Abstract
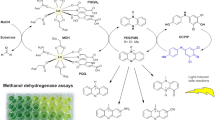
Previous work showed that random mutagenesis produced a mutant of toluene ortho-monooxygenase (TOM) of Burkholderia cepacia G4 containing the V106A substitution in the hydroxylase α-subunit (TomA3) that changed the color of the cell suspension from wild-type brown to green in rich medium. Here, DNA shuffling was used to isolate a random TOM mutant that turned blue due to mutation TomA3 A113V. To better understand the TOM reaction mechanism, we studied the specificity of indole hydroxylation using a spectrum of colored TOM mutants expressed in Escherichia coli TG1 and formed as a result of saturation mutagenesis at TomA3 positions A113 and V106. Colonies expressing these altered enzymes ranged in color from blue through green and purple to orange; and the enzyme products were identified using thin-layer chromatography, high performance liquid chromatography, and liquid chromatography–mass spectroscopy. Derived from the single TOM template, enzymes were identified that produced primarily isoindigo (wild-type TOM), indigo (A113V), indirubin (A113I), and isatin (A113H and V106A/A113G). The discovery that wild-type TOM formed isoindigo via C-2 hydroxylation of the indole pyrrole ring makes this the first oxygenase shown to form this compound. Variant TOM A113G was unable to form indigo, indirubin, or isoindigo (did not hydroxylate the indole pyrrole ring), but produced 4-hydroxyindole and unknown yellow compounds from C-4 hydroxylation of the indole benzene ring. Mutations at V106 in addition to A113G restored C-3 indole oxidation, so along with C-2 indole oxidation, isatin, indigo, and indirubin were formed. Other TomA3 V106/A113 mutants with hydrophobic, polar, or charged amino acids in place of the Val and/or Ala residues hydroxylated indole at the C-3 and C-2 positions, forming isatin, indigo, and indirubin in a variety of distributions. Hence, for the first time, a single enzyme was genetically modified to produce a wide range of colors from indole.


Similar content being viewed by others
References
Adachi J, Mori Y, Matsui S, Takigami H, Fujino J, Kitagawa H, Miller III CA, Kato T, Saeki K, Matsuda T (2001) Indirubin and indigo are potent aryl hydrocarbon receptor ligands present in human urine. J Biol Chem 276:31475–31478
Berry A, Dodge TC, Pepsin M, Weyler W (2002) Application of metabolic engineering to improve both the production and use of biotech indigo. J Ind Microbiol Biotechnol 28:127–133
Bhushan B, Samanta SK, Jain RK (2000) Indigo production by naphthalene-degrading bacteria. Lett Appl Microbiol 31:5–9
Bialy H (1997) Biotechnology, bioremediation, and blue genes. Nat Biotechnol 15:110
Brannigan JA, Wilkinson AJ (2002) Timeline: protein engineering 20 years on. Nat Rev Mol Cell Biol 3:964–970
Buolamwini JK (2000) Cell cycle molecular targets in novel anticancer drug discovery. Curr Pharm Des 6:379–392
Canada KA, Iwashita S, Shim H, Wood TK (2002) Directed evolution of toluene ortho-monooxygenase for enhanced 1-naphthol synthesis and chlorinated ethene degradation. J Bacteriol 184:344–349
Damani LA, Crooks PA (1982) Oxidative metabolism of heterocyclic ring systems. In: Jakoby WB, Bend JR, Caldwell J (eds) Metabolic basis of detoxication. Academic, New York, pp 69–89
Eaton RW, Chapman PJ (1995) Formation of indigo and related compounds from indolecarboxylic acids by aromatic acid-degrading bacteria: chromogenic reactions for cloning genes encoding dioxygenases that act on aromatic acids. J Bacteriol 177:6983–6988
Ensley BD, Ratzkin BJ, Osslund TD, Simon MJ, Wackett LP, Gibson DT (1983) Expression of naphthalene oxidation genes in Escherichia coli results in the biosynthesis of indigo. Science 222:167–169
Frost JW, Lievense J (1994) Prospects for biocatalytic synthesis of aromatics in the 21st century. New J Chem 18:341–348
Gillam EMJ, Notley LM, Cai H, De Voss JJ, Guengerich FP (2000) Oxidation of indole by cytochrome P450 enzymes. Biochemistry 39:13817–13824
Glover V, Halket JM, Watkins PJ, Clow A, Goodwin BL, Sandler M (1988) Isatin: identity with the purified endogenous monoamine oxidase inhibitor tribulin. J Neurochem 51:656–659
Hoessel R, Leclerc S, Endicott JA, Noble MEM, Lawrie A, Tunnah P, Leost M, Damiens E, Marie D, Marko D, Niederberger E, Tang W, Eisenbrand G, Meijer L (1999) Indirubin, the active constituent of a Chinese antileukaemia medicine, inhibits cyclin-dependent kinases. Nat Cell Biol 1:60–67
Luu PP, Yung CW, Sun AK, Wood TK (1995) Monitoring trichloroethylene mineralization by Pseudomonas cepacia G4 PR1. Appl Microbiol Biotechnol 44:259–264
Maugard T, Enaud E, Choisy P, Legoy MD (2001) Identification of an indigo precursor from leaves of Isatis tinctoria (Woad). Phytochemistry 58:897–904
Maugard T, Enaud E, de La Sayette A, Choisy P, Legoy MD (2002) Beta-glucosidase-catalyzed hydrolysis of indican from leaves of Polygonum tinctorium. Biotechnol Prog 18:1104–1108
Mermod N, Harayama S, Timmis KN (1986) New route to bacterial production of indigo. Bio/Technology 4:321–324
Meyer A, Würsten M, Schmid A, Kohler H-PE, Witholt B (2002) Hydroxylation of indole by laboratory-evolved 2-hydroxybiphenyl 3-monooxygenase. J Biol Chem 277:34161–34167
Murdock D, Ensley BD, Serdar C, Thalen M (1993) Construction of metabolic operons catalyzing the de novo biosynthesis of indigo in Escherichia coli. Bio/Technology 11:381–386
Nelson MJK, Montgomery SO, O’Neill EJ, Pritchard PH (1986) Aerobic metabolism of trichloroethylene by a bacterial isolate. Appl Environ Microbiol 52:383–384
Nelson MJK, Montgomery SO, Mahaffey WR, Pritchard PH (1987) Biodegradation of trichloroethylene and involvement of an aromatic biodegradative pathway. Appl Environ Microbiol 53:949–954
Newman LM, Wackett LP (1995) Purification and characterization of toluene 2-monooxygenase from Burkholderia cepacia G4. Biochemistry 34:14066–14076
O’Connor KE, Hartmans S (1998) Indigo formation by aromatic hydrocarbon-degrading bacteria. Biotechnol Lett 20:219–223
O’Connor KE, Dobson AD, Hartmans S (1997) Indigo formation by microorganisms expressing styrene monooxygenase activity. Appl Environ Microbiol 63:4287–4291
Rui L, Kwon YM, Fishman A, Reardon KF, Wood TK (2004) Saturation mutagenesis of toluene ortho-monooxygenase for enhanced 1-naphthol synthesis and chloroform degradation. Appl Environ Microbiol 70:3246–3252
Sakamoto T, Joern JM, Arisawa A, Arnold FH (2001) Laboratory evolution of toluene dioxygenase to accept 4-picoline as a substrate. Appl Environ Microbiol 67:3882–3887
Sambrook J, Fritsch EF, Maniatis T (1989) Molecular cloning: a laboratory manual, 2nd edn. Cold Spring Harbor Laboratory, Cold Spring Harbor, N.Y.
Shields MS, Montgomery SO, Chapman PJ, Cuskey SM, Pritchard PH (1989) Novel pathway of toluene catabolism in the trichloroethylene-degrading bacterium G4. Appl Environ Microbiol 55:1624–1629
Shim H, Wood TK (2000) Aerobic degradation of mixtures of chlorinated aliphatics by cloned toluene-o-xylene monooxygenase and toluene o-monooxygenase in resting cells. Biotechnol Bioeng 70:693–698
Stemmer WPC (1994) DNA shuffling by random fragmentation and reassembly: in vitro recombination for molecular evolution. Proc Natl Acad Sci USA 91:10747–10751
Sundberg RJ (1970) The chemistry of indoles. Academic, New York
Sundberg RJ (1996) Indoles. Academic, San Diego
Tao Y, Fishman A, Bentley WE, Wood TK (2004a) Altering toluene 4-monooxygenase by active site engineering for the synthesis of 3-methoxycatechol, methoxyhydroquinone, and methylhydroquinone. J Bacteriol 186:4705–4713
Tao Y, Fishman A, Bentley WE, Wood TK (2004b) Oxidation of benzene to phenol, catechol, and 1,2,3-trihydroxybenzene by toluene 4-monooxygenase of Pseudomonas mendocina KR1 and toluene 3-monooxygenase of Ralstonia pickettii PKO1. Appl Environ Microbiol 70:3814–3820
Vardar G, Wood TK (2004) Protein engineering of toluene-o-xylene monooxygenase from Pseudomonas stutzeri OX1 for synthesizing 4-methylresorcinol, methylhydroquinone, and pyrogallol. Appl Environ Microbiol 70:3253–3262
Wallar BJ, Lipscomb JD (1996) Dioxygen activation by enzymes containing binuclear non-heme iron clusters. Chem Rev 96:2625–2657
Wick CB (1995) Genencor international takes a green route to blue dye. Genet Eng News 15:22
Zhao H, Arnold FH (1997) Optimization of DNA shuffling for high fidelity recombination. Nucleic Acids Res 25:1307–1308
Acknowledgements
This research was supported by the National Science Foundation (BES-9911469). We thank Albert Kind, University of Connecticut, for helping with LC-MS and appreciate the donation of 6-hydroxyindole from Matrix Scientific (Columbia, S.C.).
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Rui, L., Reardon, K.F. & Wood, T.K. Protein engineering of toluene ortho-monooxygenase of Burkholderia cepacia G4 for regiospecific hydroxylation of indole to form various indigoid compounds. Appl Microbiol Biotechnol 66, 422–429 (2005). https://doi.org/10.1007/s00253-004-1698-z
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00253-004-1698-z