1, PAPOLA, PSMC1, PYGL), respectively, although only BUB1B is localized to human chromosomes 15 (National Center for Biotechnology Information, http://www.ncbi.nlm.nih.gov/). The genetic linkage of S. crassicollis X and Y chromosomes therefore have been highly conserved in amniotes; however, S. crassicollis XY sex chromosomes have no homology with the ZW sex chromosomes of P. sinensis, which is homologous to chicken chromosome 15 (Kawagoshi et al. 2009), suggesting that their sex chromosomes were differentiated independently within the same order from different autosomes of the common ancestor. The eastern long-necked turtle (C. longicollis) of Chelidae has micro-XY sex chromosomes, which are detectable by C banding, reverse fluorescent staining, or comparative genomic hybridization (Ezaz et al. 2006). In contrast, Murray river turtle (E. macquarii) and Brazilian radiolated swamp turtle (A. radiolata) belonging to the same family have large-sized (submetaentric X and metacentric Y) and middle-sized (metacentric X and acrocentric Y) sex chromosomes, respectively (McBee et al. 1985; Martinez et al. 2008). This significant size differences among three species implied independent origins of sex chromosomes even within the same family. Comparative mapping of these three species with S. crassicollis is required to understand the diversity of sex chromosomal origins in XY-type turtles. Male heterogamety is also found in two species of Kinosternidae, Chiapas giant musk turtle (S. salvinii) and Mexican giant musk turtle (S. triporcatus) (Bull et al. 1974), which are the most closely related to S. crassicollis among XY-type turtles. S. salvini and S. triporcatus also have the fourth largest X chromosome, and the centromere and short arms of their X chromosomes were heterochromatized. These characteristics are similar to those of S. crassicollis X chromosome, and thus it would be considerable interest to investigate sex chromosomal homologies among three species.
The order of 14 chicken chromosome 5-linked genes on S. crassicollis Y chromosome was the same as that on the long arm of X chromosome, although two genes (DDB1 and DEAF1) were localized to the proximal region of the Yp. The X and Y chromosomes of S. crassicollis therefore are at an early stage of sex chromosome differentiation with no deletion of the Y homologues. The gene order of S. crassicollis X chromosome was largely different from that of chicken chromosome 5, indicating that intra-chromosomal rearrangements have occurred extensively between two lineages, although the gene order in the common ancestor is still unknown.
Genomic organization of sex chromosome repeats and XY differentiation
Two novel families of repetitive sequences were isolated, which served as good cytogenetic markers to delineate the differentiation process of S. crassicollis X and Y chromosomes. SCR-AluI-09 was located in the pericentromeric region of the Xq and the interstitial region of the Yp. SCR-AluI-01 was co-localized with SCR-AluI-09 and also localized to the distal ends of both the short and long arms of the X and Y chromosomes. Centromere repositioning is most likely to explain this result. Centromere repositioning occurs without the disruption of gene order (Montefalcone et al. 1999) and is caused by the following phenomena: the activation of a neocentromere, acquisition and/or amplification of neocentromeric DNA, and inactivation and/or deletion of the old centromere (Wong and Choo 2001). According to our schematic representation, the SCR-AluI-01 and -09 sequences and DDB1 and DEAF1 in the pericentromeric region of the proto-X long arm moved to the Y short arm with no change to the order of SCR-AluI-01/SCR-AluI-09–DDB1–DEAF1 (Fig. 10). Meiotic recombination should be suppressed around the subcentromeric regions containing DDB1 and DEAF1 genes between the X and Y chromosomes, and then the X chromosome-specific repetitive sequences may have been amplified in the pericentromeric region of the X long arm, which are intensely stained with PI and with Giemsa by C banding. The alternative hypothesis is that a pericentric inversion occurred in the pericentromeric region; however, the order of SCR-AluI-01/SCR-AluI-09–DDB1–DEAF1 on the Xq was the same as that on the Yp.
Schematic representation for the differentiation process of the X and Y chromosomes in S. crassicollis. Chromosomal locations of DDB1 and DEAF1 are represented by red and blue circles, respectively. Yellow ellipse and yellow-and blue-striped ellipse indicate the chromosomal location of SCR-AluI-01 and the co-location of SCR-AluI-01 and SCR-AluI-09, respectively. Two different types of heterochromatic regions are each gray- and black-painted. Arrows show the differentiation into the X (SCRX) and Y (SCRY) chromosomes from the proto-X chromosome, showing that X chromosome-specific repetitive sequences were amplified in the pericentromeric region of the X long arm and that the co-localized SCR-AluI-01 and -09 sequences and DDB1 and DEAF1 genes in the pericentromeric region of the proto-X long arm moved to the Y short arm as a result of centromere repositioning
SCR-AluI-01 was conserved through three testudinian families, Geoemydidae, Kinosternidae, and Trionychidae; however, weak signals were observed in Crocodilia, Aves, and Squamata. These results suggested that this sequence originated in the genome of the common ancestor of sauropsids and diverged and/or was amplified in Testudines. In contrast, the signal of SCR-AluI-09 was only found in S. crassicollis, indicating that this sequence was amplified species-specifically as a rapidly evolving molecule. The repetitive sequences comprising the heterochromatized regions of the X and Y short arms and the pericentromeric C-positive heterochromatin block of the X long arm have not been isolated yet. Molecular characterization of these sequences is important to understand the differentiation process of S. crassicollis sex chromosome.
The Y-specific region that is responsible for male sex determination has not been identified in S. crassicollis, but the sex-determining locus is more likely to be located near the pericentromeric region where it is structurally differentiated between the X and Y chromosomes. There were no known genes related to sexual differentiation in the chicken chromosome segment homologous to this region; however, WT1, which is a significant nuclear zinc-finger transcription factor that is required for early kidney and gonad development in mammals (Haber et al. 1991; Armstong et al. 1992; Kreidberg et al. 1993; Rackley et al. 1993; Roberts 2005), was localized to the subcentromeric region of the long arm of Y chromosome in this species. The three-spined stickleback (Gasterosteus aculeatus) and Nile tilapia (Oreochromis niloticus) have duplicated WT1 genes, and one of them is located on the X and Y sex chromosomes in both species (Ross and Peichel 2008; Lee and Kocher 2007). Three inversions occurred in the chromosomal region containing the Y-linked WT1 gene in the stickleback (Ross and Peichel 2008); however, the role of the Y-linked WT1 gene for gonadal differentiation is unknown. In turtles, WT1 expression has been examined in two TSD and one GSD species (Spotila et al. 1998; Valenzuela 2007). In the painted turtle (C. picta), sexual dimorphic expression of WT1 was found in the developing adrenal-kidney-gonad (AKG) complex of embryos incubated at 25°C (male-producing temperature) and 30°C (female-producing temperature) during developmental stage 12 (Valenzuela 2007); the expression level was much higher at 25°C than 30°C. A GSD turtle, the smooth soft-shell turtle (Apalone mutica) also exhibited the thermal sensitive expression pattern (i.e., sexual dimorphic expression) of WT1 between incubation temperatures of 25°C and 30°C during developmental stage 12 (Valenzuela 2007), indicating that this species retain the thermal-sensitivity to sexual differentiation after it acquired the GSD system. Neither expression stages nor expression levels of sexual differentiation-related genes during gonadal development has been examined in S. crassicollis. It is unclear whether the WT1 clone isolated in the present study was derived from the X or Y chromosome; however, it will be of great interest to compare the expression of the X-linked WT1 with that of the Y copy, which may have become functionally divergent from the X copy, as the sex chromosomes differentiated.
Abbreviations
- AMI:
-
Alligator mississippiensis
- ASI:
-
Alligator sinensis
- BCO:
-
Boa constrictor
- BrdU:
-
5-bromo-2′-deoxyuridine
- BSG:
-
Barium hydroxide/saline/Giemsa
- CCD:
-
Charge-coupled device
- CCR:
-
Caiman crocodilus
- cDNA:
-
Complementary DNA
- CLA:
-
Caiman latirostris
- CSI:
-
Crocodylus siamensis
- DDBJ:
-
DNA Data Bank of Japan
- Dig:
-
Digoxigenin
- dNTP:
-
Deoxynucleotide triphosphate
- dUTP:
-
2′-deoxyuridine 5′-triphosphate
- EQU:
-
Elaphe quadrivirgata
- FISH:
-
Fluorescence in situ hybridization
- FITC:
-
Fluorescein isothiocyanate
- GGA:
-
Gallus gallus
- GGA:
-
Gavialis gangeticus
- GHO:
-
Gekko hokouensis
- GSD:
-
Genotypic sex determination
- kb:
-
Kilo base pairs
- MYA:
-
Million years ago
- PCR:
-
Polymerase chain reaction
- PFL:
-
Protobothrops flavoviridis
- PMB:
-
Python molurus bivittatus
- PI:
-
Propidium iodide
- PSI:
-
Pelodiscus sinensis
- SCA:
-
Struthio camelus
- SCR:
-
Siebenrockiella crassicollis
- SDS:
-
Sodium dodecyl sulfate
- SSC:
-
Saline sodium citrate
- STR:
-
Staurotypus triporcatus
- TSC:
-
Tomistoma schlegelii
- TSD:
-
Temperature-dependent sex determination
- UV:
-
Ultraviolet
References
Armstrong JF, Pritchard-Jones K, Bickmore WA, Hastie ND, Bard JB (1992) The expression of the Wilms' tumour gene, WT1, in the developing mammalian embryo. Mech Dev 40:85–97
Ayres M, Sampaio MM, Barros RMS, Dias LB, Cunha OR (1969) A karyological study of turtles from the Brazilian Amazon region. Cytogenetics 8:401–409
Bickham JW, Baker RJ (1976) Chromosome homology and evolution of emydid turtles. Chromosoma 54:201–219
Bickham JW, Bull JJ, Legler JM (1983) Karyotypes and evolutionary relationships of trionychoid turtles. Cytologia 48:177–183
Bull JJ, Moon RG, Legler JM (1974) Male heterogamety in kinosternid turtles (genus Staurotypus). Cytogenet Cell Genet 13:419–425
Carr JL, Bickham JW (1981) Sex-chromosomes of the Asian black pond turtle, Siebenrockiella crassicollis (Testudines: Emydidae). Cytogenet Cell Genet 31:178–183
Carr JL, Bickham JW (1986) Phylogenetic implications of karyotypic variation in the Batagurinae (Testudines: Emydidae). Genetica 70:89–106
Ezaz T, Valenzuela N, Grützner F, Miura I, Georges A, Burke RL, Graves JA (2006) An XX/XY sex microchromosome system in a freshwater turtle, Chelodina longicollis (Testudines: Chelidae) with genetic sex determination. Chromosome Res 14:139–150
Haber DA, Sohn RL, Buckler AJ, Pelletier J, Call KM, Housman DE (1991) Alternative splicing and genomic structure of the Wilms tumor gene WT1. Proc Natl Acad Sci USA 88:9618–9622
Head G, May RM, Pendleton L (1987) Environmental determination of sex in the reptiles. Nature (London) 329:198–199
Iwabe N, Hara Y, Kumazawa Y, Shibamoto K, Saito Y, Miyata T, Katoh K (2005) Sister group relationship of turtles to the bird–crocodilian clade revealed by nuclear DNA-coded proteins. Mol Biol Evol 22:810–813
Janke A, Erpenbeck D, Nilsson M, Arnason U (2001) The mitochondrial genomes of the iguana (Iguana iguana) and the caiman (Caiman crocodylus): implications for amniote phylogeny. Proc R Soc Lond B 268:623–631
Kawagoshi T, Nishida N, Ota H, Kumazawa Y, Endo H, Matsuda Y (2008) Molecular structures of centromeric heterochromatin and karyotypic evolution in the Siamese crocodile (Crocodylus siamensis) (Crocodylidae, Crocodylia). Chromosome Res 16:1119–1132
Kawagoshi T, Uno Y, Matsubara K, Matsuda Y, Nishida C (2009) The ZW micro-sex chromosomes of the Chinese soft-shelled turtle (Pelodiscus sinensis, Trionychidae, Testudines) have the same origin as chicken chromosome 15. Cytogenet Genome Res 125:125–131
Kawai A, Nishida-Umehara C, Ishijima J, Tsuda Y, Ota H, Matsuda Y (2007) Different origins of bird and reptile sex chromosomes inferred from comparative mapping of chicken Z-linked genes. Cytogenet Genome Res 117:92–102
Kawai A, Ishijima J, Nishida C, Kosaka A, Ota H, Kohno S, Matsuda Y (2009) The ZW sex chromosomes of Gekko hokouensis (Gekkonidae, Squamata) represent highly conserved homology with those of avian species. Chromosoma 118:43–51
Killebrew (1977) Mitotic chromosomes of turtles. Tex J Sci 29:245–253
Kreidberg JA, Sariola H, Loring JM, Maeda M, Pelletier J, Housman D, Jaenisch R (1993) WT-1 is required for early kidney development. Cell 74:679–691
Kumazawa Y, Nishida M (1999) Complete mitochondrial DNA sequences of the green turtle and blue-tailed mole skink: statistical evidence for Archosaurian affinity of turtles. Mol Biol Evol 16:784–792
Lee BY, Kocher TD (2007) Exclusion of Wilms tumour (WT1b) and ovarian cytochrome P450 aromatase (CYP19A1) as candidates for sex determination genes in Nile tilapia (Oreochromis niloticus). Anim Genet 38:85–86
Martinez PA, Ezaz T, Valenzuela N, Georges A, Marshall Graves JA (2008) An XX/XY heteromorphic sex chromosome system in the Australian chelid turtle Emydura macquarii: a new piece in the puzzle of sex chromosome evolution in turtles. Chromosome Res 16:815–825
Matsubara K, Nishida-Umehara C, Kuroiwa A, Tsuchiya K, Matsuda Y (2003) Identification of chromosome rearrangements between the laboratory mouse (Mus musculus) and the Indian spiny mouse (Mus platythrix) by comparative FISH analysis. Chromosome Res 11:57–64
Matsuda Y, Chapman VM (1995) Application of fluorescence in situ hybridization in genome analysis of the mouse. Electrophoresis 16:261–272
Matsuda Y, Nishida-Umehara C, Tarui H et al (2005) Highly conserved linkage homology between birds and turtles: bird and turtle chromosomes are precise counterparts of each other. Chromosome Res 13:601–615
McBee K, Bickham JW, Rhodin AGJ, Mittermeier RA (1985) Karyotypic variation in the genus Platemys (Testudines, Pleurodira). Copeia 2:445–449
Montefalcone G, Tempesta S, Rocchi M, Archidiacono N (1999) Centromere repositioning. Genome Res 9:1184–1188
Olmo E, Signorino G (2005) Chromorep: a reptile chromosomes database. Internet references. Retrieved from: http://193.206.118.100/professori/chromorep.pdf
Rackley RR, Flenniken AM, Kuriyan NP, Kessler PM, Stoler MH, Williams BRG (1993) Expression of the Wilms' tumor suppressor gene WT1 during mouse embryogenesis. Cell Growth Different 4:1023–1031
Rest JS, Ast JC, Austin CC, Waddell PJ, Tibbetts EA, Hay JM, Mindell DP (2003) Molecular systematics of primary reptilian lineages and the tuatara mitochondrial genome. Mol Phylogent Evol 29:289–297
Roberts SG (2005) Transcriptional regulation by WT1 in development. Curr Opin Genet Dev 15:542–547
Ross JA, Peichel CL (2008) Molecular cytogenetic evidence of rearrangements on the Y chromosome of the threespine stickleback fish. Genetics 179:2173–2182
Sarre SD, Georges A, Quinn A (2004) The ends of a continuum: genetic and temperature-dependent sex determination in reptiles. BioEssays 26:639–645
Sharma GP, Kaur P, Nakhasi U (1975) Female heterogamety in the Indian crytodiran Chelonian, Kachuga Smithi. Gray In Tiwari KK, Srivistava CB, eds. Dr. B.S. Chauhan Commemoration Volume 359–368. Zoological Society of India, Orissa
Spotila LD, Spotila JR, Hall SE (1998) Sequence and expression analysis of Wt1 and Sox9 in the red-eared slider turtle, Trachemys scripta. J Exp Zool 281:417–427
Sumner AT (1972) A simple technique for demonstrating centromeric heterochromatin. Exp Cell Res 75:304–306
Valenzuela N (2007) Relic thermosensitive gene expression in a turtle with genotypic sex determination. Evolution 62:234–240
Valenzuela N, Lance V (eds) (2004) Temperature-dependent sex determination in vertebrates. Smithonian Books, Washington
Wong LH, Choo KHA (2001) Centromere on the move. Genome Res 11:513–516
Acknowledgements
We are grateful for Fengtang Yang and Patricia O'Brien, Department of Veterinary Medicine, Cambridge University, UK for providing chromosome-specific DNA probes of Pelodiscus sinensis. This work was supported by Grant-in-Aid for Scientific Research on Innovative Areas (No. 23113004) and Grant-in-Aid for Scientific Research (B) (No. 22370081) from the Ministry of Education, Culture, Sports, Science and Technology (MEXT), Japan.
Author information
Authors and Affiliations
Corresponding author
Additional information
Responsible Editors: Tariq Ezaz and Jennifer Graves.
Rights and permissions
About this article
Cite this article
Kawagoshi, T., Nishida, C. & Matsuda, Y. The origin and differentiation process of X and Y chromosomes of the black marsh turtle (Siebenrockiella crassicollis, Geoemydidae, Testudines). Chromosome Res 20, 95–110 (2012). https://doi.org/10.1007/s10577-011-9267-7
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10577-011-9267-7