Abstract
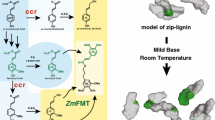
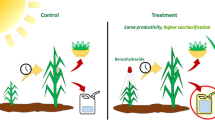
Maize (Zea mays) caffeate 3-O-methyltransferase (ZmaysCOMT, EC 2.1.1.68), a key enzyme of the phenylpropanoid pathway, catalyzes the O-methylation of caffeic acid to ferulic acid, a precursor of lignin polymer and a crucial component of the cell wall structure. Plant cell wall recalcitrance is due to lignin, and the discovery of specific inhibitors of ZmaysCOMT could be useful to increase the digestibility of the lignocellulose biomass and improve the production of cellulosic biofuels. In this work, we have modeled the three-dimensional structure of ZmaysCOMT and prospected promising inhibitors by using virtual screening techniques. A set of 1668 putative candidates was screened from a virtual library and docked in the active site of the enzyme, and nitecapone was selected as one of the most promising enzyme inhibitors. Details of the mode of inhibition were assessed by in silico simulation and in vitro assays of nitecapone on the enzyme. In comparison with the nitecapone-free control, kinetics parameters showed different values of Vmax and KM, suggesting a kinetic profile such as mixed inhibition of the ZmaysCOMT. In brief, we suggest that the nitecapone-induced inhibition of ZmaysCOMT may serve as a non-transgenic strategy to explore the biosynthesis of ferulic acid and lignin, their relationships with the recalcitrance of lignocellulosic biomass, and, possibly, to improve bioethanol production.







Similar content being viewed by others
References
Boerjan W, Ralph J, Baucher M (2003) Lignin biosynthesis. Annu Rev Plant Biol 54:519–546. https://doi.org/10.1146/annurev.arplant.54.031902.134938
Bonawitz ND, Chapple C (2013) Can genetic engineering of lignin deposition be accomplished without an unacceptable yield penalty? Curr Opin Biotechnol 24:336–343. https://doi.org/10.1016/j.copbio.2012.11.004
Bonifácio MJ, Palma PN, Almeida L, Soares-da-Silva P (2007) Catechol-O-methyltransferase and its inhibitors in Parkinson’s disease. CNS Drug Rev 13:352–379. https://doi.org/10.1111/j.1527-3458.2007.00020.x
Borges N, Vieira-Coelho MA, Parada A, Soares-da-Silva P (1997) Studies on the tight-binding nature of tolcapone inhibition of soluble and membrane-bound rat brain catechol-O-methyltransferase. J Pharmacol Exp Ther 282:812–817
Bradford MM (1976) A rapid and sensitive method for the quantitation of microgram quantities of protein utilizing the principle of protein-dye binding. Anal Biochem 72:248–254. https://doi.org/10.1016/0003-2697(76)90527-3
Bubna GA, Lima RB, Zanardo DYL, dos Santos WD, Ferrarese MLL, Ferrarese-Filho O (2011) Exogenous caffeic acid inhibits the growth and enhances the lignification of the roots of soybean (Glycine max). J Plant Physiol 168:1627–1633. https://doi.org/10.1016/j.jplph.2011.03.005
Calvo-Flores FG, Dobado JA, Isac-Garcia J, Martín-Martínez FJ (2015) Lignin and lignans as renewable raw materials. Chemistry, Technology and Applications. John Wiley & Sons, New York
Carpita NC (2012) Progress in the biological synthesis of the plant cell wall: new ideas for improving biomass for bioenergy. Curr Opin Biotechnol 23:330–337. https://doi.org/10.1016/j.copbio.2011.12.003
CCP4 (1994) The CCP4 suite: programs for protein crystallography. Acta Crystallogr Sect D Biol Crystallogr 50:760–763. https://doi.org/10.1107/S0907444994003112
Chen F, Dixon RA (2007) Lignin modification improves fermentable sugar yields for biofuel production. Nat Biotechnol 25:759–761. https://doi.org/10.1038/nbt1316
Collazo P, Montoliu L, Puigdomènech P, Rigau J (1992) Structure and expression of the lignin O-methyltransferase gene from Zea mays L. Plant Mol Biol 20:857–867. https://doi.org/10.1007/BF00027157
Copeland RA (2013) Evaluation of enzyme inhibitors in drug discovery: a guide for medicinal chemists and pharmacologists, 2nd edn. Wiley-Interscience, New York
Dong J, Wu F, Zhang G (2006) Influence of cadmium on antioxidant capacity and four microelement concentrations in tomato seedlings (Lycopersicon esculentum). Chemosphere 64:1659–1666. https://doi.org/10.1016/j.chemosphere.2006.01.030
dos Santos WD, Ferrarese MLL, Nakamura CV, Mourão KSM, Mangolin CA, Ferrarese-Filho O (2008) Soybean (Glycine max) root lignification induced by ferulic acid. The possible mode of action. J Chem Ecol 34:1230–1241. https://doi.org/10.1007/s10886-008-9522-3
Eswar N, Webb B, Marti-Renom MA, Madhusudhan MS, Eramian E, Shen M-Y, Pieper U, Sali A (2014) Comparative protein structure modeling using Modeller. Curr Protoc Bioinformatics 15:5.6.1–5.6.30
Eudes A, Liang Y, Mitra P, Loqué D (2014) Lignin bioengineering. Curr Opin Biotechnol 26:189–198. https://doi.org/10.1016/j.copbio.2014.01.002
Ferrarese MLL, Rodrigues JD, Ferrarese-Filho O (2000) Phenylalanine ammonia-lyase activity in soybean roots extract measured by reverse-phase high performance liquid chromatography. Plant Biol 2:152–153. https://doi.org/10.1055/s-2000-9162
Fornalé S, Rencoret J, García-Calvo L, Encina A, Rigau J, Gutiérrez A, del Río JC, Caparros-Ruiz D (2017) Changes in cell wall polymers and degradability in maize mutants lacking 3- and 5-O-methyltransferases involved in lignin biosynthesis. Plant Cell Physiol 58:240–255. https://doi.org/10.1093/pcp/pcw198
Ganellin CR, Triggle DJ (1996) Dictionary of pharmacological agents. Chapman and Hall/CRC, Portlan
Grabber JH, Ralph J, Lapierre C, Barrière Y (2004) Genetic and molecular basis of grass cell-wall degradability. I. Lignin-cell wall matrix interactions. C R Biol 327:455–465
Homem DP, Flores R Jr, Tosqui P, Rozada TC, Basso EA, Gasparotto A Jr, Seixas FAV (2013) Homology modeling of dihydrofolate reductase from T. gondii bonded to antagonists: molecular docking and molecular dynamics simulations. Mol BioSyst 9:1308–1315. https://doi.org/10.1039/c3mb25530a
Inoue K, Sewalt VJ, Murray GB, Ni W, Stürzer C, Dixon RA (1998) Developmental expression and substrate specificities of alfalfa caffeic acid 3-O-methyltransferase and caffeoyl coenzyme A 3-O-methyltransferase in relation to lignification. Plant Physiol 117:761–770. https://doi.org/10.1104/pp.117.3.761
Irwin JJ, Sterling T, Mysinger MM, Bolstad ES, Coleman RG (2012) ZINC: a free tool to discover chemistry for biology. J Chem Inf Model 52:1757–1768. https://doi.org/10.1021/ci3001277
Krissinel E, Henrick K (2007) Inference of macromolecular assemblies from crystalline state. J Mol Biol 372:774–797. https://doi.org/10.1016/j.jmb.2007.05.022
Lima RB, Salvador VH, dos Santos WD, Bubna GA, Finger-Teixeira A, Soares AR, Marchiosi R, Ferrarese MLL, Ferrarese-Filho O (2013) Enhanced lignin monomer production caused by cinnamic acid and its hydroxylated derivatives inhibits soybean root growth. PLoS One 8:e80542. https://doi.org/10.1371/journal.pone.0080542
Liu T, Lin Y, Wen X, Jorissen RN, Gilson MK (2007) BindingDB: a web-accessible database of experimentally determined protein-ligand binding affinities. Nucleic Acids Res 35:198–201. https://doi.org/10.1093/nar/gkl999
Lobley A, Sadowski MI, Jones DT (2009) pGenTHREADER and pDomTHREADER: new methods for improved protein fold recognition and superfamily discrimination. Bioinformatics 25:1761–1767. https://doi.org/10.1093/bioinformatics/btp302
Lotta T, Vidgren J, Tilgmann C, Ulmanen I, Melén K, Julkunen I, Taskinen J (1995) Kinetics of human soluble and membrane-bound catechol O-methyltransferase: a revised mechanism and description of the thermolabile variant of the enzyme. Biochemistry 34:4202–4210. https://doi.org/10.1021/bi00013a008
Louie GV, Bowman ME, Tu Y, Mouradov A, Spangenberg G, Noel JP (2010) Structure-function analyses of a caffeic acid O-methyltransferase from perennial ryegrass reveal the molecular basis for substrate preference. Plant Cell 22:4114–4127. https://doi.org/10.1105/tpc.110.077578
Mackerell AD, Feig M, Brooks CL (2004) Extending the treatment of backbone energetics in protein force fields: limitations of gas-phase quantum mechanics in reproducing protein conformational distributions in molecular dynamics simulation. J Comput Chem 25:1400–1415. https://doi.org/10.1002/jcc.20065
Marchiosi R, dos Santos WD, Constantin RP, de Lima RB, Soares AR, Finger-Teixeira A, Mota TR, de Oliveira DM, Foletto-Felipe MP, Abrahão J, Ferrarese-Filho O (2020) Biosynthesis and metabolic actions of simple phenolic acids in plants. Phytochem Rev. https://doi.org/10.1007/s11101-020-09689-2
Marriott PE, Gómez LD, McQueen-Mason SJ (2016) Tansley review unlocking the potential of lignocellulosic biomass through plant science. New Phytol 209:1366–1381. https://doi.org/10.1111/nph.13684
Meents MJ, Watanabe Y, Samuels AL (2018) The cell biology of secondary cell wall biosynthesis. Ann Bot 121:1107–1125. https://doi.org/10.1093/aob/mcy005
Miteva MA, Violas S, Montes M, Gomez D, Tuffery P, Villoutreix BO (2006) FAF-drugs: free ADME/tox filtering of compound collections. Nucleic Acids Res 34:738–744. https://doi.org/10.1093/nar/gkl065
Moinuddin SGA, Jourdes M, Laskar DD, Ki C, Cardenas CL, Kim KW, Zhang D, Davin LB, Lewis NG (2010) Insights into lignin primary structure and deconstruction from Arabidopsis thaliana COMT (caffeic acid O-methyl transferase) mutant Atomt1. Org Biomol Chem 8:3928–3946. https://doi.org/10.1039/c004817h
Mottiar Y, Vanholme R, Boerjan W, Ralph J, Mansfield SD (2016) Designer lignins: harnessing the plasticity of lignification. Curr Opin Biotechnol 37:190–200. https://doi.org/10.1016/j.copbio.2015.10.009
Oliveira DM, Finger-Teixeira A, Mota TR, Salvador VH, Moreira-Vilar FC, Molinari HBC, Mitchell RAC, Marchiosi R, Ferrarese-Filho O, dos Santos WD (2015) Ferulic acid: a key component in grass lignocellulose recalcitrance to hydrolysis. Plant Biotechnol J 13:1224–1232. https://doi.org/10.1111/pbi.12292
Phillips JC, Braun R, Wang W, Gumbart J, Tajkhorshid E, Villa E, Chipot C, Skeel RD, Kalé L, Schulten K (2005) Scalable molecular dynamics with NAMD. J Comput Chem 26:1781–1802. https://doi.org/10.1002/jcc.20289
Quevillon E, Silventoinen V, Pillai S, Harte N, Mulder N, Apweiler R, Lopez R (2005) InterProScan: protein domains identifier. Nucleic Acids Res 33:116–120. https://doi.org/10.1093/nar/gki442
Rost (1999) Twilight zone of protein sequence alignments. Protein Eng 12:85–94
Salvador VH, Lima RB, dos Santos WD, Soares AR, Böhm PAF, Marchiosi R, Ferrarese MLL, Ferrarese-Filho O (2013) Cinnamic acid increases lignin production and inhibits soybean root growth. PLoS One 8:e80542. https://doi.org/10.1371/journal.pone.0069105
Schomburg I, Chang A, Placzek S, Söhngen C, Rother M, Lang M, Munaretto C, Ulas S, Stelzer M, Grote A, Scheer M, Schomburg D (2013) BRENDA in 2013: integrated reactions, kinetic data, enzyme function data, improved disease classification: new options and contents in BRENDA. Nucleic Acids Res 41:764–772. https://doi.org/10.1093/nar/gks1049
Schultz E, Nissinen E (1989) Inhibition of rat liver and duodenum soluble catechol-O-methyltransferase by a tight-binding inhibitor OR-462. Biochem Pharmacol 38:3953–3956. https://doi.org/10.1016/0006-2952(89)90673-4
Seixas FAV, Santini TD, Moura VP, Gandra EA (2008) Evaluation of the (haem)Fe-Nε2(HisF8) bond distances from haemoglobin structures deposited in the Protein Data Bank. Acta Crystallogr Sect D Biol Crystallogr 64:971–976. https://doi.org/10.1107/S0907444908022208
Shimada M, Kuroda H, Higuchi T (1973) Evidence for the formation of methoxyl groups of ferulic and sinapic acids in Bambusa by the same O-methyltransferase. Phytochemistry 12:2873–2875. https://doi.org/10.1016/0031-9422(73)80498-4
Shinde SD, Meng X, Kumar R, Ragauskas AJ (2018) Recent advances in understanding the pseudo-lignin formation in a lignocellulosic biorefinery. Green Chem 20:2192–2205. https://doi.org/10.1039/C8GC00353J
Simmons BA, Loqué D, Ralph J (2010) Advances in modifying lignin for enhanced biofuel production. Curr Opin Plant Biol 13:313–320. https://doi.org/10.1016/j.pbi.2010.03.001
Souza WR, Martins PK, Freeman J et al (2018) Suppression of a single BAHD gene in Setaria viridis causes large, stable decreases in cell wall feruloylation and increases biomass digestibility. New Phytol 218:81–93. https://doi.org/10.1111/nph.14970
Tsuji E, Okazaki K, Isaji M, Takeda K (2009) Crystal structures of the Apo and Holo form of rat catechol-O-methyltransferase. J Struct Biol 165:133–139. https://doi.org/10.1016/j.jsb.2008.11.012
Umezawa T (2018) Lignin modification in planta for valorization. Phytochem Rev 17:1305–1327. https://doi.org/10.1007/s11101-017-9545-x
Van Acker R, Vanholme R, Storme V, Mortimer JC, Dupree P, Boerjan W (2013) Lignin biosynthesis perturbations affect secondary cell wall composition and saccharification yield in Arabidopsis thaliana. Biotechnol Biofuels 6:46. https://doi.org/10.1186/1754-6834-6-46
Van Acker R, Déjardin A, Desmet S (2017) Different metabolic routes for coniferaldehyde and sinapaldehyde with cinnamyl alcohol dehydrogenase1 deficiency. Plant Physiol 175:1018–1039. https://doi.org/10.1104/pp.17.00834
Yang AS, Honig B (2000) An integrated approach to the analysis and modeling of protein sequences and structures. III. A comparative study of sequence conservation in protein structural families using multiple structural alignments. J Mol Biol 301:691–711. https://doi.org/10.1006/jmbi.2000.3975
Zanardo DIL, Lima RB, Ferrarese MLL, Bubna GA, Ferrarese-Filho O (2009) Soybean root growth inhibition and lignification induced by p-coumaric acid. Environ Exp Bot 66:25–30. https://doi.org/10.1016/j.envexpbot.2008.12.014
Zoete V, Cuendet MA, Grosdidier A, Michielin O (2011) SwissParam: a fast force field generation tool for small organic molecules. J Comput Chem 32:2359–2368. https://doi.org/10.1002/jcc.21816
Zubieta C, Kota P, Ferrer J, Dixon RA, Noel JP (2002) Structural basis for the modulation of lignin monomer methylation by caffeic acid/5-hydroxyferulic acid 3/5-O-methyltransferase. Plant Cell 14:1265–1277. https://doi.org/10.1105/tpc.001412.nylpropanoids
Acknowledgments
A.V. Parizotto was a recipient of Coordination of Enhancement of Higher Education Personnel (CAPES, code 001) fellowship. A.P. Ferro was a recipient of The Brazilian Council for Scientific and Technological Development (CNPq/PNPD) fellowship. O. Ferrarese-Filho and R. Marchiosi are research fellows of CNPq. The authors also acknowledge the National Center for High Performance Processing (CENAPAD, Brazil) for providing computational facilities.
Funding
This study was funded by The Brazilian Council for Scientific and Technological Development (Grants n° 477075/2011-8) and Araucaria Foundation (Grants n° 20133960, 40/16 and 53/19).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of Interest
The authors declare that they have no conflict of interest.
Ethical Approval
This article does not contain any studies with human participants performed by any of the authors.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Key Message
• Three-dimensional structure of maize (Zea mays) caffeate 3-O-methyltransferase was modeled
• A set of 1668 putative inhibitors of ZmaysCOMT was screened from a virtual library
• Nitecapone was selected as one of most promising enzyme inhibitors
• A mixed inhibition of the ZmaysCOMT was obtained with nitecapone
Electronic supplementary material
ESM 1
(DOCX 984 kb)
Rights and permissions
About this article
Cite this article
Parizotto, A.V., Ferro, A.P., Marchiosi, R. et al. Inhibition of Maize Caffeate 3-O-Methyltransferase by Nitecapone as a Possible Approach to Reduce Lignocellulosic Biomass Recalcitrance. Plant Mol Biol Rep 39, 179–191 (2021). https://doi.org/10.1007/s11105-020-01242-x
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11105-020-01242-x