Abstract
In silico molecular modeling studies were carried out on some newly synthesized flavanoid analogues. Search for potential targets for these compounds was performed using pharmacophore-mapping algorithm employing inverse screening of some representative compounds to a large set of pharmacophore models constructed from human target proteins. Further, molecular docking studies were carried out to assess binding affinity of these compounds to proteins mediating tumor growth. In vitro anticancer studies were carried out on colon cancer cell lines (HCT116) to assess validity of this approach for target identification of the new compounds. Further important structural features of compounds for anticolon cancer activity were assessed using Monte Carlo-based SMILES and hydrogen graph-Based QSAR studies. In conclusion this study have depicted successful and stepwise application of pharmacophore mapping, molecular docking, and QSAR studies in target identification and lead optimization of flavonoids.




Similar content being viewed by others
References
Hichri I, Barrieu F, Bogs J, Kappel C, Delrot S, Lauvergeat V (2011) Recent advances in the transcriptional regulation of the flavonoid biosynthetic pathway. J Exp Bot 62:2465–2483
Halbwirth H (2010) The creation and physiological relevance of divergent hydroxylation patterns in the flavonoid pathway. Int J Mol Sci 11:595–621
Gupta S, Shivahare R, Korthikunta V, Singh R, Gupta S, Tadigoppula N (2014) Synthesis and biological evaluation of chalcones as potential antileishmanial agents. Eur J Med Chem 81:359–366
Rahman MA (2011) Chalcone: a valuable insight into the recent advances and potential pharmacological activities. Chem Sci J 29:1–16
Sivakumar P, Prabhakar P, Doble M (2011) Synthesis, antioxidant evaluation, and quantitative structure–activity relationship studies of chalcones. Med Chem Res 20:482–492
Lahsasni SA, Al Korbi FH, Aljaber NA-A (2014) Synthesis, characterization and evaluation of antioxidant activities of some novel chalcones analogues. Chem Cent J 8:32
Scotti L, Mendonca JB, Moreira DRM, Silva MS, Pitta IR, Scotti MT (2012) SAR, QSAR and docking of anticancer flavonoids and variants: a review. Curr Top Med Chem 12(24):2785–2809
Ibrahim A, Sobeh M, Ismail A, Alaa A, Sheashaa H, Sobh M, Badria F (2014) Free-B-Ring flavonoids as potential lead compounds for colon cancer therapy. Mol Clin Oncol 2:581–585
Morton A, Cao L (1038) Bourquin LD (2013) A flavonoid-rich apple extract inhibits growth of HCT116 human colon cancer cells. FASEB J 27:1079
Carmona SJ, Lopez S, Abia R, Arcos RR, Jimenez A, Guillen R, Muriana FJG (2014) Combination of quercetin and kaempferol enhances in vitro cytotoxicity on human colon cancer (HCT-116) cells. Rec Nat Prod 8(3):262–271
Liu X, Ouyang S, Yu B, Liu Y, Huang K, Gong J, Zheng S, Li Z, Li H, Jiang H (2010) PharmMapper server: a web server for potential drug target identification using pharmacophore mapping approach. Nucleic Acids Res 38:W609–W614
Shewchuk L, Hassell A, Wisely B, Rocque W, Holmes W, Veal J, Kuyper LF (2000) Binding mode of the 4-anilinoquinazoline class of protein kinase inhibitor: X-ray crystallographic studies of 4-anilinoquinazolines bound to cyclin-dependent kinase 2 and p38 kinase. J Med Chem 43:133–138
Thomsen R, Christensen MH (2006) MolDock: a new technique for high-accuracy molecular docking. J Med Chem 49:3315–3321
Simon L, Srinivasan K, Kumar N, Reddy N, Biswas S, Rao CM, Moorkoth S (2016) Selected novel 5′-amino-2′-hydroxy-1, 3-diaryl-2-propen-1-ones arrest cell cycle of HCT 116 in G0/G1 phase. EXCLI J Exp Clin Sci 15:21–32
Simon L, Srinivasan K, Rao CM, Kumar N, Reddy N, Biswas S, Moorkoth S (2015) Synthesis and evaluation of anti-cancer activity of some 6-Aminoflavones. Int J Pharm Chem 5:240–246
Veselinović J, Veselinović A, Toropov A, Toropova A, Damnjanović I, Nikolić G (2014) Monte Carlo method based QSAR modeling of coumarin derivates as potent HIV-1 integrase inhibitors and molecular docking studies of selected 4-phenyl hydroxycoumarins. Acta Fac Medicae Naissensis 31:95–103
Veselinović JB, Toropov AA, Toropova AP, Nikolić GM, Veselinović AM (2015) Monte carlo method-based QSAR modeling of penicillins binding to human serum proteins. Arch Pharm (Weinheim) 348(1):62–67
Toropova AP, Toropov AA, Veselinović JB, Miljković FN, Veselinović AM (2014) QSAR models for HEPT derivates as NNRTI inhibitors based on Monte Carlo method. Eur J Med Chem 77:298–305
Weininger D (1988) SMILES, a chemical language and information system. 1. Introduction to methodology and encoding rules. J Chem Inf Comput Sci 28:31–36
Basak S, Gute B (1997) Characterization of molecular structures using topological indices. SAR QSAR Environ Res 7:1–21
Roy PP, Roy K (2009) QSAR studies of CYP2D6 inhibitor aryloxypropanolamines using 2D and 3D descriptors. Chem Biol Drug Des 73:442–455
Ojha PK, Mitra I, Das RN, Roy K (2011) Further exploring r m 2 metrics for validation of QSPR models. Chemom Intell Lab Syst 107:194–205
Ojha PK, Roy K (2011) Comparative QSARs for antimalarial endochins: importance of descriptor-thinning and noise reduction prior to feature selection. Chemom Intell Lab Syst 109:146–161
Ambasta RK, Jha SK, Kumar D, Sharma R, Jha NK, Kumar P (2015) Comparative study of anti-angiogenic activities of luteolin, lectin and lupeol biomolecules. J Transl Med 13:307
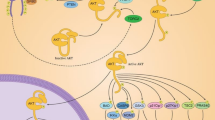
Orellana AM, Vasconcelos AR, Leite JA, de Sá Lima L, Andreotti DZ, Munhoz CD, Kawamoto EM, Scavone C (2015) Age-related neuroinflammation and changes in AKT-GSK-3β and WNT/β-CATENIN signaling in rat hippocampus. Aging 7(12):1094–1111
Golpich M, Amini E, Hemmati F, Ibrahim NM, Rahmani B, Mohamed Z, Raymond AA, Dargahi L, Ghasemi R, Ahmadiani A (2015) Glycogen synthase kinase-3 beta (GSK-3β) signaling: implications for Parkinson's disease. Pharmacol Res 97:16–26
Lei P, Ayton S, Bush AI, Adlard PA (2011) GSK-3 in neurodegenerative diseases. Int J Alzheimer’s Dis 2011:189246
Li B, Thrasher JB, Terranova P (2015) Glycogen synthase kinase-3: a potential preventive target for prostate cancer management. Urol Oncol Semin Orig Investig 33:456–463
McCubrey JA, Abrams SL, Fitzgerald TL, Cocco L, Martelli AM, Montalto G, Cervello M, Scalisi A, Candido S, Libra M (2015) Roles of signaling pathways in drug resistance, cancer initiating cells and cancer progression and metastasis. Adv Biol Regul 57:75–101
Greuber EK, Smith-Pearson P, Wang J, Pendergast AM (2013) Role of ABL family kinases in cancer: from leukaemia to solid tumours. Nat Rev Cancer 13:559–571
Sivaraman VS, Wang H, Nuovo GJ, Malbon CC (1997) Hyperexpression of mitogen-activated protein kinase in human breast cancer. J Clin Investig 99(7):1478–1483
Foloppe N, Fisher LM, Howes R, Potter A, Robertson AG, Surgenor AE (2006) Identification of chemically diverse Chk1 inhibitors by receptor-based virtual screening. Bioorganic Med Chem 14:4792–4802
Chohan TA, Qian H, Pan Y, Chen J-Z (2015) Cyclin-dependent kinase-2 as a target for cancer therapy: progress in the development of CDK2 inhibitors as anti-cancer agents. Curr Med Chem 22:237–263
Porter AG, Jänicke RU (1999) Emerging roles of caspase-3 in apoptosis. Cell Death Differ 6:99–104
Tarling CA, Woods K, Zhang R, Brastianos HC, Brayer GD, Andersen RJ, Withers SG (2008) The search for novel human pancreatic α-amylase inhibitors: high-throughput screening of terrestrial and marine natural product extracts. ChemBioChem 9:433–438
McCormick F (1995) Ras-related proteins in signal transduction and growth control. Mol Reprod Dev 42:500–506
Lamson RE, Winters MJ, Pryciak PM (2002) Cdc42 regulation of kinase activity and signaling by the yeast p21-activated kinase Ste20. Mol Cell Biol 22:2939–2951
Mikhail S, Albanese C, Pishvaian MJ (2015) Cyclin-dependent kinase inhibitors and the treatment of gastrointestinal cancers. Am J Pathol 185:1185–1197
Laskowski RA, Swindells MB (2011) LigPlot + : multiple ligand–protein interaction diagrams for drug discovery. J Chem Inf Model 51:2778–2786
Roy K, Chakraborty P, Mitra I, Ojha PK, Kar S, Das RN (2013) Some case studies on application of “rm2″ metrics for judging quality of quantitative structure–activity relationship predictions: emphasis on scaling of response data. J Comput Chem 34:1071–1082
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Simon, L., Imane, A., Srinivasan, K.K. et al. In Silico Drug-Designing Studies on Flavanoids as Anticolon Cancer Agents: Pharmacophore Mapping, Molecular Docking, and Monte Carlo Method-Based QSAR Modeling. Interdiscip Sci Comput Life Sci 9, 445–458 (2017). https://doi.org/10.1007/s12539-016-0169-4
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12539-016-0169-4