Abstract
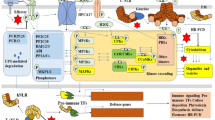
Innate immune signaling of plants is initiated by pattern recognition receptors (PRRs) at the plasma membrane. Upon pathogen attack, PRRs recognize pathogen-associated molecular patterns (PAMPs) via ectodomain and lead to signaling cascade via cytoplasmic kinase domain. PAMP-triggered immunity (PTI) activates basal defense responses sufficient to confer broad-spectrum disease resistance by inhibiting pathogen entry and growth. On the other hand, one of the major virulence factors in plant-pathogenic bacteria is type III secretion system, which can deliver effector proteins into the host cell and modulate host cellular processes. Most type III effectors are implicated in PTI suppression, and PRRs have been identified as targets of multiple type III effectors. Mutants defective in T3SS lack pathogenicity in many bacterial species, revealing that T3SS-mediated PTI suppression is critical for host colonization and subsequent disease development. This review summarizes molecular basis of bacterial pathogen perception by plant PRRs and also interaction between PRRs and type III effectors during early stages of plant-pathogen interaction.

Similar content being viewed by others
References
Abramovitch RB, Janjusevic R, Stebbins CE, Martin GB (2006) Type III effector AvrPtoB requires intrinsic E3 ubiquitin ligase activity to suppress plant cell death and immunity. Proc Natl Acad Sci USA 103:2851–2856
Albert M, Jehle AK, Mueller K, Eisele C, Lipschis M, Felix G (2010) The Arabidopsis thaliana pattern recognition receptors for bacterial elongation factor Tu and flagellin can be combined to form functional chimeric receptors. J Biol Chem 285:19035–19042
Albert I, Böhm H, Albert M, Feiler CE, Imkampe J, Wallmeroth N, Brancato C, Raaymakers TM, Oome S, Zhang H, Krol E (2015) An RLP23-SOBIR1-BAK1 complex mediates NLP-triggered immunity. Nat Plants 1:15140
Alfano JR, Collmer A (2004) Type III secretion system effector proteins: double agents in bacterial disease and plant defense. Annu Rev Phytopathol 42:385–414
Almeida NF, Yan S, Lindeberg M, Studholme DJ, Schneider DJ, Condon B, Liu H, Viana CJ, Warren A, Evans C, Kemen E (2009) A draft genome sequence of Pseudomonas syringae pv. tomato T1 reveales a type III effector repertoire significantly divergent from that of Pseudomonas syringae pv. tomato DC3000. Mol Plant-Microbe Interact 22:52–62
Balagué C, Gouget A, Bouchez O, Souriac C, Haget N, Boutet-Mercey S, Govers F, Roby D, Canut H (2017) The Arabidopsis thaliana lectin receptor kinase LecRK-I. 9 is required for full resistance to Pseudomonas syringae and affects jasmonate signalling. Mol Plant Pathol 18:937–948
Baltrus DA, Nishimura MT, Romanchuk A, Chang JH, Mukhtar MS, Cherkis K, Roach J, Grant SR, Jones CD, Dangl JL (2011) Dynamic evolution of pathogenicity revealed by sequencing and comparative genomics of 19 Pseudomonas syringae isolates. PLoS Pathog 7:e1002132
Bi G, Liebrand TW, Cordewener JH, America AH, Xu X, Joosten MH (2014) Arabidopsis thaliana receptor-like protein At RLP23 associates with the receptor-like kinase At SOBIR1. Plant Signal Behav 9:e27937
Boudsocq M, Willmann MR, McCormack M, Lee H, Shan L, He P, Bush J, Sheen J (2010) Differential innate immune signalling via Ca2+ sensor protein kinases. Nature 464:418
Brutus A, Sicilia F, Macone A, Cervone F, De Lorenzo G (2010) A domain swap approach reveals a role of the plant wall-associated kinase 1 (WAK1) as a receptor of oligogalacturonides. Proc Natl Acad Sci USA 107:9452–9457
Büttner D (2016) Behind the lines–actions of bacterial type III effector proteins in plant cells. FEMS Microbiol Rev 40:894–937
Cao Y, Liang Y, Tanaka K, Nguyen CT, Jedrzejczak RP, Joachimiak A, Stacey G (2014) The kinase LYK5 is a major chitin receptor in Arabidopsis and forms a chitin-induced complex with related kinase CERK1. Elife 3:e03766
Castañeda-Ojeda MP, Moreno-Pérez A, Ramos C, López-Solanilla E (2017) Suppression of plant immune responses by the Pseudomonas savastanoi pv. savastanoi NCPPB 3335 type III effector tyrosine phosphatases HopAO1 and HopAO2. Front Plant Sci 8:680
Choi J, Tanaka K, Cao Y, Qi Y, Qiu J, Liang Y, Lee SY, Stacey G (2014) Identification of a plant receptor for extracellular ATP. Science 343:290–294
Cunnac S, Chakravarthy S, Kvitko BH, Russell AB, Martin GB, Collmer A (2011) Genetic disassembly and combinatorial reassembly identify a minimal functional repertoire of type III effectors in Pseudomonas syringae. Proc Natl Acad Sci USA 108:2975–2980
Deng W, Marshall NC, Rowland JL, McCoy JM, Worrall LJ, Santos AS, Strynadka NC, Finlay BB (2017) Assembly, structure, function and regulation of type III secretion systems. Nat Rev Microbiol 15:323
Erbs G, Silipo A, Aslam S, De Castro C, Liparoti V, Flagiello A, Pucci P, Lanzetta R, Parrilli M, Molinaro A, Newman MA (2008) Peptidoglycan and muropeptides from pathogens Agrobacterium and Xanthomonas elicit plant innate immunity: structure and activity. Chem Biol 15:438–448
Espinoza C, Liang Y, Stacey G (2017) Chitin receptor CERK 1 links salt stress and chitin-triggered innate immunity in Arabidopsis. Plant J 89:984–995
Felix G, Duran JD, Volko S, Boller T (1999) Plants have a sensitive perception system for the most conserved domain of bacterial flagellin. Plant J 18:265–276
Frei dit Frey N, Mbengue M, Kwaaitaal M, Nitsch L, Altenbach D, Häweker H, Lozano-Duran R, Njo M, Beeckman T, Huettel B, Borst JW (2012) Plasma membrane calcium ATPases are important components of receptor-mediated signaling in plant immune responses and development. Plant Physiol 159:798–809
Fritz-Laylin LK, Krishnamurthy N, Tör M, Sjölander KV, Jones JD (2005) Phylogenomic analysis of the receptor-like proteins of rice and Arabidopsis. Plant Physiol 138:611–623
Gimenez-Ibanez S, Ntoukakis V, Rathjen JP (2009a) The LysM receptor kinase CERK1 mediates bacterial perception in Arabidopsis. Plant Signal Behav 4:539–541
Gimenez-Ibanez S, Hann DR, Ntoukakis V, Petutschnig E, Lipka V, Rathjen JP (2009b) AvrPtoB targets the LysM receptor kinase CERK1 to promote bacterial virulence on plants. Curr Biol 19:423–429
Göhre V, Spallek T, Häweker H, Mersmann S, Mentzel T, Boller T, de Torres M, Mansifield JW, Robatzek S (2008) Plant pattern-recognition receptor FLS2 is directed for degradation by the bacterial ubiquitin ligase AvrPtoB. Curr Biol 18:1824–1832
Gust AA, Biswas R, Lenz HD, Rauhut T, Ranf S, Kemmerling B, Gotz F, Glawischnig E, Keem J, Felix G, Nürnberger T (2007) Bacteria-derived peptidoglycans constitute pathogen-associated molecular patterns triggering innate immunity in Arabidopsis. J Biol Chem 282:32338–32348
Heese A, Hann DR, Gimenez-Ibanez S, Jones AM, He K, Li J, Schroeder JI, Peck SC, Rathjen JP (2007) The receptor-like kinase SERK3/BAK1 is a central regulator of innate immunity in plants. Proc Natl Acad Sci USA 104:12217–12222
Hou S, Wang X, Chen D, Yang X, Wang M, Turrà D, Di Pietro A, Zhang W (2014) The secreted peptide PIP1 amplifies immunity through receptor-like kinase 7. PLoS Pathog 10:e1004331
Hurley B, Lee D, Mott A, Wilton M, Liu J, Liu YC, Angers S, Coaker G, Desveaux D (2014) The Pseudomonas syringae type III effector HopF2 suppresses Arabidopsis stomatal immunity. PLoS One 9:e114921
Janjusevic R, Abramovitch RB, Martin GB, Stebbins CE (2006) A bacterial inhibitor of host programmed cell death defenses is an E3 ubiquitin ligase. Science 311:222–226
Jehle AK, Lipschis M, Albert M, Fallahzadeh-Mamaghani V, Fürst U, Mueller K, Felix G (2013) The receptor-like protein ReMAX of Arabidopsis detects the microbe-associated molecular pattern eMax from Xanthomonas. Plant Cell 25:2330–2340
Kadota Y, Sklenar J, Derbyshire P, Stransfeld L, Asai S, Ntoukakis V, Jones JD, Shirasu K, Menke F, Jones A, Zipfel C (2014) Direct regulation of the NADPH oxidase RBOHD by the PRR-associated kinase BIK1 during plant immunity. Mol Cell 54:43–55
Kim YJ, Lin NC, Martin GB (2002) Two distinct Pseudomonas effector proteins interact with the Pto kinase and activate plant immunity. Cell 109:589–598
Kunze G, Zipfel C, Robatzek S, Niehaus K, Boller T, Felix G (2004) The N terminus of bacterial elongation factor Tu elicits innate immunity in Arabidopsis plants. Plant Cell 16:3496–3507
Lacombe S, Rougon-Cardoso A, Sherwood E, Peeters N, Dahlbeck D, Van Esse HP, Smoker M, Rallapalli G, Thomma BP, Staskawicz B, Jones JD (2010) Interfamily transfer of a plant pattern-recognition receptor confers broad-spectrum bacterial resistance. Nat Biotechnol 28:365
Li L, Li M, Yu L, Zhou Z, Liang X, Liu Z, Cai G, Gao L, Zhang X, Wang Y, Chen S (2014) The FLS2-associated kinase BIK1 directly phosphorylates the NADPH oxidase RbohD to control plant immunity. Cell Host Microbe 15:329–338
Lindeberg M (2012) Genome-enabled perspectives on the composition, evolution, and expression of virulence determinants in bacterial plant pathogens. Annu Rev Phytopathol 50:111–132
Lindeberg M, Cunnac S, Collmer A (2012) Pseudomonas syringae type III effector repertoires: last words in endless arguments. Trends Microbiol 20:199–208
Liu T, Liu Z, Song C, Hu Y, Han Z, She J, Fan F, Wang J, Jin C, Chang J, Zhou JM (2012) Chitin-induced dimerization activates a plant immune receptor. Science 336:1160–1164
Macho AP, Schwessinger B, Ntoukakis V, Brutus A, Segonzac C, Roy S, Kadota Y, Oh MH, Sklenar J, Derbyshire P, Lozano-Durán R (2014) A bacterial tyrosine phosphatase inhibits plant pattern recognition receptor activation. Science 343:1509–1512
Miya A, Albert P, Shinya T, Desaki Y, Ichimura K, Shirasu K, Narusaka Y, Kawakami N, Kaku H, Shibuya N (2007) CERK1, a LysM receptor kinase, is essential for chitin elicitor signaling in Arabidopsis. Proc Natl Acad Sci USA 104:19613–19618
Mott GA, Thakur S, Smakowska E, Wang PW, Belkhadir Y, Desveaux D, Guttman DS (2016) Genomic screens identify a new phytobacterial microbe-associated molecular pattern and the cognate Arabidopsis receptor-like kinase that mediates its immune elicitation. Genome Biol 17:98
Nicaise V, Joe A, Jeong BR, Korneli C, Boutrot F, Westedt I, Staiger D, Alfano JR, Zipfel C (2013) Pseudomonas HopU1 modulates plant immune receptor levels by blocking the interaction of their mRNAs with GRP7. EMBO J 32:701–712
Oh MH, Wang X, Kota U, Goshe MB, Clouse SD, Huber SC (2009) Tyrosine phosphorylation of the BRI1 receptor kinase emerges as a component of brassinosteroid signaling in Arabidopsis. Proc Natl Acad Sci USA 106:658–663
Petnicki-Ocwieja T, Schneider DJ, Tam VC, Chancey ST, Shan L, Jamir Y, Schechter LM, Janes MD, Buell CR, Tang X, Collmer A (2002) Genome-wide identification of proteins secreted by the Hrp type III protein secretion system of Pseudomonas syringae pv. tomato DC3000. Proc Natl Acad Sci USA. 99:7652–7657
Postel S, Küfner I, Beuter C, Mazzotta S, Schwedt A, Borlotti A, Halter T, Kemmerling B, Nürnberger T (2010) The multifunctional leucine-rich repeat receptor kinase BAK1 is implicated in Arabidopsis development and immunity. Eur J Cell Biol 89:169–174
Ranf S, Gisch N, Schäffer M, Illig T, Westphal L, Knirel YA, Sanchez-Carballo PM, Zahringer U, Huckelhoven R, Lee J, Scheel D (2015) A lectin S-domain receptor kinase mediates lipopolysaccharide sensing in Arabidopsis thaliana. Nat Immunol 16:426
Ronald PC, Salmeron JM, Carland FM, Staskawicz BJ (1992) The cloned avirulence gene avrPto induces disease resistance in tomato cultivars containing the Pto resistance gene. J Bacteriol 174:1604–1611
Roux M, Schwessinger B, Albrecht C, Chinchilla D, Jones A, Holton N, Malinovsky FG, Tor M, de Vries S, Zipfel C (2011) The Arabidopsis leucine-rich repeat receptor-like kinases BAK1/SERK3 and BKK1/SERK4 are required for innate immunity to hemibiotrophic and biotrophic pathogens. Plant Cell 23:2440–2455
Saijo Y, Loo EPI, Yasuda S (2018) Pattern recognition receptors and signaling in plant-microbe interactions. Plant J 93:592–613
Schulze B, Mentzel T, Jehle AK, Mueller K, Beeler S, Boller T, Felix G, Chinchilla D (2010) Rapid heteromerization and phosphorylation of ligand-activated plant transmembrane receptors and their associated kinase BAK1. J Biol Chem 285:9444–9451
Schwessinger B, Ronald PC (2012) Plant innate immunity: perception of conserved microbial signatures. Annu Rev Plant Biol 63:451–482
Schwessinger B, Bahar O, Thomas N, Holton N, Nekrasov V, Ruan D, Canlas PE, Daudi A, Petzold CJ, Singan VR, Kuo R (2015) Transgenic expression of the dicotyledonous pattern recognition receptor EFR in rice leads to ligand-dependent activation of defense responses. PLoS Pathog 11:e1004809
Shan L, Thara VK, Martin GB, Zhou JM, Tang X (2000) The Pseudomonas AvrPto protein is differentially recognized by tomato and tobacco and is localized to the plant plasma membrane. Plant Cell 12:2323–2337
Shan L, He P, Li J, Heese A, Peck SC, Nürnberger T, Martin GB, Sheen J (2008) Bacterial effectors target the common signaling partner BAK1 to disrupt multiple MAMP receptor-signaling complexes and impede plant immunity. Cell Host Microbe 4:17–27
Shinya T, Yamaguchi K, Desaki Y, Yamada K, Narisawa T, Kobayashi Y, Maeda K, Suzuki M, Tanimoto T, Takeda J, Nakashima M (2014) Selective regulation of the chitin-induced defense response by the Arabidopsis receptor-like cytoplasmic kinase PBL 27. Plant J 79:56–66
Sugiyama N, Nakagami H, Mochida K, Daudi A, Tomita M, Shirasu K, Ishihama Y (2008) Large-scale phosphorylation mapping reveals the extent of tyrosine phosphorylation in Arabidopsis. Mol Syst Biol 4:193
Wang G, Ellendorff U, Kemp B, Mansfield JW, Forsyth A, Mitchell K, Mitchell K, Bastas K, Liu CM, Woods-Tor A, Zipfel C, De Wit PJ (2008) A genome-wide functional investigation into the roles of receptor-like proteins in Arabidopsis. Plant Physiol 147:503–517
Wang Y, Li J, Hou S, Wang X, Li Y, Ren D, Chen S, Tang X, Zhou JM (2010) A Pseudomonas syringae ADP-ribosyltransferase inhibits Arabidopsis mitogen-activated protein kinase kinases. Plant Cell 22:2033–2044
Willmann R, Lajunen HM, Erbs G, Newman MA, Kolb D, Tsuda K, Katagiri F, Fliegmann J, Bono JJ, Cullimore JV, Jehle AK (2011) Arabidopsis lysin-motif proteins LYM1 LYM3 CERK1 mediate bacterial peptidoglycan sensing and immunity to bacterial infection. Proc Natl Acad Sci USA 108:19824–19829
Xiang T, Zong N, Zou Y, Wu Y, Zhang J, Xing W, Li Y, Tang X, Zhu L, Chai J, Zhou JM (2008) Pseudomonas syringae effector AvrPto blocks innate immunity by targeting receptor kinases. Curr Biol 18:74–80
Xiang T, Zong N, Zhang J, Chen J, Chen M, Zhou JM (2011) BAK1 is not a target of the Pseudomonas syringae effector AvrPto. Mol Plant Microbe Interact 24:100–107
Xing W, Zou Y, Liu Q, Liu J, Luo X, Huang Q, Chen S, Zhu L, Bi R, Hao Q, Wu JW (2007) The structural basis for activation of plant immunity by bacterial effector protein AvrPto. Nature 449:243
Yamada K, Yamaguchi K, Shirakawa T, Nakagami H, Mine A, Ishikawa K, Fujiwara M, Narusaka M, Narusaka Y, Ichimura K, Kobayashi Y (2016) The Arabidopsis CERK1-associated kinase PBL27 connects chitin perception to MAPK activation. EMBO J 35:2468–2483
Yamaguchi Y, Pearce G, Ryan CA (2006) The cell surface leucine-rich repeat receptor for AtPep1, an endogenous peptide elicitor in Arabidopsis, is functional in transgenic tobacco cells. Proc Natl Acad Sci USA 103:10104–10109
Yamaguchi Y, Huffaker A, Bryan AC, Tax FE, Ryan CA (2010) PEPR2 is a second receptor for the Pep1 and Pep2 peptides and contributes to defense responses in Arabidopsis. Plant Cell 22:508–522
Yeh YH, Panzeri D, Kadota Y, Huang YC, Huang PY, Tao CN, Roux M, Chien HC, Chin TC, Chu PW, Zipfel C (2016) The Arabidopsis malectin-like/LRR-RLK IOS1 is critical for BAK1-dependent and BAK1-independent pattern-triggered immunity. Plant Cell 28:1701–1721
Zhang W, Fraiture M, Kolb D, Löffelhardt B, Desaki Y, Boutrot FF, Tor M, Zipfel C, Gust AA, Brunner F (2013) Arabidopsis receptor-like protein30 and receptor-like kinase suppressor of BIR1-1/EVERSHED mediate innate immunity to necrotrophic fungi. Plant Cell 25:4227–4241
Zhou J, Wu S, Chen X, Liu C, Sheen J, Shan L, He P (2014) The Pseudomonas syringae effector HopF2 suppresses Arabidopsis immunity by targeting BAK1. Plant J 77:235–245
Zipfel C (2009) Early molecular events in PAMP-triggered immunity. Curr Opin Plant Biol 12:414–420
Zipfel C (2014) Plant pattern-recognition receptors. Trends Immunol 35:345–351
Zipfel C, Kunze G, Chinchilla D, Caniard A, Jones JD, Boller T, Felix G (2006) Perception of the bacterial PAMP EF-Tu by the receptor EFR restricts Agrobacterium-mediated transformation. Cell 125:749–760
Acknowledgements
This work was supported in part by the Basic Science Research Program through the National Research Foundation of Korea (NRF), funded by the Ministry of Education (No. NRF-2017R1A2B4004620), and by Korea Institute of Planning and Evaluation for Technology in Food, Agriculture, Forestry and Fisheries (IPET) through the Golden Seed Project, Ministry of Agriculture, Food and Rural Affairs (MAFRA) (213006-05-3-SBC30).
Author information
Authors and Affiliations
Corresponding author
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Lee, J.H., Kim, H., Chae, W.B. et al. Pattern recognition receptors and their interactions with bacterial type III effectors in plants. Genes Genom 41, 499–506 (2019). https://doi.org/10.1007/s13258-019-00801-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13258-019-00801-1