Abstract
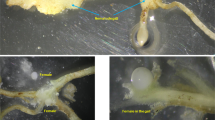
Root-knot nematodes (RKNs, Meloidogyne spp.) are damaging pests that can infect thousands of plant species and cause enormous crop losses worldwide. Panax notoginseng is a common host of root-knot nematodes. In this study, we surveyed notoginseng gardens and determined the incidence of RKNs. Among the gardens surveyed, 71 % were infected with RKNs, and the RKN incidence index ranged from 8 % to 47 % in three randomly infected gardens. Meloidogyne hapla was identified as the pathogenic nematode based on 18S ribosomal RNA analysis by DNA barcoding. The results were qualitatively and quantitatively confirmed using a real-time PCR assay according to variations in the ITS1 and ITS2 regions. These results indicated that the combination of DNA barcoding and real-time PCR is a reliable and precise method for identifying parasitic nematodes from mixed-infected plant roots in the field. In addition, the abundance of ITS1 and ITS2 displayed a similar trend to the numbers of RKNs in the three gardens, which suggests that the results of real-time PCR can be used to determine the damage caused by M. hapla in the field. Our studies show that RKNs are common and can cause serious damage to notoginseng. We present an integrated method of detecting mixed nematode species in the field and confirm M. hapla as the target for parasitic nematode control in notoginseng gardens. Our results contribute to the improvement of RKN control in notoginseng and further promote the sustainable development of medicinal plants.





Similar content being viewed by others
References
Abad, P., Gouzy, J., Aury, J. M., Castagnone-Sereno, P., Danchin, E. G. C., Deleury, E., et al. (2008). Genome sequence of the metazoan plant-parasitic nematode Meloidogyne incognita. Nature Biotechnology, 26, 909–915.
Bakhetia, M., Charlton, W., Atkinson, H. J., & McPherson, M. J. (2005). RNA interference of dual oxidase in the plant nematode Meloidogyne incognita. Molecular Plant-Microbe Interactions, 18, 1099–1106.
Berry, S. D., Fargette, M., Spaull, V. M., Morand, S., & Cadet, P. (2008). Detection and quantification of root-knot nematode (Meloidogyne javanica), lesion nematode (Pratylenchus zeae) and dagger nematode (Xiphinema elongatum) parasites of sugarcane using real-time PCR. Molecular and Cellular Probes, 22, 168–176.
Bhadury, P., & Austen, M. C. (2010). Barcoding marine nematodes: an improved set of nematode 18S rRNA primers to overcome eukaryotic co-interference. Hydrobiologia, 641, 245–251.
Bird, D. M., Opperman, C. H., & Williamson, V. M. (2009). Plant infection by root-knot nematode. In R. H. Berg & C. G. Taylor (Eds.), Cell biology of plant nematode parasitism. Berlin/Heidelberg: Springer.
Blaxter, M. L., De Ley, P., Garey, J. R., Liu, L. X., Scheldeman, P., Vierstraete, A., et al. (1998). A molecular evolutionary framework for the phylum Nematoda. Nature, 392, 71–75.
Chen, L. J., Wang, Y., Ke, J. H., & Zhou, J. M. (2002). Investigation of root-knot nematodes in Panax notoginseng. China Journal of Chinese Materia Medica, 27, 380–381.
De Ley, P., De Ley, T. I., Morris, K., Abebe, E., Mundo-Ocampo, M., Yoder, M., et al. (2005). An integrated approach to fast and informative morphological vouchering of nematodes for applications in molecular barcoding. Philosophical Transactions of the Royal Society of London. Series B, Biological Sciences, 360, 1945–1958.
De Oliveira, C. M. G., Montriro, A. R., & Blok, V. C. (2011). Morphological and molecular diagnostics for plant-parasitic nematodes: working together to get the identification done. Tropical Plant Pathology, 36, 65–73.
Derycke, S., Vanaverbeke, J., Rigaux, A., Backeljau, T., & Moens, T. (2010). Exploring the use of Cytochrome Oxidase c Subunit 1 (COI) for DNA barcoding of free-living marine nematodes. PLoS ONE, 5, e13716.
Dong, L. L., Huang, C. D., Huang, L., Li, X. L., & Zuo, Y. M. (2012). Screening plants resistant against Meloidogyne incognita and integrated management of plant resources for nematode control. Crop Protection, 33, 34–39.
Feng, G. Q., Dong, L. Y., Chen, Y. J., Shang, H., Liu, Y. Z., Li, J. R., et al. (2008). PCR detection of nematode isolated from Panax notoginseng. Southwest China Journal of Agricultural Sciences, 21, 100–102.
Fierer, N., Jackson, J. A., Vilgalys, R., & Jackson, R. B. (2005). Assessment of soil microbial community structure by use of Taxon-specific quantitative PCR assays. Applied and Environmental Microbiology, 71, 4117–4120.
Floyd, R., Abebe, E., Papert, A., & Blaxter, M. (2002). Molecular barcodes for soil nematode identification. Molecular Ecology, 11, 839–850.
Floyd, R. M., Rogers, A. D., Lambshead, P. J. D., & Smith, C. R. (2005). Nematode-specific PCR primers for the 18S small subunit rRNA gene. Molecular Ecology Notes, 5, 611–612.
Guo, R. J., Liu, X. Z., Li, S. D., & Miao, Z. Q. (2009). In vitro inhibition of fungal root-rot pathogens of Panax notoginseng by rhizobacteria. Plant Pathology Journal, 25, 70–76.
Hu, X. Q., Yang, Y. L., & Yu, S. F. (1998). Studies on the pathogen of root-knot disease on Panax notoginseng. Journal of Yunnan Agricultural University, 13, 375–379.
Hu, M. X., Zhuo, K., & Liao, J. L. (2011). Multiplex PCR for the simultaneous identification and detection of Meloidogyne incognita, M. enterolobill, and M. javanica using DNA extracted directly from individual galls. Phytopathology, 101, 1270–1277.
Jeanmougin, F., Thompson, J. D., Gouy, M., Higgins, D. G., & Gibson, T. J. (1998). Multiple sequence alignment with Clustal X. Trends in Biochemical Sciences, 23, 403–405.
Karssen, G., & Van Hoenselaar, T. (1998). Revision of the genus Meloidogyne Göldi, 1892(Nematode: Heteroderidae) in Europe. Nematologica, 44, 713–788.
Min, Y. Y., Toyota, K., Goto, K., Sato, E., Mizuguchi, S., Abe, N., et al. (2011). Development of a direct quantitative detection method for Meloidogyne incognita in sandy soils and its application to sweet potato cultivated fields in Tokushima prefecture, Japan. Nematology, 13, 95–102.
Murphy, L. L., & Lee, T. J. (2002). Ginseng, sex behavior, and nitric oxide. Annals of the New York Academy of Sciences, 962, 372–377.
Oiu, J. J., Westerdahl, B. B., Anderson, C., & Williamson, V. M. (2006). Sensitive PCR detection of Meloidogyne arenaria, M. incognita and M. javanica extracted from soil. Journal of Nematology, 38, 434–441.
Oka, Y., & Mizukubo, T. (2008). Tomato culture filtrate stimulates hatching and activity of Meloidogyne incognita juveniles. Nematology, 11, 51–61.
Onkendi, E. M., & Moleleki, L. N. (2013). Detection of Meloidogyne enterolobii in potatoes in South Africa and phylogenetic analysis based on intergenic region and the mitochondrial DNA sequences. European Journal of Plant Pathology, doi:10.1007/s10658-012-0142-y.
Powers, T. O., Neher, D. A., Mullin, P., Esquivel, A., Giblin-Davis, R. M., Kanzaki, N., et al. (2009). Tropical nematode diversity: vertical stratification of nematode communities in a Costa Rican humid lowland rainforest. Molecular Ecology, 18, 1–12.
Rousk, J., Bååth, E., Brookes, P. C., Lauber, C. L., Catherine, L., Caporaso, J. G., et al. (2010). Soil bacterial and fungal communities across a pH gradient in an arable soil. The ISME Journal, 4, 1–12.
Sasser, J. N. (1980). Root-knot nematodes: a global menace to crop production. Plant Disease, 64, 36–41.
Sasser, J. N., Eisenback, J. D., & Carter, C. C. (1983). The international Meloidogyne project-its goals and accomplishments. Annual Review of Phytopathology, 21, 271–288.
Sayed Abdul Rahman, S. A., Mohamed, Z., Othman, R. Y., Swennen, R., Panis, B., De Waele, D., et al. (2010). In planta PCR-based detection of early infection of plant-parasitic nematodes in the roots: a step towards the understanding of infection and plant defence. European Journal of Plant Pathology, 128, 343–351.
Siddall, M. E., Kvist, S., Phillips, A., & Oceguera-Figuero, A. (2012). DNA barcoding of parasitic nematodes: is it kosher? Journal of Parasitology, 98, 692–694.
Tamura, K., Peterson, D., Peterson, N., Stecher, G., Nei, M., & Kumar, S. (2011). MEGA5: molecular evolutionary genetics analysis using maximum likelihood, evolutionary distance, and maximum. Molecular Biology and Evolution, 28, 2731–2739.
Thompson, J. D., Gibson, T. J., Plewniak, F., Jeanmougin, F., & Higgins, D. G. (1997). The CLUSTAL_X windows interface: flexible strategies for multiple sequence alignment aided by quality analysis tools. Nucleic Acids Research, 25, 4876–4882.
Toyota, K., Shirakashi, T., Sato, E., Wada, S., & Min, Y. Y. (2008). Development of a real-time PCR method for the potato-cyst nematode Globodera rostochiensis and the root-knot nematode Meloidogyne incognita. Soil Science and Plant Nutrition, 54, 72–76.
Trudgill, D. L., & Blok, V. C. (2001). Apomictic, polyphagous root-knot nematodes: exceptionally successful and damaging root pathogens. Annual Review of Phytopathology, 39, 53–77.
Tsay, T. T., Wu, S. T., & Lin, Y. Y. (2004). Evaluation of asteraceae plants for control of Meloidogyne incognita. Journal of Nematology, 36, 36–41.
Vovlas, N., Trisciuzzi, N., Troccoli, A., De Luca, F., Cantalapiedra-Navarrete, C., & Castillo, P. (2013). Integrative diagnosis and parasitic habits of Cryphodera brinkmani a non-cyst forming heteroderid nematode intercepted on Japanese white pine bonsai trees imported into Italy. European Journal of Plant Pathology, 135, 717–726.
Vrain, T. C., Wakarchuk, D. A., Levesque, A. C., & Hamilton, R. I. (1992). Intraspecific rDNA restriction fragment length polymorphism in the Xiphinema americanum group. Fundamental and Applied Nematology, 15, 563–573.
Yang, P. W., Cui, X. M., Dong, L. Y., Jin, L., Chen, Y. J., Liu, S. F., et al. (2008). Identification of root-knot nematode harmful to Panax notoginseng and its distribution. Journal of Yunnan Agricultural University, 23, 479–482.
Yeh, G. Y., Eisenberg, D. M., Kaptchuk, T. J., & Phillips, R. S. (2003). Systematic review of herbs and dietary supplements for glycemic control in diabetes. Diabetes Care, 26, 1277–1294.
Ying, Y. X., Ding, W. L., & Li, Y. (2012). Characterization of soil bacterial communities in rhizospheric and nonrhizospheric soil of Panax ginseng. Biochemical Genetics, 50, 848–859.
Zijlstra, C. (1997). A fast PCR assay to identify Meloidogyne hapla, M. chitwoodi, and M. fallax, and to sensitively differentiate them from each other and from M. incognita in mixtures. Fundamental and Applied Nematology, 20, 505–511.
Zijlstra, C., & Van Hoof, R. A. (2006). A multiplex real-time polymerase chain reaction (TaqMan) assay for the simultaneous detection of Meloidogyne chiwoodi and M. fallax. Phytopathology, 96, 1255–1262.
Acknowledgments
This work was supported by the 52nd General Financial Grant from the China Postdoctoral Science Foundation (grant no. 2012M520203), the Key National Natural Science Foundation of China (grant no. 81130069), the National Key Technology R&D Program (grant no. 2012BAI29B01) and the Program for Changjiang Scholars and Innovative Research Team in the University of the Ministry of Education of China (grant no. IRT1150).
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Dong, L., Yao, H., Li, Q. et al. Investigation and integrated molecular diagnosis of root-knot nematodes in Panax notoginseng root in the field. Eur J Plant Pathol 137, 667–675 (2013). https://doi.org/10.1007/s10658-013-0277-5
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10658-013-0277-5