Abstract
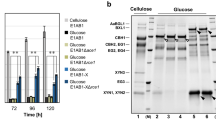
The role of the transcription factor creA-mediating carbon catabolite repression in Trichoderma orientalis EU7-22 was investigated for cellulase and hemicellulase production. The binary vector pUR5750G/creA::hph was constructed to knock out creA by homologous integration, generating the ΔcreA mutant Trichoderma orientalis CF1D. For strain CF1D, the filter paper activities (FPA), endoglucanase activities (CMC), cellobiohydrolase activity(CBH), β-glucosidase activity (BG), xylanase activity (XYN), and extracellular protein concentration were 1.45-, 1.15-, 1.71-, 2.51-, 2.72, and 1.95-fold higher in inducing medium and were 6.41-, 7.50-, 10.27-, 11.79-, 9.25-, and 3.77-fold higher in glucose repressing medium, respectively, than those in the parent strain after 4 days. SDS–PAGE demonstrated that the extracellular proteins were largely secreted in the mutant CF1D. Quantitative reverse-transcription polymerase chain reaction indicated that the expressions of cbh1, cbh2, eg1, eg2, bgl1, xyn1, and xyn2 were significantly increasing for the mutant CF1D not only in the inducing medium but also in the repressing medium. Those results indicated that creA was a valid target gene in strain engineering for improved enzyme production in T. orientalis.





Similar content being viewed by others
References
Kamm, B., & Kamm, M. (2007). Biorefineries—Multi product processes. Advances in Biochemical Engineering/Biotechnology, 105, 175–204.
Kumar, R., Singh, S., & Singh, O. V. (2008). Bioconversion of lignocellulosic biomass: Biochemical and molecular perspectives. Journal of Industrial Microbiology and Biotechnology, 35, 377–391.
Kovács, K., Szakacs, G., & Zacchi, G. (2009). Comparative enzymatic hydrolysis of pretreated spruce by supernatants, whole fermentation broths and washed mycelia of Trichoderma reesei and Trichoderma atroviride. Bioresource Technology, 100, 1350–1357.
Sipos, B., Benko, Z., Dienes, D., Réczey, K., Viikari, L., & Siika-aho, M. (2010). Characterisation of specific activities and hydrolytic properties of cell-wall degrading enzymes produced by Trichoderma reesei Rut C30 on different carbon sources. Applied Biochemistry and Biotechnology, 161, 347–364.
Portnoy, T., Margeot, A., Linke, R., Atanasova, L., Fekete, E., Sándor, E., et al. (2011). The CRE1 carbon catabolite repressor of the fungus Trichoderma reesei: A master regulator of carbon assimilation. BMC Genomics, 12(269), 1–12.
Takashima, S., Iikura, H., Nakamura, A., Masaki, H., & Uozumi, T. (1996). Analysis of Cre1 binding sites in the Trichoderma reesei cbh1 upstream region. FEMS Microbiology Letters, 145, 361–366.
Cubero, B., & Scazzocchio, C. (1994). Two different, adjacent and divergent zinc finger binding sites are necessary for CREA-mediated carbon catabolite repression in the proline gene cluster of Aspergillus nidulans. EMBO Journal, 13, 407–415.
Drysdale, M. R., Kolze, S. E., & Kelly, J. M. (1993). The Aspergillus niger carbon catabolite repressor encoding gene, creA. Gene, 130, 241–245.
Sun, J., & Glass, N. L. (2011). Identification of the CRE-1 cellulolytic regulon in Neurospora crassa. PLoS ONE, 6, e25654.
Long, C. N., Cheng, Y. J., Gan, L. H., Liu, J., & Long, M. N. (2013). Identification of a genomic region containing a novel promoter resistant to glucose repression and over-expression of β-glucosidase gene in Hypocrea orientalis EU7-22. International Journal of Molecular Sciences, 14(4), 8479–8490.
Penttilä, M., Nevalainen, H., Rättö, M., Salminen, E., & Knowles, J. (1987). A versatile transformation system for the cellulolytic filamentous fungus Trichoderma reesei. Gene, 61, 155–164.
Ghose, T. K. (1987). Measurement of cellulase activities. Pure and Applied Chemistry, 59, 257–268.
Saha, B. C. (2014). Production, purification and properties of endoglucanase from a newly isolated strain of Mucor circinelloides. Process Biochemistry, 39, 1871–1876.
Bailey, M. J., Biely, P., & Poutanen, K. (1992). Interlaboratory testing of methods for assay of xylanase activity. Journal of Biotechnology, 23, 257–270.
Nakari-Setälä, T., Paloheimo, M., Kallio, J., Vehmaanperä, J., Penttilä, M., & Saloheimo, M. (2009). Genetic modification of carbon catabolite repression in Trichoderma reesei for improved protein production. Applied and Environment Microbiology, 75, 4853–4860.
Mach-Aigner, A. R., Pucher, M. E., Steiger, M. G., Bauer, G. E., Preis, S. J., & Mach, R. L. (2008). Transcriptional regulation of xyr1, encoding the main regulator of the xylanolytic and cellulolytic enzyme system in Hypocrea jecorina. Applied and Environment Microbiology, 74, 6554–6562.
Ries, L., Belshaw, N. J., Ilmén, M., Penttilä, M. E., Alapuranen, M., & Archer, D. B. (2014). The role of CRE1 in nucleosome positioning within the cbh1 promoter and coding regions of Trichoderma reesei. Applied Microbiology and Biotechnology, 98, 749–762.
Acknowledgements
This work was supported by the National Natural Science Foundation of China (Grant No. 31170067, 21303142), Jiangxi Province Science Foundation for Youths (Grant No. 20161BAB214177).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no conflict of interest.
Rights and permissions
About this article
Cite this article
Long, C., Cheng, Y., Cui, J. et al. Enhancing Cellulase and Hemicellulase Production in Trichoderma orientalis EU7-22 via Knockout of the creA . Mol Biotechnol 60, 55–61 (2018). https://doi.org/10.1007/s12033-017-0046-3
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12033-017-0046-3