Abstract
Introduction
This work describes the Medication and Adverse Drug Events from Electronic Health Records (MADE 1.0) corpus and provides an overview of the MADE 1.0 2018 challenge for extracting medication, indication, and adverse drug events (ADEs) from electronic health record (EHR) notes.
Objective
The goal of MADE is to provide a set of common evaluation tasks to assess the state of the art for natural language processing (NLP) systems applied to EHRs supporting drug safety surveillance and pharmacovigilance. We also provide benchmarks on the MADE dataset using the system submissions received in the MADE 2018 challenge.
Methods
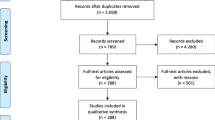
The MADE 1.0 challenge has released an expert-annotated cohort of medication and ADE information comprising 1089 fully de-identified longitudinal EHR notes from 21 randomly selected patients with cancer at the University of Massachusetts Memorial Hospital. Using this cohort as a benchmark, the MADE 1.0 challenge designed three shared NLP tasks. The named entity recognition (NER) task identifies medications and their attributes (dosage, route, duration, and frequency), indications, ADEs, and severity. The relation identification (RI) task identifies relations between the named entities: medication-indication, medication-ADE, and attribute relations. The third shared task (NER-RI) evaluates NLP models that perform the NER and RI tasks jointly. In total, 11 teams from four countries participated in at least one of the three shared tasks, and 41 system submissions were received in total.
Results
The best systems F1 scores for NER, RI, and NER-RI were 0.82, 0.86, and 0.61, respectively. Ensemble classifiers using the team submissions improved the performance further, with an F1 score of 0.85, 0.87, and 0.66 for the three tasks, respectively.
Conclusion
MADE results show that recent progress in NLP has led to remarkable improvements in NER and RI tasks for the clinical domain. However, some room for improvement remains, particularly in the NER-RI task.
Similar content being viewed by others
Notes
The complete annotation guideline and dataset is available at bio-nlp.org/dataset/made1.
The evaluation script is included with the MADE data release.
References
Donaldson MS, Corrigan JM, Kohn LT, et al. To err is human: building a safer health system, vol. 6. Washington: National Academies Press; 2000.
Bates DW, Cullen DJ, Laird N, Petersen LA, Small SD, Servi D, Laffel G, Sweitzer BJ, Shea BF, Hallisey R, et al. Incidence of adverse drug events and potential adverse drug events: implications for prevention. JAMA. 1995;274(1):29–34.
Lazarou J, Pomeranz BH, Corey PN. Incidence of adverse drug reactions in hospitalized patients: a meta-analysis of prospective studies. JAMA. 1998;279(15):1200–5.
Bates DW, Spell N, Cullen DJ, Burdick E, Laird N, Petersen LA, Small SD, Sweitzer BJ, Leape LL. The costs of adverse drug events in hospitalized patients. JAMA. 1997;277(4):307–11.
Nebeker JR, Hoffman JM, Weir CR, Bennett CL, Hurdle JF. High rates of adverse drug events in a highly computerized hospital. Arch Intern Med. 2005;165(10):1111–6.
Gurwitz JH, Field TS, Harrold LR, Rothschild J, Debellis K, Seger AC, Cadoret C, Fish LS, Garber L, Kelleher M, et al. Incidence and preventability of adverse drug events among older persons in the ambulatory setting. JAMA. 2003;289(9):1107–16.
Johnson J, Booman L. Drug-related morbidity and mortality. J Manag Care Pharm. 1996;2(1):39–47.
Haas JS, Iyer A, Orav EJ, Schiff GD, Bates DW. Participation in an ambulatory e-pharmacovigilance system. Pharmacoepidemiol Drug Saf. 2010;19(9):961–9.
Frank C, Himmelstein DU, Woolhandler S, Bor DH, Wolfe SM, Heymann O, Zallman L, Lasser KE. Era of faster FDA drug approval has also seen increased black-box warnings and market withdrawals. Health Aff. 2014;33(8):1453–9.
WHO. WHO | Pharmacovigilance; 2017. http://www.who.int/medicines/areas/quality_safety/safety_efficacy/pharmvigi/en/. Accessed 10 May 2018.
Edlavitch SA. Adverse drug event reporting: improving the low us reporting rates. Arch Intern Med. 1988;148(7):1499–503.
Hasford J, Goettler M, Munter K-H, Müller-Oerlinghausen B. Physicians’ knowledge and attitudes regarding the spontaneous reporting system for adverse drug reactions. J Clin Epidemiol. 2002;55(9):945–50.
Begaud B, Moride Y, Tubert-Bitter P, Chaslerie A, Haramburu F. False-positives in spontaneous reporting: should we worry about them? Br J Clin Pharmacol. 1994;38(5):401–4.
Xu R, Wang Q. Comparing a knowledge-driven approach to a super-vised machine learning approach in large-scale extraction of drug-side effect relation-ships from free-text biomedical literature. BMC Bioinform. 2015;16:S6.
Butt TF, Cox AR, Oyebode JR, Ferner RE. Internet accounts of serious adverse drug reactions. Drug Saf. 2012;35(12):1159–70.
Rossi AC, Knapp DE, Anello C, O’Neill RT, Graham CF, Mendelis PS, Stanley GR. Discovery of adverse drug reactions: a comparison of selected phase IV studies with spontaneous reporting methods. JAMA. 1983;249(16):2226–8.
Lardon J, Abdellaoui R, Bellet F, Asfari H, Souvignet J, Texier N, Jaulent M-C, Beyens M-N, Burgun A, Bousquet C. Adverse drug reaction identification and extraction in social media: a scoping review. J Med Internet Res. 2015;17(7):e171.
Smythe MA, Fanikos J, Gulseth MP, Wittkowsky AK, Spinler SA, Dager WE, Nutescu EA. Rivaroxaban: practical consider-ations for ensuring safety and efficacy. Pharmacotherapy. 2013;33(11):1223–45.
McGraw D, Rosati K, Evans B. A policy framework for public health uses of electronic health data. Pharmacoepidemiol Drug Saf. 2012;21(S1):18–22.
Yih WK, Lieu TA, Kulldorff M, Martin D, McMahill-Walraven CN, Platt R, Selvam N, Selvan M, Lee GM, Nguyen M. Intussusception risk after rotavirus vaccination in us infants. N Engl J Med. 2014;370(6):503–51.
Peissig PL, Costa VS, Caldwell MD, Rottscheit C, Berg RL, Mendonca EA, Page D. Relational machine learning for electronic health record-driven phenotyping. J Biomed Informat. 2014;52:260–70.
Wu J, Roy J, Stewart WF. Prediction modeling using EHR data: challenges, strategies, and a comparison of machine learning approaches. Med Care. 2010;48:S106–13.
Jha AK, Kuperman GJ, Teich JM, Leape L, Shea B, Rittenberg E, Burdick E, Seger DL, Vliet MV, Bates DW. Identifying adverse drug events: development of a computer-based monitor and comparison with chart review and stimulated voluntary report. J Am Med Inform Assoc. 1998;5(3):305–14.
Skentzos S, Shubina M, Plutzky J, Turchin A. Structured vs. unstructured: factors affecting adverse drug reaction documentation in an EMR repository. In: AMIA annual symposium proceedings, vol. 2011. American Medical Informatics Association.
Schulman S, Kearon C. Subcommittee on Control of Anticoagulation of the Scientific, Standardization Committee of the International Society on Thrombosis, and Haemostasis. Definition of major bleeding in clinical investigations of antihemostatic medicinal products in non-surgical patients. J Thromb Haemost. 2005;3(4):692–4.
Murtaugh MA, Gibson BS, Redd D, Zeng-Treitler Q. Regular expression-based learning to extract bodyweight values from clinical notes. J Biomed Inform. 2015;54:186–90.
Classen DC, Pestotnik SL, Evans RS, Burke JP. Computerized surveillance of adverse drug events in hospital patients. BMJ Qual Saf. 2005;14(3):221–6.
Aronson AR. Effective mapping of biomedical text to the umls metathesaurus: the metamap program. In: Proceedings of the AMIA symposium, p. 17. American Medical Informatics Association; 2001.
Xu H, Stenner SP, Doan S, Johnson KB, Waitman LR, Denny JC. Medex: a medication information extraction system for clinical narratives. J Am Med Inform Assoc. 2010;17(1):19–24.
Friedman C, Kra P, Yu H, Krauthammer M, Rzhetsky A. Genies: a natural-language processing system for the extraction of molecular pathways from journal articles. In: ISMB (supplement of bioinformatics), p. 74–82; 2001.
Hahn U, Romacker M, Schulz S. Creating knowledge repositories from biomedical reports: the medsyndikate text mining system. In: Biocomputing 2002, pp. 338–349. World Scientific; 2001.
Hong Y, Lee M. Accessing bioscience images from abstract sentences. Bioinformatics. 2006;22(14):e547–56.
Yu H. Towards answering biological questions with experimental evidence: automatically identifying text that summarize image content in full-text articles. In: AMIA annual symposium proceedings, vol. 2006, p. 834. American Medical Informatics Association; 2006.
Kim J-D, Ohta T, Pyysalo S, Kano Y, Tsujii J. Overview of BioNLP’09 shared task on event extraction. In: Proceedings of the workshop on current trends in biomedical natural language processing: shared task, pp. 1–9. Association for Computational Linguistics; 2009.
Hirschman L, Yeh A, Blaschke C, Valencia A. Overview of biocreative: critical assessment of information extraction for biology; 2005.
Li Z, Cao Y, Antieau L, Agarwal S, Zhang Q, Yu H. Extracting medication information from patient discharge summaries. In: Proceedings of the third i2b2 workshop on challenges in natural language processing for clinical data; 2009.
Pradhan S, Elhadad N, South BR, Martinez D, Christensen L, Vogel A, Suominen H, Chapman WW, Savova G. Evaluating the state of the art in disorder recognition and normalization of the clinical narrative. J Am Med Inform Assoc. 2014;22(1):143–54.
Li Q, Melton K, Lingren T, Kirkendall ES, Hall E, Zhai H, Ni Y, Kaiser M, Stoutenborough L, Solti I. Phenotyping for patient safety: algorithm development for electronic health record based automated adverse event and medical error detection in neonatal intensive care. J Am Med Inform Assoc. 2014;21(5):776–84.
Melton GB, Hripcsak G. Automated detection of adverse events using natural language processing of discharge summaries. J Am Med Inform Assoc. 2005;12(4):448–57.
Humphreys BL, Lindberg DAB, Schoolman HM, Barnett GO. The unified medical language system: an informatics research collaboration. J Am Med Inform Assoc. 1998;5(1):1–11.
Bodenreider O. The unified medical language system (UMLS): integrating biomedical terminology. Nucleic Acids Res. 2004;32(suppl 1):D267–70.
Rochefort CM, Verma AD, Eguale T, Lee TC, Buckeridge DL. A novel method of adverse event detection can accurately identify venous thromboembolisms (VTES) from narrative electronic health record data. J Am Med Inform Assoc. 2014;22(1):155–65.
Haerian K, Salmasian H, Friedman C. Methods for identifying suicide or suicidal ideation in EHRS. In: AMIA annual symposium proceedings, vol. 2012, p. 1244. American Medical Informatics Association; 2012.
Wang S, Li Y, Ferguson D, Zhai C. Side effect PTM: an unsupervised topic model to mine adverse drug reactions from health forums. In: Proceedings of the 5th ACM conference on bioinformatics, computational biology, and health informatics, p. 321–330. ACM; 2014.
Nikfarjam Azadeh, Sarker Abeed, O’Connor Karen, Ginn Rachel, Gon-zalez Graciela. Pharmacovigilance from social media: mining adverse drug reaction mentions using sequence labeling with word embedding cluster features. J Am Med Inform Assoc. 2015;22(3):671–81.
Li Q, Deleger L, Lingren T, Zhai H, Kaiser M, Stoutenborough L, Jegga AG, Cohen KB, Solti I. Mining FDA drug labels for medical conditions. BMC Med Inform Decis Making. 2013;13(1):53.
Duke JD, Friedlin J. ADESSA: a real-time decision support service for de-livery of semantically coded adverse drug event data. In: AMIA Annual symposium proceedings, vol. 2010, p. 177. American Medical Informatics Association; 2010.
Kim J-D, Ohta T, Tateisi Y, Tsujii J. Genia corpus—a semantically annotated corpus for bio-textmining. Bioinformatics. 2003;19(suppl 1):i180–2.
Cohen AM, Hersh WR. The TREC 2004 genomics track categorization task: classifying full text biomedical documents. J Biomed Discov Collab. 2006;1(1):4.
Doğan RI, Lu Z. An improved corpus of disease mentions in Pubmed citations. In: Proceedings of the 2012 workshop on biomedical natural language processing, p. 91–99. Association for Computational Linguistics; 2012.
Vincze V, Szarvas G, Farkas R, Móra G, Csirik J. The bioscope corpus: biomedical texts annotated for uncertainty, negation and their scopes. BMC Bioinform. 2008;9(11):S9.
Gurulingappa H, Rajput AM, Roberts A, Fluck J, Hofmann-Apitius M, Toldo L. Development of a benchmark corpus to support the automatic extraction of drug-related adverse effects from medical case reports. J Biomed Inform. 2012;45(5):885–92.
Uzuner Ö, Solti I, Cadag E. Extracting medication information from clinical text. J Am Med Inform Assoc. 2010;17(5):514–8.
Henriksson A, Kvist M, Dalianis H, Duneld M. Identifying adverse drug event information in clinical notes with distributional semantic representations of context. J Biomed Inform. 2015;57:333–49.
Fleiss JL. Measuring nominal scale agreement among many raters. Psychol Bull. 1971;76(5):378.
Landis JR, Koch GG. The measurement of observer agreement for categorical data. Biometrics. 1977;33:159–74.
Liu Z, Chen Y, Tang B, Wang X, Chen Q, Li H, Wang J, Deng Q, Zhu S. Automatic de-identification of electronic medical records using token-level and character-level conditional random fields. J Biomed Inform. 2015;58:S47–52. https://doi.org/10.1016/j.jbi.2015.06.009.
Wunnava S, Qin X, Kakar T, Rundensteiner EA, Kong X. Bidirectional LSTM-CRF for adverse drug event tagging in electronic health records. In: Liu F, Jagannatha A, Yu H, editors. Proceedings of the 1st International Workshop on Medication and Adverse Drug Event Detection, Proceedings of Machine Learning Research, vol. 90, p. 48–56. PMLR; 2018. http://proceedings.mlr.press/v90/wunnava18a.html. Accessed 10 May 2018.
Dandala B, Joopudi V, Devarakonda M. Adverse drug events detection in clinical notes by jointly modeling entities and relations using neural networks. Drug Saf. 2019. https://doi.org/10.1007/s40264-018-0764-x.
Yang X, Bian J, Gong Y, Hogan WR, Wu Y. MADEx: a system for detecting medications, adverse drug events, and their relations from clinical notes. Drug Saf. 2019. https://doi.org/10.1007/s40264-018-0761-0.
Xu D, Yadav V, Bethard S. Uarizona at the made 1.0 NLP challenge. In: Liu F, Jagannatha A, Yu H, editors, Proceedings of the 1st international workshop on medication and adverse drug event detection, Proceedings of machine learning research, vol. 90, pp. 57–65. PMLR; 2018. http://proceedings.mlr.press/v90/xu18a.html. Accessed 10 May 2018.
Chapman AB, Peterson KS, Alba PR, DuVall SL, Patterson OV. Detecting adverse drug events with rapidly trained classification models. Drug Saf. 2019. https://doi.org/10.1007/s40264-018-0763-y.
Ngo D-H, Metke-Jimenez A, Nguyen A. Knowledge-based feature engineering for detecting medication and adverse drug events from electronic health records. In: Liu F, Jagannatha A, Yu H, editors, Proceedings of the 1st international workshop on medication and adverse drug event detection, Proceedings of machine learning research, vol. 90, pp. 31–38. PMLR; 2018. http://proceedings.mlr.press/v90/ngo18a.html. Accessed 10 May 2018.
Magge A, Scotch M, Gonzalez-Hernandez G. Clinical NER and relation extraction using bi-char-LSTMs and random forest classifiers. In: Liu F, Jagannatha A, Yu H, editors. Proceedings of the 1st international workshop on medication and adverse drug event detection, Proceedings of machine learning research, vol. 90, p. 25–30. PMLR; 2018. http://proceedings.mlr.press/v90/magge18a.html. Accessed 10 May 2018.
Florez E, Precioso F, Riveill M, Pighetti R. Named entity recognition using neural networks for clinical notes. In: Liu F, Jagannatha A, Yu H, editors. Proceedings of the 1st international workshop on medication and adverse drug event detection, Proceedings of machine learning research, vol. 90, p. 7–15. PMLR; 2018. http://proceedings.mlr.press/v90/florez18a.html. Accessed 10 May 2018.
Lafferty J, McCallum A, Pereira FCN. Conditional random fields: probabilistic models for segmenting and labeling sequence data. In: Proceedings of the 18th international conference on machine learning. San Francisco: Morgan Kaufmann Publishers Inc.; 2001. pp. 282–9.
McCallum A, Freitag D, Pereira FCN. Maximum entropy markov models for information extraction and segmentation. In: ICML, vol. 17, pp. 591–598; 2000.
Zhou GD, Su J. Named entity recognition using an hmm-based chunk tagger. In: Proceedings of the 40th annual meeting on association for computational linguistics, p. 473–480. Association for Computational Linguistics; 2002.
Gers FA, Schmidhuber J, Cummins F. Learning to forget: continual prediction with LSTM. Neural Comput. 2000;12(10):2451–71.
Chung J, Gulcehre C, Cho K, Bengio Y. Gated feedback recurrent neural networks. In: International conference on machine learning, p. 2067–2075; 2015.
Jagannatha AN, Yu H. Bidirectional RNN for medical event detection in electronic health records. In: Proceedings of the conference. Association for Computational Linguistics. North American Chapter. Meeting, vol. 2016, p. 473. NIH Public Access; 2016.
Sundermeyer M, Schlüter R, Ney H. LSTM neural networks for language modeling. In: Thirteenth annual conference of the international speech communication association; 2012.
Huang Z, Xu W, Yu K. Bidirectional LSTM-CRF models for sequence tagging. arXiv preprint, arXiv:1508.01991; 2015.
Cristianini N, Shawe-Taylor J, et al. An introduction to support vector machines and other kernel-based learning methods. Cambridge: Cambridge University Press; 2000.
Breiman Leo. Random forests. Mach Learn. 2001;45(1):5–32.
Acknowledgements
The authors are extremely thankful to the MADE 1.0 annotation team: Elaine Freund, Heather Keating, Nadya Frid, Edgard Granillo, Raelene Goodwin, Brian Corner, Zuofeng Li, Rashmi Prasad, Balaji Ramesh, Victoria Wang, and Steven Belknap for their contributions to the MADE project. They were an essential part of the data curation, annotation, and research process for MADE 1.0. They are also the authors of the annotation guideline used throughout the development of this corpus.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Funding
Research reported in this publication was supported by the National Heart, Lung, and Blood Institute (NHLBI) of the National Institutes of Health under award number R01HL125089.
Declaration
The content is solely the responsibility of the authors and does not necessarily represent the official views of the National Institutes of Health.
Conflict of interest
Abhyuday Jagannatha, Feifan Liu, Weisong Liu, and Hong Yu have no conflicts of interest that are directly relevant to the content of this article.
Dataset
The data used are from the MADE 1.0 corpus available at http://bio-nlp.org/index.php/projects/39-nlp-challenges.
Additional information
Part of a theme issue on “NLP Challenge for Detecting Medication and Adverse Drug Events from Electronic Health Records (MADE 1.0)” guest edited by Feifan Liu, Abhyuday Jagannatha and Hong Yu.
Rights and permissions
About this article
Cite this article
Jagannatha, A., Liu, F., Liu, W. et al. Overview of the First Natural Language Processing Challenge for Extracting Medication, Indication, and Adverse Drug Events from Electronic Health Record Notes (MADE 1.0). Drug Saf 42, 99–111 (2019). https://doi.org/10.1007/s40264-018-0762-z
Published:
Issue Date:
DOI: https://doi.org/10.1007/s40264-018-0762-z