Abstract
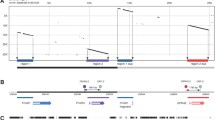
G protein-coupled receptors are the principal mediators of the sweet, umami, bitter, and fat taste qualities in mammals. Intriguingly, the taste receptors are also expressed outside of the oral cavity, including in the gut, airways, brain, and heart, where they have additional functions and contribute to disease. However, there is little known about the mechanisms governing the transcriptional regulation of taste receptor genes. Following our recent delineation of taste receptors in the heart, we investigated the genomic loci encoding for taste receptors to gain insight into the regulatory mechanisms that drive their expression in the heart. Gene expression analyses of healthy and diseased human and mouse hearts showed coordinated expression for a subset of chromosomally clustered taste receptors. This chromosomal clustering mirrored the cardiac expression profile, suggesting that a common gene regulatory block may control the taste receptor locus. We identified unique domains with strong regulatory potential in the vicinity of taste receptor genes. We also performed de novo motif enrichment in the proximal promoter regions and found several overrepresented DNA motifs in cardiac taste receptor gene promoters corresponding to ubiquitous and cardiac-specific transcription factor binding sites. Thus, combining cardiac gene expression data with bioinformatic analyses, this study has provided insights into the noncoding regulatory landscape for taste GPCRs. These findings also have broader relevance for the study of taste GPCRs outside of the classical gustatory system, where understanding the mechanisms controlling the expression of these receptors may have implications for future therapeutic development.










Similar content being viewed by others
Abbreviations
- bHLH:
-
Basic helix-loop-helix
- bZIP:
-
Basic leucine zipper
- ChIP:
-
Chromatin immunoprecipitation
- ENCODE:
-
Encyclopedia of DNA elements
- GPCRs:
-
G protein-coupled receptors
- I/R:
-
Ischemia/reperfusion
- LAD:
-
Left anterior descending
- LV:
-
Left ventricular
- PWM:
-
Position weight matrix
- TAS1R/T1R:
-
Taste receptor type 1
- TAS2R/T2R:
-
Taste receptor type 2
- TFBS:
-
Transcription factor binding sites
References
Adler E, Hoon MA, Mueller KL, Chandrashekar J, Ryba NJP, Zuker CS (2000) A novel family of mammalian taste receptors. Cell 100:693–702
Aurora AB, Mahmoud AI, Luo X, Johnson BA, van Rooij E, Matsuzaki S, Humphries KM, Hill JA, Bassel-Duby R, Sadek HA, Olson EN (2012) MicroRNA-214 protects the mouse heart from ischemic injury by controlling Ca(2)(+) overload and cell death. J Clin Invest 122:1222–1232
Berger MF, Badis G, Gehrke AR, Talukder S, Philippakis AA, Pena-Castillo L, Alleyne TM, Mnaimneh S, Botvinnik OB, Chan ET, Khalid F, Zhang W, Newburger D, Jaeger SA, Morris QD, Bulyk ML, Hughes TR (2008) Variation in homeodomain DNA binding revealed by high-resolution analysis of sequence preferences. Cell 133:1266–1276
Bryne JC, Valen E, Tang MH, Marstrand T, Winther O, da Piedade I, Krogh A, Lenhard B, Sandelin A (2008) JASPAR, the open access database of transcription factor-binding profiles: new content and tools in the 2008 update. Nucleic Acids Res 36:D102–D106
Cartoni C, Yasumatsu K, Ohkuri T, Shigemura N, Yoshida R, Godinot N, le Coutre J, Ninomiya Y, Damak S (2010) Taste preference for fatty acids is mediated by GPR40 and GPR120. J Neurosci 30:8376–8382
Chandrashekar J, Kuhn C, Oka Y, Yarmolinsky DA, Hummler E, Ryba NJP, Zuker CS (2010) The cells and peripheral representation of sodium taste in mice. Nature 464:297–301
Chen MC, Wu SV, Reeve JR, Rozengurt E (2006) Bitter stimuli induce Ca2+ signaling and CCK release in enteroendocrine STC-1 cells: role of L-type voltage-sensitive Ca2+ channels. Am J Physiol Cell Physiol 291:C726–C739
Cogliati T, Good DJ, Haigney M, Delgado-Romero P, Eckhaus MA, Koch WJ, Kirsch IR (2002) Predisposition to arrhythmia and autonomic dysfunction in Nhlh1-deficient mice. Mol Cell Biol 22:4977–4983
Collins FS, Green ED, Guttmacher AE, Guyer MS (2003) A vision for the future of genomics research. Nature 422:835–847
Conway SJ, Firulli B, Firulli AB (2010) A bHLH code for cardiac morphogenesis. Pediatr Cardiol 31:318–324
Damak S, Mosinger B, Margolskee RF (2008) Transsynaptic transport of wheat germ agglutinin expressed in a subset of type II taste cells of transgenic mice. BMC Neurosci 9:96
Dekker J, Rippe K, Dekker M, Kleckner N (2002) Capturing chromosome conformation. Science 295:1306–1311
Dekker J, Marti-Renom MA, Mirny LA (2013) Exploring the three-dimensional organization of genomes: interpreting chromatin interaction data. Nat Rev Genet 14:390–403
Deshpande DA, Wang WCH, McIlmoyle EL, Robinett KS, Schillinger RM, An SS, Sham JSK, Liggett SB (2010) Bitter taste receptors on airway smooth muscle bronchodilate by localized calcium signaling and reverse obstruction. Nat Med 16:1299–1304
Dostie J, Richmond TA, Arnaout RA, Selzer RR, Lee WL, Honan TA, Rubio ED, Krumm A, Lamb J, Nusbaum C, Green RD, Dekker J (2006) Chromosome Conformation Capture Carbon Copy (5C): a massively parallel solution for mapping interactions between genomic elements. Genome Res 16:1299–1309
ENCODE Project Consortium (2004) The ENCODE (ENCyclopedia Of DNA Elements) Project. Science (New York, NY) 306:636–640
ENCODE Project Consortium (2011) A user's guide to the encyclopedia of DNA elements (ENCODE). PLoS Biol 9:e1001046
Ettwiller L, Paten B, Ramialison M, Birney E, Wittbrodt J (2007) Trawler: de novo regulatory motif discovery pipeline for chromatin immunoprecipitation. Nat Methods 4:563–565
Fentzke RC, Korcarz CE, Lang RM, Lin H, Leiden JM (1998) Dilated cardiomyopathy in transgenic mice expressing a dominant-negative CREB transcription factor in the heart. J Clin Invest 101:2415–2426
Flegel C, Manteniotis S, Osthold S, Hatt H, Gisselmann G (2013) Expression profile of ectopic olfactory receptors determined by deep sequencing. PLoS One 8:e55368
Flicek P, Ahmed I, Amode MR, Barrell D, Beal K, Brent S, Carvalho-Silva D, Clapham P, Coates G, Fairley S, Fitzgerald S, Gil L, Garcia-Giron C, Gordon L, Hourlier T, Hunt S, Juettemann T, Kahari AK, Keenan S, Komorowska M, Kulesha E, Longden I, Maurel T, McLaren WM, Muffato M, Nag R, Overduin B, Pignatelli M, Pritchard B, Pritchard E, Riat HS, Ritchie GR, Ruffier M, Schuster M, Sheppard D, Sobral D, Taylor K, Thormann A, Trevanion S, White S, Wilder SP, Aken BL, Birney E, Cunningham F, Dunham I, Harrow J, Herrero J, Hubbard TJ, Johnson N, Kinsella R, Parker A, Spudich G, Yates A, Zadissa A, Searle SM (2013) Ensembl 2013. Nucleic Acids Res 41:D48–D55
Foster SR, Porrello ER, Purdue B, Chan H-W, Voigt A, Frenzel S, Hannan RD, Moritz KM, Simmons DG, Molenaar P, Roura E, Boehm U, Meyerhof W, Thomas WG (2013) Expression, regulation and putative nutrient-sensing function of taste GPCRs in the heart. PLoS One 8:e64579
Foster SR, Blank K, See Hoe LE, Behrens M, Meyerhof W, Peart JN, Thomas WG (2014a) Bitter taste receptor agonists elicit G-protein-dependent negative inotropy in the murine heart. The FASEB J
Foster SR, Roura E, Thomas WG (2014b) Extrasensory perception: Odorant and taste receptors beyond the nose and mouth. Pharmacol Ther 142:41–61
Galindo MM, Voigt N, Stein J, van Lengerich J, Raguse JD, Hofmann T, Meyerhof W, Behrens M (2012) G protein-coupled receptors in human fat taste perception. Chem Senses 37:123–139
Haudry Y, Ramialison M, Paten B, Wittbrodt J, Ettwiller L (2010) Using Trawler_standalone to discover overrepresented motifs in DNA and RNA sequences derived from various experiments including chromatin immunoprecipitation. Nat Protoc 5:323–334
Huang AL, Chen X, Hoon MA, Chandrashekar J, Guo W, Trankner D, Ryba NJP, Zuker CS (2006) The cells and logic for mammalian sour taste detection. Nature 442:934–938
John S, Sabo PJ, Canfield TK, Lee K, Vong S, Weaver M, Wang H, Vierstra J, Reynolds AP, Thurman RE, Stamatoyannopoulos JA (2013) Genome-scale mapping of DNase I hypersensitivity. Curr Protoc Mol Biol Chapter 27: Unit 21 27
Karolchik D, Hinrichs AS, Kent WJ (2012) The UCSC Genome Browser. Curr Protoc Bioinformatics Chapter 1: Unit1 4
Kaul A, Koster M, Neuhaus H, Braun T (2000) Myf-5 revisited: loss of early myotome formation does not lead to a rib phenotype in homozygous Myf-5 mutant mice. Cell 102:17–19
Knuppel R, Dietze P, Lehnberg W, Frech K, Wingender E (1994) TRANSFAC retrieval program: a network model database of eukaryotic transcription regulating sequences and proteins. J Comput Biol 1:191–198
Lander ES, Linton LM, Birren B, Nusbaum C, Zody MC, Baldwin J, Devon K, Dewar K, Doyle M, FitzHugh W, Funke R, Gage D, Harris K, Heaford A, Howland J, Kann L, Lehoczky J, LeVine R, McEwan P, McKernan K, Meldrim J, Mesirov JP, Miranda C, Morris W, Naylor J, Raymond C, Rosetti M, Santos R, Sheridan A, Sougnez C, Stange-Thomann N, Stojanovic N, Subramanian A, Wyman D, Rogers J, Sulston J, Ainscough R, Beck S, Bentley D, Burton J, Clee C, Carter N, Coulson A, Deadman R, Deloukas P, Dunham A, Dunham I, Durbin R, French L, Grafham D, Gregory S, Hubbard T, Humphray S, Hunt A, Jones M, Lloyd C, McMurray A, Matthews L, Mercer S, Milne S, Mullikin JC, Mungall A, Plumb R, Ross M, Shownkeen R, Sims S, Waterston RH, Wilson RK, Hillier LW, McPherson JD, Marra MA, Mardis ER, Fulton LA, Chinwalla AT, Pepin KH, Gish WR, Chissoe SL, Wendl MC, Delehaunty KD, Miner TL, Delehaunty A, Kramer JB, Cook LL, Fulton RS, Johnson DL, Minx PJ, Clifton SW, Hawkins T, Branscomb E, Predki P, Richardson P, Wenning S, Slezak T, Doggett N, Cheng JF, Olsen A, Lucas S, Elkin C, Uberbacher E, Frazier M, Gibbs RA, Muzny DM, Scherer SE, Bouck JB, Sodergren EJ, Worley KC, Rives CM, Gorrell JH, Metzker ML, Naylor SL, Kucherlapati RS, Nelson DL, Weinstock GM, Sakaki Y, Fujiyama A, Hattori M, Yada T, Toyoda A, Itoh T, Kawagoe C, Watanabe H, Totoki Y, Taylor T, Weissenbach J, Heilig R, Saurin W, Artiguenave F, Brottier P, Bruls T, Pelletier E, Robert C, Wincker P, Rosenthal A, Platzer M, Nyakatura G, Taudien S, Rump A, Yang HM, Yu J, Wang J, Huang GY, Gu J, Hood L, Rowen L, Madan A, Qin SZ, Davis RW, Federspiel NA, Abola AP, Proctor MJ, Myers RM, Schmutz J, Dickson M, Grimwood J, Cox DR, Olson MV, Kaul R, Shimizu N, Kawasaki K, Minoshima S, Evans GA, Athanasiou M, Schultz R, Roe BA, Chen F, Pan HQ, Ramser J, Lehrach H, Reinhardt R, McCombie WR, de la Bastide M, Dedhia N, Blocker H, Hornischer K, Nordsiek G, Agarwala R, Aravind L, Bailey JA, Bateman A, Batzoglou S, Birney E, Bork P, Brown DG, Burge CB, Cerutti L, Chen HC, Church D, Clamp M, Copley RR, Doerks T, Eddy SR, Eichler EE, Furey TS, Galagan J, Gilbert JGR, Harmon C, Hayashizaki Y, Haussler D, Hermjakob H, Hokamp K, Jang WH, Johnson LS, Jones TA, Kasif S, Kaspryzk A, Kennedy S, Kent WJ, Kitts P, Koonin EV, Korf I, Kulp D, Lancet D, Lowe TM, McLysaght A, Mikkelsen T, Moran JV, Mulder N, Pollara VJ, Ponting CP, Schuler G, Schultz JR, Slater G, Smit AFA, Stupka E, Szustakowki J, Thierry-Mieg D, Thierry-Mieg J, Wagner L, Wallis J, Wheeler R, Williams A, Wolf YI, Wolfe KH, Yang SP, Yeh RF, Collins F, Guyer MS, Peterson J, Felsenfeld A, Wetterstrand KA, Patrinos A, Morgan MJ (2001) Initial sequencing and analysis of the human genome. Nature 409:860–921
Lee RJ, Xiong G, Kofonow JM, Chen B, Lysenko A, Jiang P, Abraham V, Doghramji L, Adappa ND, Palmer JN, Kennedy DW, Beauchamp GK, Doulias PT, Ischiropoulos H, Kreindler JL, Reed DR, Cohen NA (2012) T2R38 taste receptor polymorphisms underlie susceptibility to upper respiratory infection. J Clin Invest 122:4145–4159
Lelli KM, Slattery M, Mann RS (2012) Disentangling the many layers of eukaryotic transcriptional regulation. Annu Rev Genet 46:43–68
Li Q, Peterson KR, Fang X, Stamatoyannopoulos G (2002) Locus control regions. Blood 100:3077–3086
Liman ER, Zhang YV, Montell C (2014) Peripheral coding of taste. Neuron 81:984–1000
Livak KJ, Schmittgen TD (2001) Analysis of relative gene expression data using real-time quantitative PCR and the 2(T)(-Delta Delta C) method. Methods 25:402–408
Mahony S, Benos PV (2007) STAMP: a web tool for exploring DNA-binding motif similarities. Nucleic Acids Res 35:W253–W258
Maston GA, Evans SK, Green MR (2006) Transcriptional regulatory elements in the human genome. Annu Rev Genom Hum G 7:29–59
Meyerhof W, Batram C, Kuhn C, Brockhoff A, Chudoba E, Bufe B, Appendino G, Behrens M (2010) The Molecular Receptive Ranges of Human TAS2R Bitter Taste Receptors. Chem Senses 35:157–170
Millar RP, Newton CL (2010) The year in G protein-coupled receptor research. Mol Endocrinol 24:261–274
Molkentin JD, Lin Q, Duncan SA, Olson EN (1997) Requirement of the transcription factor GATA4 for heart tube formation and ventral morphogenesis. Genes Dev 11:1061–1072
Molkentin JD, Lu JR, Antos CL, Markham B, Richardson J, Robbins J, Grant SR, Olson EN (1998) A calcineurin-dependent transcriptional pathway for cardiac hypertrophy. Cell 93:215–228
Mollova M, Bersell K, Walsh S, Savla J, Das LT, Park SY, Silberstein LE, dos Remedios CG, Graham D, Colan S, Kuhn B (2013) Cardiomyocyte proliferation contributes to heart growth in young humans. Proc Natl Acad Sci U S A 110:1446–1451
Mueller KL, Hoon MA, Erlenbach I, Chandrashekar J, Zuker CS, Ryba NJP (2005) The receptors and coding logic for bitter taste. Nature 434:225–229
Naya FJ, Black BL, Wu H, Bassel-Duby R, Richardson JA, Hill JA, Olson EN (2002) Mitochondrial deficiency and cardiac sudden death in mice lacking the MEF2A transcription factor. Nat Med 8:1303–1309
Nei M, Niimura Y, Nozawa M (2008) The evolution of animal chemosensory receptor gene repertoires: roles of chance and necessity. Nat Rev Genet 9:951–963
Nelson G, Hoon MA, Chandrashekar J, Zhang YF, Ryba NJP, Zuker CS (2001) Mammalian sweet taste receptors. Cell 106:381–390
Nelson G, Chandrashekar J, Hoon MA, Feng LX, Zhao G, Ryba NJP, Zuker CS (2002) An amino-acid taste receptor. Nature 416:199–202
Niu Z, Yu W, Zhang SX, Barron M, Belaguli NS, Schneider MD, Parmacek M, Nordheim A, Schwartz RJ (2005) Conditional mutagenesis of the murine serum response factor gene blocks cardiogenesis and the transcription of downstream gene targets. J Biol Chem 280:32531–32538
Ohmoto M, Matsumoto I, Yasuoka A, Yoshihara Y, Abe K (2008) Genetic tracing of the gustatory and trigeminal neural pathways originating from T1R3-expressing taste receptor cells and solitary chemoreceptor cells. Mol Cell Neurosci 38:505–517
Ohmoto M, Maeda N, Abe K, Yoshihara Y, Matsumoto I (2010) Genetic tracing of the neural pathway for bitter taste in t2r5-WGA transgenic mice. Biochem Biophys Res Commun 400:734–738
Oka Y, Butnaru M, von Buchholtz L, Ryba NJP, Zuker CS (2013) High salt recruits aversive taste pathways. Nature 494:472–475
Rajkumar P, Aisenberg WH, Acres OW, Protzko RJ, Pluznick JL (2014) Identification and characterization of novel renal sensory receptors. PLoS One 9:e111053
Rosenbloom KR, Dreszer TR, Long JC, Malladi VS, Sloan CA, Raney BJ, Cline MS, Karolchik D, Barber GP, Clawson H, Diekhans M, Fujita PA, Goldman M, Gravell RC, Harte RA, Hinrichs AS, Kirkup VM, Kuhn RM, Learned K, Maddren M, Meyer LR, Pohl A, Rhead B, Wong MC, Zweig AS, Haussler D, Kent WJ (2012) ENCODE whole-genome data in the UCSC Genome Browser: update 2012. Nucleic Acids Res 40:D912–D917
Sawzdargo M, George SR, Nguyen T, Xu S, Kolakowski LF, O'Dowd BF (1997) A cluster of four novel human G protein-coupled receptor genes occurring in close proximity to CD22 gene on chromosome 19q13.1. Biochem Biophys Res Commun 239:543–547
Schauer IE, Knaub LA, Lloyd M, Watson PA, Gliwa C, Lewis KE, Chait A, Klemm DJ, Gunter JM, Bouchard R, McDonald TO, O'Brien KD, Reusch JE (2010) CREB downregulation in vascular disease: a common response to cardiovascular risk. Arterioscler Thromb Vasc Biol 30:733–741
Schulte JS, Seidl MD, Nunes F, Freese C, Schneider M, Schmitz W, Muller FU (2012) CREB critically regulates action potential shape and duration in the adult mouse ventricle. Am J Physiol Heart Circ Physiol 302:H1998–H2007
Smith CM, Finger JH, Hayamizu TF, McCright IJ, Xu J, Berghout J, Campbell J, Corbani LE, Forthofer KL, Frost PJ, Miers D, Shaw DR, Stone KR, Eppig JT, Kadin JA, Richardson JE, Ringwald M (2014) The mouse Gene Expression Database (GXD): 2014 update. Nucleic Acids Res 42:D818–D824
Spitz F, Duboule D (2008) Global control regions and regulatory landscapes in vertebrate development and evolution. Adv Genet 61:175–205
Tizzano M, Cristofoletti M, Sbarbati A, Finger T (2011) Expression of taste receptors in Solitary Chemosensory Cells of rodent airways. BMC Pulm Med 11:3
Toyono T, Seta Y, Kataoka S, Toyoshima K (2007) CCAAT/Enhancer-binding protein beta regulates expression of human T1R3 taste receptor gene in the bile duct carcinoma cell line, HuCCT1. Biochim Biophys Acta-Gene Struct Expr 1769:641–648
Vaquerizas JM, Kummerfeld SK, Teichmann SA, Luscombe NM (2009) A census of human transcription factors: function, expression and evolution. Nat Rev Genet 10:252–263
Acknowledgments
We gratefully acknowledge Dr Eric Olson (UT Southwestern Medical Center, Dallas, USA) for provision of mouse heart samples from experimental models of myocardial infarction and cardiac hypertrophy/heart failure. This work was supported by project grants awarded to WGT from the Australian National Health and Medical Research Council (NHMRC) (1024726) and the National Heart Foundation (NHF) of Australia (G-12B-6532). MR was supported by a Career Development Fellowship from the NHMRC and the NHF (1049980). SRF was supported by an Australian Postgraduate Award scholarship.
Authors’ contributions
SRF conceived of the study, participated in its design, performed and analyzed gene expression studies, and drafted the manuscript. EP participated in gene expression studies in mouse heart tissues. MS assisted with human heart sample preparation. NJS provided reagents and samples for human gene expression analyses. PM and CGR provided human myocardial tissues. WGT participated in the study design and coordination and helped to draft the manuscript. MR conceived of the study, and participated in its design and coordination and helped to draft the manuscript. All authors read and approved the final manuscript.
Author information
Authors and Affiliations
Corresponding authors
Electronic supplementary material
Below is the link to the electronic supplementary material.
ESM 1
(DOCX 1281 kb)
Rights and permissions
About this article
Cite this article
Foster, S.R., Porrello, E.R., Stefani, M. et al. Cardiac gene expression data and in silico analysis provide novel insights into human and mouse taste receptor gene regulation. Naunyn-Schmiedeberg's Arch Pharmacol 388, 1009–1027 (2015). https://doi.org/10.1007/s00210-015-1118-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00210-015-1118-1