Abstract
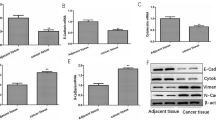
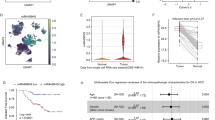
Lung cancer (LC) is a pulmonary malignant tumor with extremely low 5-year survival rate. N6-methyladenosine (m6A) is confirmed to regulate diverse pathophysiological processes including cancers. Methyltransferase-like 14 (METTL14) is an important RNA methyltransferase in m6A modification. However, researches on the regulatory mechanism of METTL14 on LC progression are relatively rare. Tumor xenograft experiment was conducted to investigate the effect of METTL14 on LC in vivo. The relative expression of METTL14, miR-30c-1-3p, and myristoylated alanine-rich C kinase substrate-like protein-1 (MARCKSL1) in LC tissues and/or cell lines was determined using qRT-PCR. Western blot assay was used to measure the protein levels of METTL14 and MARCKSL1 in tumor xenograft model and/or LC cell lines. MTT, wound healing, and transwell assays were performed to detect LC cell viability and metastasis. RNA immunoprecipitation assay and qRT-PCR were used to verify the effects of METTL14 on pri-miR-30c-1-3p. The relationship between miR-30c-1-3p and MARCKSL1 was confirmed by the dual-luciferase reporter assay. METTL14 was remarkably downregulated in LC tissues and cell lines. METTL14 mediated the maturation of miR-30c-1-3p. The overexpressed METTL14 and overexpressed miR-30c-1-3p suppressed the cell viability and metastasis in LC. Meanwhile, the increased METTL14 also repressed the growth of tumor xenograft in vivo. In addition, MARCKSL1 was confirmed to be the target gene of miR-30c-1-3p. High expression of MARCKSL1 and low expression of miR-30c-1-3p reversed the suppressive effects of METTL14 overexpression on cell viability and metastasis. METTL14 promoted the maturation of miR-30c-1-3p and mediated MARCKSL1 expression to inhibit the progression of LC. This study may provide a new insight for the LC clinical therapy.







Similar content being viewed by others
References
Travis, W. D. (2020). Lung cancer pathology: Current concepts. Clinics in Chest Medicine, 41(1), 67–85. https://doi.org/10.1016/j.ccm.2019.11.001
Siegel, R. L., Miller, K. D., & Jemal, A. (2019). Cancer statistics, 2019. CA: A Cancer Journal for Clinicians, 69(1), 7–34. https://doi.org/10.3322/caac.21551
Ridge, C. A., McErlean, A. M., & Ginsberg, M. S. (2013). Epidemiology of lung cancer. Seminars in Interventional Radiology, 30(2), 93–98. https://doi.org/10.1055/s-0033-1342949
Hussain, S. (2016). Nanomedicine for treatment of lung cancer. Advances in Experimental Medicine and Biology, 890, 137–147. https://doi.org/10.1007/978-3-319-24932-2_8
Quaratino, S., Forssmann, U., & Marschner, J. P. (2017). New approaches in immunotherapy for the treatment of lung cancer. Current Topics in Microbiology and Immunology, 405, 1–31. https://doi.org/10.1007/82_2014_428
Detterbeck, F. C., Boffa, D. J., Kim, A. W., & Tanoue, L. T. (2017). The Eighth Edition Lung Cancer Stage Classification. Chest, 151(1), 193–203. https://doi.org/10.1016/j.chest.2016.10.010
Cantara, W. A., Crain, P. F., Rozenski, J., McCloskey, J. A., Harris, K. A., Zhang, X., Vendeix, F. A., Fabris, D., & Agris, P. F. (2011). The RNA Modification Database, RNAMDB: 2011 update. Nucleic Acids Research, 39(Database Issue), D195-201. https://doi.org/10.1093/nar/gkq1028
Machnicka, M. A., Milanowska, K., Osman Oglou, O., Purta, E., Kurkowska, M., Olchowik, A., Januszewski, W., Kalinowski, S., Dunin-Horkawicz, S., Rother, K. M., Helm, M., Bujnicki, J. M., & Grosjean, H. (2013). MODOMICS: A database of RNA modification pathways—2013 Update. Nucleic Acids Research, 41(Database Issue), D262-267. https://doi.org/10.1093/nar/gks1007
Frye, M., Harada, B. T., Behm, M., & He, C. (2018). RNA modifications modulate gene expression during development. Science, 361(6409), 1346–1349. https://doi.org/10.1126/science.aau1646
Lee, M., Kim, B., & Kim, V. N. (2014). Emerging roles of RNA modification: m(6)A and U-tail. Cell, 158(5), 980–987. https://doi.org/10.1016/j.cell.2014.08.005
Gu, C., Wang, Z., Zhou, N., Li, G., Kou, Y., Luo, Y., Wang, Y., Yang, J., & Tian, F. (2019). Mettl14 inhibits bladder TIC self-renewal and bladder tumorigenesis through N(6)-methyladenosine of Notch1. Molecular Cancer, 18(1), 168. https://doi.org/10.1186/s12943-019-1084-1
Jin, D., Guo, J., Wu, Y., Du, J., Yang, L., Wang, X., Di, W., Hu, B., An, J., Kong, L., Pan, L., & Su, G. (2019). m(6)A mRNA methylation initiated by METTL3 directly promotes YAP translation and increases YAP activity by regulating the MALAT1-miR-1914-3p-YAP axis to induce NSCLC drug resistance and metastasis. Journal of Hematology and Oncology, 12(1), 135. https://doi.org/10.1186/s13045-019-0830-6
Qian, J. Y., Gao, J., Sun, X., Cao, M. D., Shi, L., Xia, T. S., Zhou, W. B., Wang, S., Ding, Q., & Wei, J. F. (2019). KIAA1429 acts as an oncogenic factor in breast cancer by regulating CDK1 in an N6-methyladenosine-independent manner. Oncogene, 38(33), 6123–6141. https://doi.org/10.1038/s41388-019-0861-z
Selberg, S., Blokhina, D., Aatonen, M., Koivisto, P., Siltanen, A., Mervaala, E., Kankuri, E., & Karelson, M. (2019). Discovery of small molecules that activate RNA methylation through cooperative binding to the METTL3-14-WTAP complex active site. Cell Reports, 26(13), 3762-3771.e3765. https://doi.org/10.1016/j.celrep.2019.02.100
Zhang, S., Zhao, B. S., Zhou, A., Lin, K., Zheng, S., Lu, Z., Chen, Y., Sulman, E. P., Xie, K., Bogler, O., Majumder, S., He, C., & Huang, S. (2017). m(6)A demethylase ALKBH5 maintains tumorigenicity of glioblastoma stem-like cells by sustaining FOXM1 expression and cell proliferation program. Cancer Cell, 31(4), 591-606.e596. https://doi.org/10.1016/j.ccell.2017.02.013
Li, Z., Weng, H., Su, R., Weng, X., Zuo, Z., Li, C., Huang, H., Nachtergaele, S., Dong, L., Hu, C., Qin, X., Tang, L., Wang, Y., Hong, G. M., Wang, X., Chen, P., Gurbuxani, S., Arnovitz, S., Li, Y., Li, S., et al. (2017). FTO plays an oncogenic role in acute myeloid leukemia as a N(6)-methyladenosine RNA demethylase. Cancer Cell, 31(1), 127–141. https://doi.org/10.1016/j.ccell.2016.11.017
Tiwari, N., Tiwari, V. K., Waldmeier, L., Balwierz, P. J., Arnold, P., Pachkov, M., Meyer-Schaller, N., Schubeler, D., van Nimwegen, E., & Christofori, G. (2013). Sox4 is a master regulator of epithelial–mesenchymal transition by controlling Ezh2 expression and epigenetic reprogramming. Cancer Cell, 23(6), 768–783. https://doi.org/10.1016/j.ccr.2013.04.020
Huang, H., Weng, H., Sun, W., Qin, X., Shi, H., Wu, H., Zhao, B. S., Mesquita, A., Liu, C., Yuan, C. L., Hu, Y. C., Huttelmaier, S., Skibbe, J. R., Su, R., Deng, X., Dong, L., Sun, M., Li, C., Nachtergaele, S., Wang, Y., et al. (2018). Recognition of RNA N(6)-methyladenosine by IGF2BP proteins enhances mRNA stability and translation. Nature Cell Biology, 20(3), 285–295. https://doi.org/10.1038/s41556-018-0045-z
Zhao, B. S., Wang, X., Beadell, A. V., Lu, Z., Shi, H., Kuuspalu, A., Ho, R. K., & He, C. (2017). m(6)A-dependent maternal mRNA clearance facilitates zebrafish maternal-to-zygotic transition. Nature, 542(7642), 475–478. https://doi.org/10.1038/nature21355
Lin, Z., Hsu, P. J., Xing, X., Fang, J., Lu, Z., Zou, Q., Zhang, K. J., Zhang, X., Zhou, Y., Zhang, T., Zhang, Y., Song, W., Jia, G., Yang, X., He, C., & Tong, M. H. (2017). Mettl3-/Mettl14-mediated mRNA N(6)-methyladenosine modulates murine spermatogenesis. Cell Research, 27(10), 1216–1230. https://doi.org/10.1038/cr.2017.117
Wen, J., Lv, R., Ma, H., Shen, H., He, C., Wang, J., Jiao, F., Liu, H., Yang, P., Tan, L., Lan, F., Shi, Y. G., Shi, Y., & Diao, J. (2018). Zc3h13 regulates nuclear RNA m(6)A methylation and mouse embryonic stem cell self-renewal. Molecular Cell, 69(6), 1028-1038.e1026. https://doi.org/10.1016/j.molcel.2018.02.015
Fustin, J. M., Doi, M., Yamaguchi, Y., Hida, H., Nishimura, S., Yoshida, M., Isagawa, T., Morioka, M. S., Kakeya, H., Manabe, I., & Okamura, H. (2013). RNA-methylation-dependent RNA processing controls the speed of the circadian clock. Cell, 155(4), 793–806. https://doi.org/10.1016/j.cell.2013.10.026
Xiang, Y., Laurent, B., Hsu, C. H., Nachtergaele, S., Lu, Z., Sheng, W., Xu, C., Chen, H., Ouyang, J., Wang, S., Ling, D., Hsu, P. H., Zou, L., Jambhekar, A., He, C., & Shi, Y. (2017). RNA m(6)A methylation regulates the ultraviolet-induced DNA damage response. Nature, 543(7646), 573–576. https://doi.org/10.1038/nature21671
Zhang, C., Chen, Y., Sun, B., Wang, L., Yang, Y., Ma, D., Lv, J., Heng, J., Ding, Y., Xue, Y., Lu, X., Xiao, W., Yang, Y. G., & Liu, F. (2017). m(6)A modulates haematopoietic stem and progenitor cell specification. Nature, 549(7671), 273–276. https://doi.org/10.1038/nature23883
Li, H. B., Tong, J., Zhu, S., Batista, P. J., Duffy, E. E., Zhao, J., Bailis, W., Cao, G., Kroehling, L., Chen, Y., Wang, G., Broughton, J. P., Chen, Y. G., Kluger, Y., Simon, M. D., Chang, H. Y., Yin, Z., & Flavell, R. A. (2017). m(6)A mRNA methylation controls T cell homeostasis by targeting the IL-7/STAT5/SOCS pathways. Nature, 548(7667), 338–342. https://doi.org/10.1038/nature23450
Du, C., Lv, C., Feng, Y., & Yu, S. (2020). Activation of the KDM5A/miRNA-495/YTHDF2/m6A-MOB3B axis facilitates prostate cancer progression. Journal of Experimental and Clinical Cancer Research, 39(1), 223. https://doi.org/10.1186/s13046-020-01735-3
Strick, A., von Hagen, F., Gundert, L., Klumper, N., Tolkach, Y., Schmidt, D., Kristiansen, G., Toma, M., Ritter, M., & Ellinger, J. (2020). The N(6)-methyladenosine (m(6) A) erasers alkylation repair homologue 5 (ALKBH5) and fat mass and obesity-associated protein (FTO) are prognostic biomarkers in patients with clear cell renal carcinoma. BJU International, 125(4), 617–624. https://doi.org/10.1111/bju.15019
Lin, S., Choe, J., Du, P., Triboulet, R., & Gregory, R. I. (2016). The m(6)A methyltransferase METTL3 promotes translation in human cancer cells. Molecular Cell, 62(3), 335–345. https://doi.org/10.1016/j.molcel.2016.03.021
Weng, L., Qiu, K., Gao, W., Shi, C., & Shu, F. (2020). LncRNA PCGEM1 accelerates non-small cell lung cancer progression via sponging miR-433-3p to upregulate WTAP. BMC Pulmonary Medicine, 20(1), 213. https://doi.org/10.1186/s12890-020-01240-5
Wang, H., Zhao, X., & Lu, Z. (2021). m(6)A RNA methylation regulators act as potential prognostic biomarkers in lung adenocarcinoma. Frontiers in Genetics, 12, 622233. https://doi.org/10.3389/fgene.2021.622233
Zhu, Z., Qian, Q., Zhao, X., Ma, L., & Chen, P. (2020). N(6)-methyladenosine ALKBH5 promotes non-small cell lung cancer progress by regulating TIMP3 stability. Gene, 731, 144348. https://doi.org/10.1016/j.gene.2020.144348
Li, J., Han, Y., Zhang, H., Qian, Z., Jia, W., Gao, Y., Zheng, H., & Li, B. (2019). The m6A demethylase FTO promotes the growth of lung cancer cells by regulating the m6A level of USP7 mRNA. Biochemical and Biophysical Research Communications, 512(3), 479–485. https://doi.org/10.1016/j.bbrc.2019.03.093
Zhang, Y., Liu, X., Liu, L., Li, J., Hu, Q., & Sun, R. (2020). Expression and prognostic significance of m6A-related genes in lung adenocarcinoma. Medical Science Monitor, 26, e919644. https://doi.org/10.12659/MSM.919644
Chen, X., Xu, M., Xu, X., Zeng, K., Liu, X., Pan, B., Li, C., Sun, L., Qin, J., Xu, T., He, B., Pan, Y., Sun, H., & Wang, S. (2020). METTL14-mediated N6-methyladenosine modification of SOX4 mRNA inhibits tumor metastasis in colorectal cancer. Molecular Cancer, 19(1), 106. https://doi.org/10.1186/s12943-020-01220-7
Li, F., Wang, H., Huang, H., Zhang, L., Wang, D., & Wan, Y. (2020). m6A RNA methylation regulators participate in the malignant progression and have clinical prognostic value in lung adenocarcinoma. Frontiers in Genetics, 11, 994. https://doi.org/10.3389/fgene.2020.00994
Guo, H., Ingolia, N. T., Weissman, J. S., & Bartel, D. P. (2010). Mammalian microRNAs predominantly act to decrease target mRNA levels. Nature, 466(7308), 835–840. https://doi.org/10.1038/nature09267
Cai, X., Wang, X., Cao, C., Gao, Y., Zhang, S., Yang, Z., Liu, Y., Zhang, X., Zhang, W., & Ye, L. (2018). HBXIP-elevated methyltransferase METTL3 promotes the progression of breast cancer via inhibiting tumor suppressor let-7g. Cancer Letters, 415, 11–19. https://doi.org/10.1016/j.canlet.2017.11.018
Ma, J. Z., Yang, F., Zhou, C. C., Liu, F., Yuan, J. H., Wang, F., Wang, T. T., Xu, Q. G., Zhou, W. P., & Sun, S. H. (2017). METTL14 suppresses the metastatic potential of hepatocellular carcinoma by modulating N(6)-methyladenosine-dependent primary microRNA processing. Hepatology, 65(2), 529–543. https://doi.org/10.1002/hep.28885
Yi, D., Wang, R., Shi, X., Xu, L., Yilihamu, Y., & Sang, J. (2020). METTL14 promotes the migration and invasion of breast cancer cells by modulating N6methyladenosine and hsamiR146a5p expression. Oncology Reports, 43(5), 1375–1386. https://doi.org/10.3892/or.2020.7515
Bi, X., Lv, X., Liu, D., Guo, H., Yao, G., Wang, L., Liang, X., & Yang, Y. (2020). METTL3-mediated maturation of miR-126-5p promotes ovarian cancer progression via PTEN-mediated PI3K/Akt/mTOR pathway. Cancer Gene Therapy. https://doi.org/10.1038/s41417-020-00222-3
Chen, X., Xu, M., Xu, X., Zeng, K., Liu, X., Sun, L., Pan, B., He, B., Pan, Y., Sun, H., Xia, X., & Wang, S. (2020). METTL14 suppresses CRC progression via regulating N6-methyladenosine-dependent primary miR-375 processing. Molecular Therapy, 28(2), 599–612. https://doi.org/10.1016/j.ymthe.2019.11.016
Zhang, B. Y., Han, L., Tang, Y. F., Zhang, G. X., Fan, X. L., Zhang, J. J., Xue, Q., & Xu, Z. Y. (2020). METTL14 regulates M6A methylation-modified primary miR-19a to promote cardiovascular endothelial cell proliferation and invasion. European Review for Medical and Pharmacological Sciences, 24(12), 7015–7023. https://doi.org/10.26355/eurrev_202006_21694
Kordass, T., Weber, C. E. M., Eisel, D., Pane, A. A., Osen, W., & Eichmuller, S. B. (2018). miR-193b and miR-30c-1(*) inhibit, whereas miR-576-5p enhances melanoma cell invasion in vitro. Oncotarget, 9(65), 32507–32522. https://doi.org/10.18632/oncotarget.25986
Zhang, X., Zhang, G., Huang, H., Li, H., Lin, S., & Wang, Y. (2020). Differentially expressed microRNAs in radioresistant and radiosensitive atypical meningioma: A clinical study in Chinese patients. Frontiers in Oncology, 10, 501. https://doi.org/10.3389/fonc.2020.00501
Chen, W., Yao, G., & Zhou, K. (2019). miR-103a-2-5p/miR-30c-1-3p inhibits the progression of prostate cancer resistance to androgen ablation therapy via targeting androgen receptor variant 7. Journal of Cellular Biochemistry, 120(8), 14055–14064. https://doi.org/10.1002/jcb.28680
Wu, M., Liang, G., Duan, H., Yang, X., Qin, G., & Sang, N. (2019). Synergistic effects of sulfur dioxide and polycyclic aromatic hydrocarbons on pulmonary pro-fibrosis via mir-30c-1-3p/ transforming growth factor beta type II receptor axis. Chemosphere, 219, 268–276. https://doi.org/10.1016/j.chemosphere.2018.12.016
Li, T., Li, D., Sha, J., Sun, P., & Huang, Y. (2009). MicroRNA-21 directly targets MARCKS and promotes apoptosis resistance and invasion in prostate cancer cells. Biochemical and Biophysical Research Communications, 383(3), 280–285. https://doi.org/10.1016/j.bbrc.2009.03.077
Salem, O., Erdem, N., Jung, J., Munstermann, E., Worner, A., Wilhelm, H., Wiemann, S., & Korner, C. (2016). The highly expressed 5′isomiR of hsa-miR-140-3p contributes to the tumor-suppressive effects of miR-140 by reducing breast cancer proliferation and migration. BMC Genomics, 17, 566. https://doi.org/10.1186/s12864-016-2869-x
Liu, H., Su, P., Zhi, L., & Zhao, K. (2017). miR34c3p acts as a tumor suppressor gene in osteosarcoma by targeting MARCKS. Molecular Medicine Reports, 15(3), 1204–1210. https://doi.org/10.3892/mmr.2017.6108
Liang, W., Gao, R., Yang, M., Wang, X., Cheng, K., Shi, X., He, C., Li, Y., Wu, Y., Shi, L., Chen, J., & Yu, X. (2020). MARCKSL1 promotes the proliferation, migration and invasion of lung adenocarcinoma cells. Oncology Letters, 19(3), 2272–2280. https://doi.org/10.3892/ol.2020.11313
Liu, X., Xiao, M., Zhang, L., Li, L., Zhu, G., Shen, E., Lv, M., Lu, X., & Sun, Z. (2020). The m6A methyltransferase METTL14 inhibits the proliferation, migration, and invasion of gastric cancer by regulating the PI3K/AKT/mTOR signaling pathway. Journal of Clinical Laboratory Analysis. https://doi.org/10.1002/jcla.23655
Jonsdottir, K., Zhang, H., Jhagroe, D., Skaland, I., Slewa, A., Bjorkblom, B., Coffey, E. T., Gudlaugsson, E., Smaaland, R., Janssen, E. A., & Baak, J. P. (2012). The prognostic value of MARCKS-like 1 in lymph node-negative breast cancer. Breast Cancer Research Treatment, 135(2), 381–390. https://doi.org/10.1007/s10549-012-2155-9
Funding
Not applicable.
Author information
Authors and Affiliations
Contributions
Conceptualization: FL and YC; Methodology: W L; Formal analysis and investigation: FL, XH, and LW; Writing-original draft preparation: FL; Writing-review and editing: WL, YC, and JZ; Funding acquisition: FL; Resources: XH and LW; Supervision: YC.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflicts of interest.
Ethical Approval
This study was performed in line with the principles of the Declaration of Helsinki. The present study was approved by Ethics Committee of Peoples Hospital of Rizhao, and the relevant informed consents were written by each patient.
Informed Consent
Informed consent was obtained from all individual participants included in the study.
Consent to Publish
Not applicable.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Li, F., Zhao, J., Wang, L. et al. METTL14-Mediated miR-30c-1-3p Maturation Represses the Progression of Lung Cancer via Regulation of MARCKSL1 Expression. Mol Biotechnol 64, 199–212 (2022). https://doi.org/10.1007/s12033-021-00406-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12033-021-00406-8