Abstract
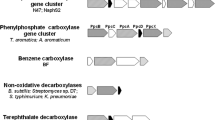
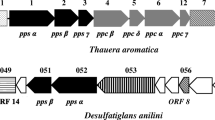
The sulfate-reducing highly enriched culture N47 is capable to anaerobically degrade naphthalene, 2-methylnaphthalene, and 2-naphthoic acid. A proteogenomic investigation was performed to elucidate the initial activation reaction of anaerobic naphthalene degradation. This lead to the identification of an alpha-subunit of a carboxylase protein that was two-fold up-regulated in naphthalene-grown cells compared to 2-methylnaphthalene-grown cells. The putative naphthalene carboxylase subunit showed 48% similarity to the anaerobic benzene carboxylase from an iron-reducing, benzene-degrading culture and 45% to alpha-subunit of phenylphosphate carboxylase of Aromatoleum aromaticum EbN1. A gene for the beta-subunit of putative naphthalene carboxylase was located nearby on the genome and was expressed with naphthalene. Similar to anaerobic benzene carboxylase, there were no genes for gamma- and delta-subunits of a putative carboxylase protein located on the genome which excludes participation in degradation of phenolic compounds. The genes identified for putative naphthalene carboxylase subunits showed only weak similarity to 4-hydroxybenzoate decarboxylase excluding ATP-independent carboxylation. Several ORFs were identified that possibly encode a 2-naphthoate-CoA ligase, which is obligate for activation before the subsequent ring reduction by naphthoyl-CoA reductase. One of these ligases was exclusively expressed on naphthalene and 2-naphthoic acid and might be the responsible naphthoate-CoA-ligase.



Similar content being viewed by others
References
Abu Laban N, Selesi D, Rattei T, Tischler P, Meckenstock RU (2010) Identification of enzymes involved in anaerobic benzene degradation by a strictly anaerobic iron-reducing enrichment culture. Environ Microbiol 12:2783–2796
Altschul SF et al (1997) Gapped BLAST and PSI-BLAST: a new generation of protein database search programs. Nucleic Acids Res 25:3389–3402
Annweiler E et al (2000) Anaerobic degradation of 2-methylnaphthalene by a sulfate-reducing enrichment culture. Appl Environ Microb 66:5329–5333
Bergmann FD, Selesi D, Weinmaier T, Tischler P, Rattei T, Meckenstock RU (2010) Genomic insights into the metabolic potential of the polycyclic aromatic hydrocarbon degrading sulphate-reducing Deltaproteobacterium N47. Environ Microbiol
Boll M, Fuchs G, Heider J (2002) Anaerobic oxidation of aromatic compounds and hydrocarbons. Curr Opin Chem Biol 6:604–611
Cline JD (1969) Spectrophotometric determination of hydrogen sulfide in natural waters. Limnol Oceanogr 14:454–458
Galushko A, Minz D, Schink B, Widdel F (1999) Anaerobic degradation of naphthalene by a pure culture of a novel type of marine sulphate-reducing bacterium. Environ Microbiol 1:415–420
Gibson J, Dispensa M, Fogg GC, Evans DT, Harwood CS (1994) 4-Hydroxybenzoate-coenzyme A ligase from Rhodopseudomonas palustris: purification, gene sequence, and role in anaerobic degradation. J Bacteriol 176:634–641
Habe H, Omori T (2003) Genetics of polycyclic aromatic hydrocarbon metabolism in diverse aerobic bacteria. Biosci Biotech Bioch 67:225–243
He Z, Wiegel J (1995) Purification and characterization of an oxygen-sensitive reversible 4-hydroxybenzoate decarboxylase from Clostridium hydroxybenzoicum. Eur J Biochem 229:77–82
Keller A, Nesvizhskii AI, Kolker E, Aebersold R (2002) Empirical statistical model to estimate the accuracy of peptide identifications made by MS/MS and database search. Anal Chem 74:5383–5392
Leuthner B et al (1998) Biochemical and genetic characterization of benzylsuccinate synthase from Thauera aromatica: a new glycyl radical enzyme catalysing the first step in anaerobic toluene metabolism. Mol Microbiol 28:615–628
Lueders T, Kindler R, Miltner A, Friedrich MW, Kaestner M (2006) Identification of bacterial micropredators distinctively active in a soil microbial food web. Appl Environ Microb 72:5342–5348
Meckenstock RU, Annweiler E, Michaelis W, Richnow HH, Schink B (2000) Anaerobic naphthalene degradation by a sulfate-reducing enrichment culture. Appl Environ Microb 66:2743–2747
Musat F et al (2009) Anaerobic degradation of naphthalene and 2-methylnaphthalene by strains of marine sulfate-reducing bacteria. Environ Microbiol 11:209–219
Muyzer G, Teske A, Wirsen C, Jannasch H (1995) Phylogenetic relationships of Thiomicrospira species and their identification in deep-sea hydrothermal vent samples by denaturing gradient gel electrophoresis of 16S rDNA fragments. Arch Microbiol 164:165–172
Nesvizhskii AI, Keller A, Kolker E, Aebersold R (2003) A statistical model for identifying proteins by tandem mass spectrometry. Anal Chem 75:4646–4658
Rabus R et al (2005) The genome sequence of an anaerobic aromatic-degrading denitrifying bacterium, strain EbN1. Arch Microbiol 183:27–36
Rappsilber J, Siniossoglou S, Hurt EC, Mann M (2000) A generic strategy to analyze the spatial organization of multi-protein complexes by cross-linking and mass spectrometry. Anal Chem 72:267–275
Rockne KJ, Chee-Sanford JC, Sanford RA, Hedlund BP, Staley JT, Strand SE (2000) Anaerobic naphthalene degradation by microbial pure cultures under nitrate-reducing conditions. Appl Environ Microb 66:1595–1601
Safinowski M, Meckenstock RU (2004) Enzymatic reactions in anaerobic 2-methylnaphthalene degradation by the sulphate-reducing enrichment culture N47. FEMS Microbiol Lett 240:99–104
Safinowski M, Meckenstock RU (2006) Methylation is the initial reaction in anaerobic naphthalene degradation by a sulfate-reducing enrichment culture. Environ Microbiol 8:347–352
Schuhle K, Fuchs G (2004) Phenylphosphate carboxylase: a new C-C lyase involved in anaerobic in phenol metabolism in Thauera aromatica. J Bacteriol 186:4556–4567
Selesi D et al (2010) Combined genomic and proteomic approaches identify gene clusters involved in anaerobic 2-methylnaphthalene degradation in the sulfate-reducing enrichment culture N47. J Bacteriol 192:295–306
Ting L, Cowley MJ, Hoon SL, Guilhaus M, Raftery MJ, Cavicchioli R (2009) Normalization and statistical analysis of quantitative proteomics data generated by metabolic labeling. Mol Cell Proteomics 8:2227–2242
Walter MC et al (2009) PEDANT covers all complete RefSeq genomes. Nucleic Acids Res 37:D408–D411
Weisburg WG, Barns SM, Pelletier DA, Lane DJ (1991) 16S ribosomal DNA amplification for phylogenetic study. J Bacteriol 173:697–703
Wu LF, Han DK (2006) Overcoming the dynamic range problem in mass spectrometry-based shotgun proteomics. Expert Rev Proteomic 3:611–619
Zehr BD, Savin TJ, Hall RE (1989) A one-step, low background Coomassie staining procedure for polyacrylamide gels. Anal Biochem 182:157–159
Zhang XM, Young LY (1997) Carboxylation as an initial reaction in the anaerobic metabolism of naphthalene and phenanthrene by sulfidogenic consortia. Appl Environ Microb 63:4759–4764
Author information
Authors and Affiliations
Corresponding author
Additional information
Communicated by Joerg Overmann.
Rights and permissions
About this article
Cite this article
Bergmann, F.D., Selesi, D. & Meckenstock, R.U. Identification of new enzymes potentially involved in anaerobic naphthalene degradation by the sulfate-reducing enrichment culture N47. Arch Microbiol 193, 241–250 (2011). https://doi.org/10.1007/s00203-010-0667-4
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00203-010-0667-4