Abstract
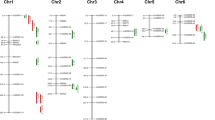
Micro satellite markers located in the Saltol QTL of 5 Mb region (10.4–15.6 Mb) in chromosome 1 confering seedling stage salt tolerance were used to evaluate 94 rice genotypes. Out of 21, eight SSR markers at Saltol region of Chromosome were found polymorphic. Based on the phenotypic screening, 94 genotypes were grouped as highly tolerant (20), tolerant (18) moderately tolerant (32), sensitive (19) and highly sensitive (5). The marker RM3412 appears to be diagnostic of salinity tolerance and associate to salinity tolerance at seedling stage as it is closely linked to SKC gene. Based on Saltol markers study, CSR 31, CSR 38, CSR 41, CSR 32, Wild 11, CSR 18, Azgo, Pant Dhan 4, Trichi 1, CSR 10 and IR64426-4B-11-1 could not be identified as tolerant genotypes though had expressed tolerant to highly tolerant phenotype to salinity stress at seedling stage, suggesting that QTLs other than Saltol might be controlling their salinity tolerance. It is suggested that these genotypes could serve as potentially novel germplasm and could be exploited for the development of new breeding lines with high level of salinity tolerance by pyramiding of the Saltol and other QTLs.




Similar content being viewed by others
Abbreviations
- SSR:
-
Simple sequence repeats
- QTL:
-
Quantitative trait loci
- RM:
-
Rice microsatellite
- PCR:
-
Polymerase chain reaction
- CSR:
-
Central Soil Salinity Research Institute rice line
- IR:
-
International Rice Research Institute rice line
References
Akbar M, Yabuno T, Nakao S (1972) Breeding for saline resistant varieties of rice. I. Variability for salt tolerance among some rice varieties. Jpn J Breed 22:277–284
Bonilla P, Dvorak J, Mackill D, Deal K, Gregorio G (2002) RFLP and SSLP mapping of salinity tolerance genes in chromosome 1 of rice (Oryzasativa L.) using recombinant inbred lines. Philipp Agric Sci 85:68–76
Doyle JJ, Doyle JL (1990) Isolation of plant DNA from fresh tissue. Focus 12:13–15
FAO (2006) World agriculture towards 2030/2050. FAO, Rome
FAO (2008) FAO Land and Plant Nutrition Management Service.Available at http://www.fao.org/ag/agl/agll/spush. Verified 24 May 2010
Flowers TJ, Yeo AR (1981) Variability in the resistance of sodium chloride salinity within rice (Oryzasativa L.) varieties. New Phytol 88:363–373
Gregorio GB, Senadhira D (1993) Genetic analysis of salinity tolerance in rice. Theor Appl Genet 86:333–338
Gregorio GB, Senadhira D, Mendoza RD (1997) Screening rice for salinity tolerance. IRRI Discussion Paper Series no. 22. Manila (Philippines), International Rice Research Institute, pp. 30
Gregorio GB, Senadhira D, Mendoza RD, Manigbas NL, Roxas JP, Guerta CQ (2002) Progress in breeding for salinity tolerance and associated abiotic stresses in rice. Field Crop Res 76:91–101
Hasegawa PM, Bressan RA, Zhu JK, Bonhert HJ (2000) Plant cellular and molecular responses to high salinity. Annu Rev Plant Physiol Plant Mol Biol 51:463–499
Islam MR, Salam MA, Hassan L, Collard BCY, Singh RK, Gregorio GB (2011) QTL mapping for salinity tolerance at seedling stage in rice. Emirates J Food Agric 23(2):137–146
Islam MR, Gregorio GB, Salam MA, Collard BCY, Singh RK, Hassan L (2012) Validation of SalTolLinked markers and haplotype diversity on chromosome 1 of rice. Mol Plant Breed 10(3):103–114
Ismail AM, Heue RS, Thomson MJ, Wissuwa M (2007) Genetic and genomic approaches to develop rice germplasm for problem soils. Plant Mol Biol 65:547–570
Ismail AM, Thomson MJ, Vergara GV, Rahman MA, Singh RK, Gregorio GB (2010) Designing resilient rice varieties for coastal deltas using modern breeding tools. In: Hoanh CT, Szuster BW, Pheng KS, Ismail AM, Nobel AD (eds) Tropical Deltas and coastal zones, food production, communities and environment at the land-water interface. CAB, Wallingford, pp 154–165
Krishnamurthy SL, Sharma SK, Kumar V, Tiwari S, Batra V, Singh NK (2014) Assessment of genetic diversity in rice genotypes for salinity tolerance using Saltol markers of Chromosome 1. Indian J Genet Plant Breed 74(2):243–247
Lin HX, Zhu MZ, Yano M, Gao JP, Liang ZW, Su WA (2004) QTLs for Na+ and K+ uptake of the shoots and roots controlling rice salt tolerance. Theor Appl Genet 108:253–260
Mohammadi-Nejad G, Arzani A, Rezai AM, Singh RK, Gregorio GB (2008) Assessment of rice genotypes for salt tolerance using microsatellite markers associated with the saltolQTL. Afr J Biotechnol 7:730–736
Naresh Babu N, Vinod KK, Gopala Krishnan S, Bhowmick PK, Vanaja T, Krishnamurthy SL, Nagarajan M, Singh NK, Prabhu KV, Singh AK (2014) Marker based haplotype diversity of Saltol QTL in relation to seedling stage salinity tolerance in selected genotypes of rice. Indian J Genet 74(1):16–25
Niones JM (2004) Fine mapping of the salinity tolerance gene on chromosome 1 of rice (Oryzasativa L.) using near isogenic lines, M. S. dissertation, College, Laguna, Philippines: University of the Philippines Los Baños, Laguna
Pandit A, Rai V, Bal S, Sinha S, Kumar V, Chauhan M, Gautam RK, Singh RK, Sharma PC, Singh AK, Gaikwad K, Sharma TR, Mohapatra T, Singh NK (2010) Combining QTL mapping and transcriptome profiling of bulked RILs for identification of functional polymorphism for salt tolerance genes in rice (Oryzasativa L.). Mol Genet Genomics 284(2):121–136
Pillai AM, Susan EK, Yasin J, Anandhi K, Selvi B (2011) Genetic variability and association studies in salt tolerant rice mutant. Int J Plant Breed Genet 5:339–348
Sharma SK (1986) Mechanisms of tolerance in rice varieties differing in sodicity tolerance. Plant Soil 93:141–145
Sharma SK (2010) Salt tolerant crop varieties developed at CSSRI Karnal, India: a success story. www.PlantStress.com. pp. 14
Thomson MJ, Ismail AM, McCouch SR, Mackill MJ (2010) Marker assisted breeding. In: Pareek A, Sopory SK, Bohnert HJ, Govindjee (eds) Abiotic stress adaptation in plants, physiological, molecular and genomic foundation. Springer, New York, pp 451–469
Yamaguchi Y, Blumwald E (2005) Developing salt-tolerant crop plants, challenges and opportunities. Trends Plant Sci 10:615–620
Yoshida S, Forno DA, Cook JH, Gomez KA (1976) Laboratory manual for physiological studies of rice. International Rice Research Institute (IRRI), LosBaños, pp 61–66
Acknowledgments
Authors express sincere thanks to the Indian Council of Agricultural Research, New Delhi for funding the National Network Project on Transgenic Crops –Functional Genomics in Rice (NPTC) and Director, CSSRI, Karnal for encouragement.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Supplementary table 1
(DOCX 25 kb)
Supplementary table 2
(DOCX 15 kb)
Rights and permissions
About this article
Cite this article
Krishnamurthy, S.L., Sharma, S.K., Kumar, V. et al. Analysis of genomic region spanning Saltol using SSR markers in rice genotypes showing differential seedlings stage salt tolerance. J. Plant Biochem. Biotechnol. 25, 331–336 (2016). https://doi.org/10.1007/s13562-015-0335-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13562-015-0335-5