Abstract
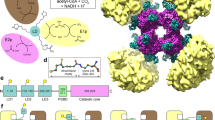
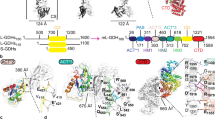
Pyruvate kinase catalyses the final step of the glycolytic pathway in central energy metabolism. The monomeric structure comprises three domains: a catalytic TIM-barrel, a regulatory domain involved in allosteric activation, and a lid domain that encloses the substrates. The lid domain is thought to close over the TIM-barrel domain forming contacts with the substrates to promote catalysis and may be involved in stabilising the activated state when the allosteric activator is bound. However, it remains unknown whether the lid domain is essential for pyruvate kinase catalytic or regulatory function. To address this, we removed the lid domain of Escherichia coli pyruvate kinase type 1 (PKTIM+Reg) using protein engineering. Biochemical analyses demonstrate that, despite the absence of key catalytic residues in the lid domain, PKTIM+Reg retains a low level of catalytic activity and has a reduced binding affinity for the substrate phosphoenolpyruvate. The enzyme retains allosteric activation, but the regulatory profile of the enzyme is changed relative to the wild-type enzyme. Analytical ultracentrifugation and small-angle X-ray scattering data show that, beyond the loss of the lid domain, the PKTIM+Reg structure is not significantly altered and is consistent with the wild-type tetramer that is assembled through interactions at the TIM and regulatory domains. Our results highlight the contribution of the lid domain for facilitating pyruvate kinase catalysis and regulation, which could aid in the development of small molecule inhibitors for pyruvate kinase and related lid-regulated enzymes.




Similar content being viewed by others
Data availability
On request.
References
Abraham MJ, Murtola T, Schulz R, Páll S, Smith JC, Hess B, Lindahl E (2015) GROMACS: high performance molecular simulations through multi-level parallelism from laptops to supercomputers. SoftwareX 1:19–25
Ainscow EK, Brand MD (1999) Top-down control analysis of ATP turnover, glycolysis and oxidative phosphorylation in rat hepatocytes. Eur J Biochem 263:671–685
Arnold K, Bordoli L, Kopp J, Schwede T (2006) The SWISS-MODEL workspace: a web-based environment for protein structure homology modelling. Bioinformatics 22:195–201
Donovan KA, Atkinson SC, Kessans SA, Peng F, Cooper TF, Griffin MD, Jameson GB, Dobson RC (2016a) Grappling with anisotropic data, pseudo-merohedral twinning and pseudo-translational noncrystallographic symmetry: a case study involving pyruvate kinase. Acta Crystallogr D Struct Biol 72:512–519
Donovan KA, Zhu S, Liuni P, Peng F, Kessans SA, Wilson DJ, Dobson RCJ (2016b) Conformational dynamics and allostery in pyruvate kinase. J Biol Chem 291:9244–9256
Emsley P, Lohkamp B, Scott WG, Cowtan K (2010) Features and development of Coot. Acta Crystallogr D Struct Biol 66:486–501
Feksa LR, Cornelio A, Dutra-Filho CS, de Souza Wyse AT, Wajner M, Wannmacher CMD (2004) Inhibition of pyruvate kinase activity by cystine in brain cortex of rats. Brain Res 1012:93–100
Fenton AW, Reinhart GD (2002) Isolation of a single activating allosteric interaction in phosphofructokinase from Escherichia coli. Biochemistry 41:13410–13416
Fleming PJ, Fleming KG (2018) HullRad: fast calculations of folded and disordered protein and nucleic acid hydrodynamic properties. Biophys J 114:856–869
Franke D, Svergun DI (2009) DAMMIF, a program for rapid ab-initio shape determination in small-angle scattering. J Appl Crystallogr 42:342–346
Franke D, Petoukhov M, Konarev P, Panjkovich A, Tuukkanen A, Mertens H, Kikhney A, Hajizadeh N, Franklin J, Jeffries C (2017) ATSAS 2.8: a comprehensive data analysis suite for small-angle scattering from macromolecular solutions. J Appl Crystallogr 50:1212–1225
Johnsen U, Hansen T, Schönheit P (2003) Comparative analysis of pyruvate kinases from the hyperthermophilicarchaeaArchaeoglobus fulgidus, Aeropyrum pernix, and Pyrobaculum aerophilum and the hyperthermophilic bacterium Thermotoga maritima. Unusual regulatory properties in hyperthermophilicarchaea. J Biol Chem 278:25417–25427
Kornberg HL, Malcovati M (1973) Control in situ of the pyruvate kinase activity of Escherichia coli. FEBS Lett 32(2):257–259
Kozin MB, Svergun DI (2001) Automated matching of high-and low-resolution structural models. J Appl Crystallogr 34:33–41
Kumar S, Tsai C-J, Nussinov R (2000) Factors enhancing protein thermostability. Protein Eng, Des Sel 13:179–191
Laue TM, Shah BD, Ridgeway TM, Pelletier SL (1992) Computer-aided interpretation of analytical sedimentation data for proteins. In: Harding S, Rowe A, Horton J (eds) Analytical ultracentrifugation in biochemistry and polymer science. Royal Society of Chemistry, Cambridge, pp 90–125
Laue TM, Stafford WF III (1999) Modern applications of analytical ultracentrifugation. Annu Rev Biophys Biomol Struct 28(1):75–100
Li F, Yu T, Zhao Y, Yu S (2012) Probing the catalytic allosteric mechanism of rabbit muscle pyruvate kinase by tryptophan fluorescence quenching. EurBiophys J 41:607–614
Mattevi A, Valentini G, Rizzi M, Speranza ML, Bolognesi M, Coda A (1995) Crystal structure of Escherichia coli pyruvate kinase type I: molecular basis of the allosteric transition. Structure 3:729–741
Mattevi A, Bolognesi M, Valentini G (1996) The allosteric regulation of pyruvate kinase. FEBS Lett 389:15–19
Mertens HD, Svergun DI (2010) Structural characterization of proteins and complexes using small-angle X-ray solution scattering. J Struct Biol 172:128–141
Morgan HP, McNae IW, Nowicki MW, Hannaert V, Michels PAM, Fothergill-Gilmore LA, Walkinshaw MD (2010) Allosteric mechanism of pyruvate kinase from Leishmania mexicana uses a rock and lock model. J Biol Chem 285:12892–12898
Nagano N, Orengo CA, Thornton JM (2002) One fold with many functions: the evolutionary relationships between TIM barrel families based on their sequences, structures and functions. J Mol Biol 321:741–765
Naithani A, Taylor P, Erman B, Walkinshaw MD (2015) A molecular dynamics study of allosteric transitions in Leishmania mexicana pyruvate kinase. Biophys J 109:1149–1156
Niesen FH, Berglund H, Vedadi M (2007) The use of differential scanning fluorimetry to detect ligand interactions that promote protein stability. Nat Protoc 2:2212
Oria-Hernández J, Cabrera N, Pérez-Montfort R, Ramírez-Silva L (2005) Pyruvate kinase revisited: The activating effect of K+. J Biol Chem 280:37924–37929
Oria-Hernández J, Riveros-Rosas H, Ramírez-Sílva L (2006) Dichotomic phylogenetic tree of the pyruvate kinase family: K+-dependent and -independent enzymes. J Biol Chem 281:30717–30724
Ortega A, Amorós D, García de la Torre J (2011) Prediction of hydrodynamic and other solution properties of rigid proteins from atomic- and residue-level models. Biophys J 101:892–898
Pearce AK, Crimmins K, Groussac E, Hewlins MJ, Dickinson JR, Francois J, Booth IR, Brown AJ (2001) Pyruvate kinase (Pyk1) levels influence both the rate and direction of carbon flux in yeast under fermentative conditions. Microbiology 147:391–401
Pendergrass DC, Williams R, Blair JB, Fenton AW (2006) Mining for allosteric information: natural mutations and positional sequence conservation in pyruvate kinase. IUBMB Life 58:31–38
Petoukhov MV, Svergun DI (2003) New methods for domain structure determination of proteins from solution scattering data. J Appl Crystallogr 36:540–544
Pizzuto R, Paventi G, Atlante A, Passarella S (2010) Pyruvate kinase in pig liver mitochondria. Arch Biochem Biophys 495:42–48
Reeves RE, Sols A (1973) Regulation of Escherichia coli phosphofructokinase in situ. Biochem Biophys Res Commun 50:459–466
Saito T, Nishi M, Lim MI, Wu B, Maeda T, Hashimoto H, Takeuchi T, Roos DS, Asai T (2008) A novel GDP-dependent pyruvate kinase isozyme from Toxoplasma gondii localizes to both the apicoplast and the mitochondrion. J Biol Chem 283:14041–14052
Schormann N, Hayden KL, Lee P, Banerjee S, Chattopadhyay D (2019) An overview of structure, function, and regulation of pyruvate kinases. Protein Sci 28:1771–1784
Schuck P (2000) Size-distribution analysis of macromolecules by sedimentation velocity ultracentrifugation and lamm equation modeling. Biophys J 78:1606–1619
Schuck P, Perugini MA, Gonzales NR, Howlett GJ, Schubert D (2002) Size-distribution analysis of proteins by analytical ultracentrifugation: strategies and application to model systems. Biophys J 82:1096–1111
Sterner R, Höcker B (2005) Catalytic versatility, stability, and evolution of the (βα) 8-barrel enzyme fold. Chem Rev 105:4038–4055
Susan-Resiga D, Nowak T (2004) Proton donor in yeast pyruvate kinase: chemical and kinetic properties of the active site Thr 298 to Cys mutant. Biochemistry 43:15230–15245
Svergun D (1992) Determination of the regularization parameter in indirect-transform methods using perceptual criteria. J Appl Crystallogr 25:495–503
Svergun D, Barberato C, Koch MH (1995) CRYSOL–a program to evaluate X-ray solution scattering of biological macromolecules from atomic coordinates. J Appl Crystallogr 28:768–773
Trewhella J, Duff AP, Durand D, Gabel F, Guss JM, Hendrickson WA, Hura GL, Jacques DA, Kirby NM, Kwan AH (2017) 2017 publication guidelines for structural modelling of small-angle scattering data from biomolecules in solution: an update. Acta Crystallogr D Struct Biol 73:710–728
Valentini G, Chiarelli L, Fortin R, Speranza ML, Galizzi A, Mattevi A (2000) The allosteric regulation of pyruvate kinase A site-directed mutagenesis study. J Biol Chem 275:18145–18152
Valentini G, Chiarelli LR, Fortin R, Dolzan M, Galizzi A, Abraham DJ, Wang C, Bianchi P, Zanella A, Mattevi A (2002) Structure and function of human erythrocyte pyruvate kinase. Molecular basis of nonspherocytic hemolytic anemia. J Biol Chem 277:23807–23814
Van Schaftingen E (1993) Glycolysis revisited. Diabetologia 36:581–588
van Wijk R, Huizinga EG, van Wesel AC, van OirschotHadders BAMA, van Solinge WW (2009) Fifteen novel mutations in PKLR associated with pyruvate kinase (PK) deficiency: structural implications of amino acid substitutions in PK. Hum Mutat 30:446–453
Vistica J, Dam J, Balbo A, Yikilmaz E, Mariuzza RA, Rouault TA, Schuck P (2004) Sedimentation equilibrium analysis of protein interactions with global implicit mass conservation constraints and systematic noise decomposition. Anal Biochem 326:234–256
Volkov VV, Svergun DI (2003) Uniqueness of ab initio shape determination in small-angle scattering. J Appl Crystallogr 36:860–864
Xie L, Wang DI (1996) Energy metabolism and ATP balance in animal cell cultivation using a stoichiometrically based reaction network. Biotechnol Bioeng 52:591–601
Zhu T, Bailey MF, Angley LM, Cooper TF, Dobson RC (2010) The quaternary structure of pyruvate kinase type 1 from Escherichia coli at low nanomolar concentrations. Biochimie 92:116–120
Acknowledgements
This research was undertaken in part using the SAXS/WAXS beamline at the Australian Synchrotron, part of ANSTO, and we thank the staff of SAXS/WAXS beamlines for their assistance in data collection. DC acknowledges the University of Canterbury Doctoral Scholarship for funding support. We acknowledge Sarah Atkinson for help during project setup and Jackie Healy for technical support.
Funding
This work is supported by the New Zealand Royal Society Marsden Fund (contract UOC1013) and the US Army Research Office under contract/grant number W911NF-11–1-0481.
Author information
Authors and Affiliations
Contributions
ES, KAD, and RCJD conceived the project. ES conducted the experiments. ES, DW, and DC analysed and interpreted the data. RCJD secured funding for the work. ES, DW, and DC drafted the manuscript. All authors read and edited the manuscript.
Corresponding author
Ethics declarations
Conflicts of interest
The authors declare that they have no conflict of interest to declare.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Special Issue: Analytical Ultracentrifugation 2019
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Sugrue, E., Coombes, D., Wood, D. et al. The lid domain is important, but not essential, for catalysis of Escherichia coli pyruvate kinase. Eur Biophys J 49, 761–772 (2020). https://doi.org/10.1007/s00249-020-01466-5
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00249-020-01466-5