Abstract
Introduction
Amyotrophic lateral sclerosis (ALS) and Parkinson’s disease (PD) are two severe neurodegenerative disorders for which the disease mechanisms are poorly understood and reliable biomarkers are absent.
Objectives
To identify metabolite biomarkers for ALS and PD, and to gain insights into which metabolic pathways are involved in disease.
Methods
Nuclear magnetic resonance (NMR) metabolomics was utilized to characterize the metabolite profiles of cerebrospinal fluid (CSF) and plasma from individuals in three age, gender, and sampling-date matched groups, comprising 22 ALS, 22 PD and 28 control subjects.
Results
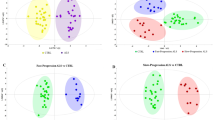
Multivariate analysis of NMR data generated robust discriminatory models for separation of ALS from control subjects. ALS patients showed increased concentrations of several metabolites in both CSF and plasma, these are alanine (CSF fold change = 1.22, p = 0.005), creatine (CSF-fc = 1.17, p = 0.001), glucose (CSF-fc = 1.11, p = 0.036), isoleucine (CSF-fc = 1.24, p = 0.002), and valine (CSF-fc = 1.17, p = 0.014). Additional metabolites in CSF (creatinine, dimethylamine and lactic acid) and plasma (acetic acid, glutamic acid, histidine, leucine, pyruvate and tyrosine) were also important for this discrimination. Similarly, panels of CSF-metabolites that discriminate PD from ALS and control subjects were identified.
Conclusions
The results for the ALS patients suggest an affected creatine/creatinine pathway and an altered branched chain amino acid (BCAA) metabolism, and suggest links to glucose and energy metabolism. Putative metabolic markers specific for ALS (e.g. creatinine and lactic acid) and PD (e.g. 3-hydroxyisovaleric acid and mannose) were identified, while several (e.g. creatine and BCAAs) were shared between ALS and PD, suggesting some overlap in metabolic alterations in these disorders.




Similar content being viewed by others
Abbreviations
- ALS:
-
Amyotrophic lateral sclerosis
- PD:
-
Parkinson’s disease
- NMR:
-
Nuclear magnetic resonance
- MS:
-
Mass spectrometry
- EDTA:
-
Ethylenediaminetetraacetic acid
- CSF:
-
Cerebrospinal fluid
- CPMG:
-
Carr-Purcell-Meiboom-Gill
- PCA:
-
Principal component analysis
- OPLS:
-
Orthogonal projection to latent structures
- DA:
-
Discriminant analysis
- EP:
-
Effect projection
- CV-ANOVA:
-
Analysis of variance testing of cross-validated predictive residuals
- CNS:
-
Central nervous system
- BCAA:
-
Branched chain amino acids
References
Andersen, P. M., Abrahams, S., Borasio, G. D., de Carvalho, M., Chio, A., Hardiman, O., et al. (2012). EFNS guidelines on the clinical management of amyotrophic lateral sclerosis (MALS)–revised report of an EFNS task force. European Journal of Neurology, 19(3), 360–375. doi:10.1111/j.1468-1331.2011.03501.x.
Aranibar, N., Ott, K. H., Roongta, V., & Mueller, L. (2006). Metabolomic analysis using optimized NMR and statistical methods. Analytical Biochemistry, 355(1), 62–70. doi:10.1016/j.ab.2006.04.014.
Averous, J., Lambert-Langlais, S., Carraro, V., Gourbeyre, O., Parry, L., B’Chir, W., et al. (2014). Requirement for lysosomal localization of mTOR for its activation differs between leucine and other amino acids. Cellular Signalling, 26(9), 1918–1927. doi:10.1016/j.cellsig.2014.04.019.
Barton, R. H., Waterman, D., Bonner, F. W., Holmes, E., Clarke, R., Procardis, C., et al. (2010). The influence of EDTA and citrate anticoagulant addition to human plasma on information recovery from NMR-based metabolic profiling studies. Molecular BioSystems, 6(1), 215–224. doi:10.1039/b907021d.
Beghi, E. (2013). Are professional soccer players at higher risk for ALS? Amyotroph Lateral Scler Frontotemporal Degener, 14(7–8), 501–506. doi:10.3109/21678421.2013.809764.
Blasco, H., Corcia, P., Moreau, C., Veau, S., Fournier, C., Vourc’h, P., et al. (2010). 1H-NMR-based metabolomic profiling of CSF in early amyotrophic lateral sclerosis. PLoS ONE, 5(10), e13223. doi:10.1371/journal.pone.0013223.
Blasco, H., Corcia, P., Pradat, P. F., Bocca, C., Gordon, P. H., Veyrat-Durebex, C., et al. (2013). Metabolomics in cerebrospinal fluid of patients with amyotrophic lateral sclerosis: an untargeted approach via high-resolution mass spectrometry. Journal of Proteome Research, 12(8), 3746–3754. doi:10.1021/pr400376e.
Bouatra, S., Aziat, F., Mandal, R., Guo, A. C., Wilson, M. R., Knox, C., et al. (2013). The human urine metabolome. PLoS ONE, 8(9), e73076. doi:10.1371/journal.pone.0073076.
Bylesjö, M., Rantalainen, M., Cloarec, O., Nicholson, J. K., Holmes, E., & Trygg, J. (2006). OPLS discriminant analysis: combining the strengths of PLS-DA and SIMCA classification. Journal of Chemometrics, 20(8–10), 341–351. doi:10.1002/cem.1006.
Dame, Z. T., Aziat, F., Mandal, R., Krishnamurthy, R., Bouatra, S., Borzouie, S., et al. (2015). The human saliva metabolome. Metabolomics, 11(6), 1864–1883. doi:10.1007/s11306-015-0840-5.
Dunn, W. B., Broadhurst, D. I., Atherton, H. J., Goodacre, R., & Griffin, J. L. (2011). Systems level studies of mammalian metabolomes: the roles of mass spectrometry and nuclear magnetic resonance spectroscopy. Chemical Society Reviews, 40(1), 387–426. doi:10.1039/b906712b.
Engelke, U. F., Tangerman, A., Willemsen, M. A., Moskau, D., Loss, S., Mudd, S. H., et al. (2005). Dimethyl sulfone in human cerebrospinal fluid and blood plasma confirmed by one-dimensional (1)H and two-dimensional (1)H-(13)C NMR. NMR in Biomedicine, 18(5), 331–336. doi:10.1002/nbm.966.
Eriksson, L., Trygg, J., & Wold, S. (2008). CV-ANOVA for significance testing of PLS and OPLS (R) models. Journal of Chemometrics, 22(11–12), 594–600. doi:10.1002/cem.1187.
Ekhtiari Bidhendi, E., Bergh, J., Zetterström, P., Andersen, P. M., Marklund, S. L., & Brännström, T. (2016). Two superoxide dismutase prion strains transmitting amyotrophic lateral sclerosis. The Journal of Investigation (in press).
Fahn, S., Elton, R., & Commitee TUD. (1987). Unified Parkinson’s disease rating scale. In S. Fahn, C. Marsden, D. Calne, & M. Goldstein (Eds.), Recent developments in Parkinson’s disease (pp. 153–163). Florham Park: Macmillan Healthcare Information.
Fernstrom, J. D. (2013). Large neutral amino acids: dietary effects on brain neurochemistry and function. Amino Acids, 45(3), 419–430. doi:10.1007/s00726-012-1330-y.
Forsberg, K., Jonsson, P. A., Andersen, P. M., Bergemalm, D., Graffmo, K. S., Hultdin, M., et al. (2010). Novel antibodies reveal inclusions containing non-native SOD1 in sporadic ALS patients. PLoS ONE, 5(7), e11552. doi:10.1371/journal.pone.0011552.
Gall, W. E., Beebe, K., Lawton, K. A., Adam, K. P., Mitchell, M. W., Nakhle, P. J., et al. (2010). Alpha-hydroxybutyrate is an early biomarker of insulin resistance and glucose intolerance in a nondiabetic population. PLoS ONE, 5(5), e10883. doi:10.1371/journal.pone.0010883.
Gibb, W. R., & Lees, A. J. (1988). A comparison of clinical and pathological features of young- and old-onset Parkinson’s disease. Neurology, 38(9), 1402–1406.
Goedert, M. (2001). Alpha-synuclein and neurodegenerative diseases. Nature Reviews Neuroscience, 2(7), 492–501. doi:10.1038/35081564.
Gray, E., Larkin, J. R., Claridge, T. D., Talbot, K., Sibson, N. R., & Turner, M. R. (2015). The longitudinal cerebrospinal fluid metabolomic profile of amyotrophic lateral sclerosis. Amyotroph Lateral Scler Frontotemporal Degener, 16(1–8), 456–463. doi:10.3109/21678421.2015.1053490.
Holmqvist, S., Chutna, O., Bousset, L., Aldrin-Kirk, P., Li, W., Bjorklund, T., et al. (2014). Direct evidence of Parkinson pathology spread from the gastrointestinal tract to the brain in rats. Acta Neuropathologica, 128(6), 805–820. doi:10.1007/s00401-014-1343-6.
Isobe, C., Abe, T., & Terayama, Y. (2010). Decrease in asymmetrical dimethylarginine, an endogenous nitric oxide synthase inhibitor, in cerebrospinal fluid during elderly aging and in patients with sporadic form of amyotrophic lateral sclerosis. Neurosignals, 18(1), 43–48. doi:10.1159/000312527.
Jewell, J. L., Kim, Y. C., Russell, R. C., Yu, F. X., Park, H. W., Plouffe, S. W., et al. (2015). Metabolism differential regulation of mTORC1 by leucine and glutamine. Science, 347(6218), 194–198. doi:10.1126/science.1259472. (New York).
Jonsson, P., Wuolikainen, A., Thysell, E., Chorell, E., Stattin, P., Wikström, P., et al. (2015). Constrained randomization and multivariate effect projections improve information extraction and biomarker pattern discovery in metabolomics studies involving dependent samples. Metabolomics, 11(6), 1667–1678. doi:10.1007/s11306-015-0818-3.
Jove, M., Portero-Otin, M., Naudi, A., Ferrer, I., & Pamplona, R. (2014). Metabolomics of human brain aging and age-related neurodegenerative diseases. Journal of Neuropathology and Experimental Neurology, 73(7), 640–657. doi:10.1097/NEN.0000000000000091.
Jupin, M., Michiels, P. J., Girard, F. C., Spraul, M., & Wijmenga, S. S. (2013). NMR identification of endogenous metabolites interacting with fatted and non-fatted human serum albumin in blood plasma: fatty acids influence the HSA-metabolite interaction. J Mag Reson, 228, 81–94. (San Diego, Calif: 1997). doi:10.1016/j.jmr.2012.12.010.
Jupin, M., Michiels, P. J., Girard, F. C., Spraul, M., & Wijmenga, S. S. (2014). NMR metabolomics profiling of blood plasma mimics shows that medium- and long-chain fatty acids differently release metabolites from human serum albumin. J Mag Reson, 239, 34–43. (San Diego, Calif: 1997). doi:10.1016/j.jmr.2013.11.019.
Keun, H. C., Ebbels, T. M., Antti, H., Bollard, M. E., Beckonert, O., Schlotterbeck, G., et al. (2002). Analytical reproducibility in (1)H NMR-based metabonomic urinalysis. Chemical Research in Toxicology, 15(11), 1380–1386. doi:10.1021/tx0255774.
Kiernan, M. C., Vucic, S., Cheah, B. C., Turner, M. R., Eisen, A., Hardiman, O., et al. (2011). Amyotrophic lateral sclerosis. Lancet, 377(9769), 942–955. doi:10.1016/S0140-6736(10)61156-7.
Knowles, T. P. J., Vendruscolo, M., & Dobson, C. M. (2014). The amyloid state and its association with protein misfolding diseases. Nature Reviews Molecular Cell Biology, 15(6), 384–396. doi:10.1038/nrm3810.
Kumar, A., Bala, L., Kalita, J., Misra, U. K., Singh, R. L., Khetrapal, C. L., et al. (2010). Metabolomic analysis of serum by (1) H NMR spectroscopy in amyotrophic lateral sclerosis. Clinica Chimica Acta, 411(7–8), 563–567. doi:10.1016/j.cca.2010.01.016.
Lakke, J. P., & Teelken, A. W. (1976). Amino acid abnormalities in cerebrospinal fluid of patients with parkinsonism and extrapyramidal disorders. Neurology, 26(5), 489–493.
Lakke, J. P., Teelken, A. W., vd Voet, H., & Wolthers, B. (1987). Amino acid abnormalities in cerebrospinal fluid and blood serum of patients with Parkinson’s disease, other heredodegenerative disorders and head injuries. Advances in Neurology, 45, 243–247.
Lang, A. E., & Obeso, J. A. (2004). Challenges in Parkinson’s disease: restoration of the nigrostriatal dopamine system is not enough. Lancet Neurology, 3(5), 309–316. doi:10.1016/S1474-4422(04)00740-9.
Lawton, K. A., Cudkowicz, M. E., Brown, M. V., Alexander, D., Caffrey, R., Wulff, J. E., et al. (2012). Biochemical alterations associated with ALS. Amyotroph Lateral Scler, 13(1), 110–118. doi:10.3109/17482968.2011.619197.
Lawton, K. A., Brown, M. V., Alexander, D., Li, Z., Wulff, J. E., Lawson, R., et al. (2014). Plasma metabolomic biomarker panel to distinguish patients with amyotrophic lateral sclerosis from disease mimics. Amyotroph Lateral Scler Frontotemporal Degener, 15(5–6), 362–370. doi:10.3109/21678421.2014.908311.
Lewitt, P. A., Li, J., Lu, M., Beach, T. G., Adler, C. H., Guo, L., et al. (2013). 3-hydroxykynurenine and other Parkinson’s disease biomarkers discovered by metabolomic analysis. Movement Disorders, 28(12), 1653–1660. doi:10.1002/mds.25555.
Li, J. V., Ashrafian, H., Bueter, M., Kinross, J., Sands, C., le Roux, C. W., et al. (2011). Metabolic surgery profoundly influences gut microbial-host metabolic cross-talk. Gut, 60(9), 1214–1223. doi:10.1136/gut.2010.234708.
Lin, M. T., & Beal, M. F. (2006). Mitochondrial dysfunction and oxidative stress in neurodegenerative diseases. Nature, 443(7113), 787–795. doi:10.1038/nature05292.
Lindon, J. C., Nicholson, J. K., Holmes, E., Antti, H., Bollard, M. E., Keun, H., et al. (2003). Contemporary issues in toxicology the role of metabonomics in toxicology and its evaluation by the COMET project. Toxicology and Applied Pharmacology, 187(3), 137–146. doi:10.1016/S0041-008X(02)00079-0.
MacAllister, R. J., Parry, H., Kimoto, M., Ogawa, T., Russell, R. J., Hodson, H., et al. (1996). Regulation of nitric oxide synthesis by dimethylarginine dimethylaminohydrolase. British Journal of Pharmacology, 119(8), 1533–1540. doi:10.1111/j.1476-5381.1996.tb16069.x.
Mally, J., Szalai, G., & Stone, T. W. (1997). Changes in the concentration of amino acids in serum and cerebrospinal fluid of patients with Parkinson’s disease. Journal of the Neurological Sciences, 151(2), 159–162. doi:10.1016/S0022-510X(97)00119-6.
Mandal, R., Guo, A. C., Chaudhary, K. K., Liu, P., Yallou, F. S., Dong, E., et al. (2012). Multi-platform characterization of the human cerebrospinal fluid metabolome: a comprehensive and quantitative update. Genome Med, 4(4), 38. doi:10.1186/gm337.
Manuel, M., & Heckman, C. J. (2011). Stronger is not always better: could a bodybuilding dietary supplement lead to ALS? Experimental Neurology, 228(1), 5–8. doi:10.1016/j.expneurol.2010.12.007.
Menzies, F. M., Fleming, A., & Rubinsztein, D. C. (2015). Compromised autophagy and neurodegenerative diseases. Nature Reviews Neuroscience, 16(6), 345–357. doi:10.1038/nrn3961.
Milewski K, Hilgier W, Albrecht J, Zielinska M (2014). The dimethylarginine (ADMA)/nitric oxide pathway in the brain and periphery of rats with thioacetamide-induced acute liver failure: modulation by histidine. http://www.ncbi.nlm.nih.gov/pubmed/25523831.
Molina, J. A., Jimenez-Jimenez, F. J., Gomez, P., Vargas, C., Navarro, J. A., Orti-Pareja, M., et al. (1997). Decreased cerebrospinal fluid levels of neutral and basic amino acids in patients with Parkinson’s disease. Journal of the Neurological Sciences, 150(2), 123–127. doi:10.1016/S0022-510X(97)00069-5.
Nicholson, J. K., & Lindon, J. C. (2008). Systems biology: metabonomics. Nature, 455(7216), 1054–1056. doi:10.1038/4551054a.
Nicholson, J. K., Foxall, P. J. D., Spraul, M., Farrant, R. D., & Lindon, J. C. (1995). 750-Mhz H-1 and H-1-C-13 Nmr-spectroscopy of human blood-plasma. Analytical Chemistry, 67(5), 793–811. doi:10.1021/ac00101a004.
Öhman, A., & Forsgren, L. (2015). NMR metabonomics of cerebrospinal fluid distinguishes between Parkinson’s disease and controls. Neuroscience Letters, 594, 36–39. doi:10.1016/j.neulet.2015.03.051.
Otto, M., Bowser, R., Turner, M., Berry, J., Brettschneider, J., Connor, J., et al. (2012). Roadmap and standard operating procedures for biobanking and discovery of neurochemical markers in ALS. Amyotroph Lateral Scler, 13(1), 1–10. doi:10.3109/17482968.2011.627589.
Paillisse, C., Lacomblez, L., Dib, M., Bensimon, G., Garcia-Acosta, S., & Meininger, V. (2005). Prognostic factors for survival in amyotrophic lateral sclerosis patients treated with riluzole. Amyotroph Lateral Scler Other Motor Neuron Disord, 6(1), 37–44. doi:10.1080/14660820510027035.
Paoli, A., Bianco, A., Damiani, E., & Bosco, G. (2014). Ketogenic diet in neuromuscular and neurodegenerative diseases. Biomed Res Int, 2014, 474296. doi:10.1155/2014/474296.
Polymenidou, M., & Cleveland, D. W. (2011). The seeds of neurodegeneration: prion-like spreading in ALS. Cell, 147(3), 498–508. doi:10.1016/j.cell.2011.10.011.
Psychogios, N., Hau, D. D., Peng, J., Guo, A. C., Mandal, R., Bouatra, S., et al. (2011). The human serum metabolome. PLoS ONE, 6(2), e16957. doi:10.1371/journal.pone.0016957.
Quinones, M. P., & Kaddurah-Daouk, R. (2009). Metabolomics tools for identifying biomarkers for neuropsychiatric diseases. Neurobiology of Diseases, 35(2), 165–176. doi:10.1016/j.nbd.2009.02.019.
Rajput, A. H., Rozdilsky, B., & Rajput, A. (1991). Accuracy of clinical diagnosis in parkinsonism—a prospective study. Canadian Journal of Neurological Sciences, 18(3), 275–278.
Romero, P., Wagg, J., Green, M. L., Kaiser, D., Krummenacker, M., & Karp, P. D. (2005). Computational prediction of human metabolic pathways from the complete human genome. Genome Biology, 6(1), R2. doi:10.1186/gb-2004-6-1-r2.
Sharma, V., Ichikawa, M., & Freeze, H. H. (2014). Mannose metabolism: more than meets the eye. Biochem Biophys Res Commun, 453(2), 220–228. doi:10.1016/j.bbrc.2014.06.021.
Shaw, P. J. (2005). Molecular and cellular pathways of neurodegeneration in motor neurone disease. J Neurol Neurosurg Psychiatr, 76(8), 1046–1057. doi:10.1136/jnnp.2004.048652.
Sinclair, A. J., Viant, M. R., Ball, A. K., Burdon, M. A., Walker, E. A., Stewart, P. M., et al. (2010). NMR-based metabolomic analysis of cerebrospinal fluid and serum in neurological diseases—a diagnostic tool? NMR in Biomedicine, 23(2), 123–132. doi:10.1002/nbm.1428.
Smolinska, A., Blanchet, L., Buydens, L. M., & Wijmenga, S. S. (2012a). NMR and pattern recognition methods in metabolomics: from data acquisition to biomarker discovery: a review. Analytica Chimica Acta, 750, 82–97. doi:10.1016/j.aca.2012.05.049.
Smolinska, A., Posma, J. M., Blanchet, L., Ampt, K. A., Attali, A., Tuinstra, T., et al. (2012b). Simultaneous analysis of plasma and CSF by NMR and hierarchical models fusion. Analytical and Bioanalytical Chemistry, 403(4), 947–959. doi:10.1007/s00216-012-5871-4.
Stoop, M. P., Coulier, L., Rosenling, T., Shi, S., Smolinska, A. M., Buydens, L., et al. (2010). Quantitative proteomics and metabolomics analysis of normal human cerebrospinal fluid samples. Mol Cell Proteom, 9(9), 2063–2075. doi:10.1074/mcp.M900877-MCP200.
Tachikawa, M., Kasai, Y., Takahashi, M., Fujinawa, J., Kitaichi, K., Terasaki, T., et al. (2008). The blood-cerebrospinal fluid barrier is a major pathway of cerebral creatinine clearance: involvement of transporter-mediated process. Journal of Neurochemistry, 107(2), 432–442. doi:10.1111/j.1471-4159.2008.05641.x.
Tandan, R., & Bradley, W. G. (1985). Amyotrophic lateral sclerosis: part 1. Clinical features, pathology, and ethical issues in management. Ann Neurol, 18(3), 271–280. doi:10.1002/ana.410180302.
Trupp, M., Jonsson, P., Ohrfelt, A., Zetterberg, H., Obudulu, O., Malm, L., et al. (2014). Metabolite and peptide levels in plasma and CSF differentiating healthy controls from patients with newly diagnosed Parkinson’s disease. J Parkinsons Dis, 4(3), 549–560. doi:10.3233/JPD-140389.
Trygg, J., & Wold, S. (2002). Orthogonal projections to latent structures (O-PLS). Journal of Chemometrics, 16(3), 119–128. doi:10.1002/cem.695.
Tsacopoulos, M., & Magistretti, P. J. (1996). Metabolic coupling between glia and neurons. Journal of Neuroscience, 16(3), 877–885.
Vallance, P., & Leiper, J. (2004). Cardiovascular biology of the asymmetric dimethylarginine:dimethylarginine dimethylaminohydrolase pathway. Arteriosclerosis, Thrombosis, and Vascular Biology, 24(6), 1023–1030. doi:10.1161/01.ATV.0000128897.54893.26.
Van Sande, M., Caers, J., & Lowenthal, A. (1971). Cerebrospinal fluid amino acids in extrapyramidal disorders before and after L-DOPA treatment. Z Neurol, 199(1), 24–29.
Wade, A. M., & Tucker, H. N. (1998). Antioxidant characteristics of l-histidine 11The work described in this manuscript was partially sponsored and funded by cytos pharmaceuticals, LLC. Journal of Nutritional Biochemistry, 9(6), 308–315. doi:10.1016/s0955-2863(98)00022-9.
Want, E. J., Wilson, I. D., Gika, H., Theodoridis, G., Plumb, R. S., Shockcor, J., et al. (2010). Global metabolic profiling procedures for urine using UPLC-MS. Nature Protocols, 5(6), 1005–1018. doi:10.1038/nprot.2010.50.
Wishart, D. S., Lewis, M. J., Morrissey, J. A., Flegel, M. D., Jeroncic, K., Xiong, Y., et al. (2008). The human cerebrospinal fluid metabolome. J Chromatogr B Analyt Technol Biomed Life Sci, 871(2), 164–173. doi:10.1016/j.jchromb.2008.05.001.
Wishart, D. S., Jewison, T., Guo, A. C., Wilson, M., Knox, C., Liu, Y., et al. (2013). HMDB 3.0—the human metabolome database in 2013. Nucleic Acids Research, 41, 801–807. doi:10.1093/nar/gks1065. (Database issue).
Wuolikainen, A., Moritz, T., Marklund, S. L., Antti, H., & Andersen, P. M. (2011). Disease-related changes in the cerebrospinal fluid metabolome in amyotrophic lateral sclerosis detected by GC/TOFMS. PLoS ONE, 6(4), e17947. doi:10.1371/journal.pone.0017947.
Wuolikainen, A., Jonsson, P., Ahnlund, M., Antti, H., Marklund, S. L., Moritz, T., et al. (2016). Multi-platform mass spectrometry analysis of the CSF and plasma metabolomes of rigorously matched amyotrophic lateral sclerosis, Parkinson’s disease and control subjects. Molecular BioSystems, 12(4), 1287–1298. doi:10.1039/c5mb00711a.
Wyss, M., & Kaddurah-Daouk, R. (2000). Creatine and creatinine metabolism. Physiological Reviews, 80(3), 1107–1213.
Xia, J., Mandal, R., Sinelnikov, I. V., Broadhurst, D., & Wishart, D. S. (2012). MetaboAnalyst 2.0—a comprehensive server for metabolomic data analysis. Nucleic Acids Research, 40(Web Server issue), W127–W133. doi:10.1093/nar/gks374.
Zhang, A., Sun, H., Wang, P., Han, Y., & Wang, X. (2012). Recent and potential developments of biofluid analyses in metabolomics. J Proteomics, 75(4), 1079–1088. doi:10.1016/j.jprot.2011.10.027.
Zhao, Z., Lange, D. J., Voustianiouk, A., MacGrogan, D., Ho, L., Suh, J., et al. (2006). A ketogenic diet as a potential novel therapeutic intervention in amyotrophic lateral sclerosis. BMC Neurosci, 7, 29. doi:10.1186/1471-2202-7-29.
Acknowledgments
We are indebted to the patients who contributed to this study. We are grateful to the Erling-Persson Family Foundation, the Kempe Foundation and Umeå University for support. Dr. Junfang Wu is grateful for a post-doctoral fellowship from the Kempe foundation. Dr. Anna Wuolikainen, Dr. Miles Trupp, Dr. Pär Jonsson and Dr. Anders Öhman are supported by grants from the Erling-Persson Family Foundation. Dr. Peter Andersen has received grants from the Swedish Research Council, the Knut and Alice Wallenberg Foundation, The Bertil Hållsten Brain Research Foundation, The Brain Research Foundation, Swedish Brain Power Consortium, The Ulla-Carin Lindquist Foundation and Neuroförbundet. Dr. Lars Forsgren has received grants from The Swedish Medical Research Council, The Swedish Parkinson Foundation, The Swedish Parkinson’s disease Association, Västerbotten County Council (ALF), and the King Gustaf V’s and Queen Victoria’s Freemason Foundation. Dr. Anders Öhman has received grants from the Parkinson Foundation in Sweden and Insamlingsstiftelsen at Umeå University.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflicts of interest
The authors report no conflicts of interest related to this study.
Ethical approval
This study was carried out in accordance with the Declaration of Helsinki (WMA 1964), with ethical approval from the Medical Ethical Review Board for northern Sweden, Umeå, Sweden.
Informed consent
Written informed consent was obtained from all subjects.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Wu, J., Wuolikainen, A., Trupp, M. et al. NMR analysis of the CSF and plasma metabolome of rigorously matched amyotrophic lateral sclerosis, Parkinson’s disease and control subjects. Metabolomics 12, 101 (2016). https://doi.org/10.1007/s11306-016-1041-6
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s11306-016-1041-6