Abstract
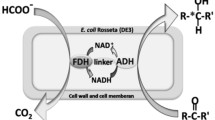
A whole-cell biotransformation system for the reduction of prochiral carbonyl compounds, such as methyl acetoacetate, to chiral hydroxy acid derivatives [methyl (R)-3-hydroxy butanoate] was developed in Escherichia coli by construction of a recombinant oxidation/reduction cycle. Alcohol dehydrogenase from Lactobacillus brevis catalyzes a highly regioselective and enantioselective reduction of several ketones or keto acid derivatives to chiral alcohols or hydroxy acid esters. The adh gene encoding for the alcohol dehydrogenase of L. brevis was expressed in E. coli. As expected, whole cells of the recombinant strain produced only low quantities of methyl (R)-3-hydroxy butanoate from the substrate methyl acetoacetate. Therefore, the fdh gene from Mycobacterium vaccae N10, encoding NAD+-dependent formate dehydrogenase, was functionally coexpressed. The resulting two-fold recombinant strain exhibited an in vitro catalytic alcohol dehydrogenase activity of 6.5 units mg−1 protein in reducing methyl acetoacetate to methyl (R)-3-hydroxy butanoate with NADPH as the cofactor and 0.7 units mg−1 protein with NADH. The in vitro formate dehydrogenase activity was 1.3 units mg−1 protein. Whole resting cells of this strain catalyzed the formation of 40 mM methyl (R)-3-hydroxy butanoate from methyl acetoacetate. The product yield was 100 mol% at a productivity of 200 μmol g−1 (cell dry weight) min−1. In the presence of formate, the intracellular [NADH]/[NAD+] ratio of the cells increased seven-fold. Thus, the functional overexpression of alcohol dehydrogenase in the presence of formate dehydrogenase was sufficient to enable and sustain the desired reduction reaction via the relatively low specific activity of alcohol dehydrogenase with NADH, instead of NADPH, as a cofactor.

Similar content being viewed by others
References
Bradford MM (1976) A rapid and sensitive method for the quantitation of microgram quantities of protein utilizing the principle of protein–dye binding. Anal Biochem 72:248–254
Doig SD, Simpson H, Alphand V, Furstoss R, Woodley JM (2003) Characterization of a recombinant Escherichia coli TOP10[pQR239] whole-cell biocatalyst for stereoselective Baeyer–Villiger oxidations. Enzyme Microb Technol 32:347–355
Endo T, Koizumi S (2001) Microbial conversion with cofactor regeneration using genetically engineered bacteria. Adv Synth Catal 343:521–526
Galkin A, Kulakova L, Tishkov V, Esaki N, Soda K (1995) Cloning of formate dehydrogenase gene from a methanol-utilizing bacterium Mycobacterium vaccae N10. Appl Microbiol Biotechnol 44:479–483
Galkin A, Kulakova L, Yoshimura T, Soda K, Esaki N (1997) Synthesis of optically active amino acids from alpha-keto acids with Escherichia coli cells expressing heterologous genes. Appl Environ Microbiol 63:4651–4656
Gul-Karaguler N, Sessions RB, Clarke AR, Holbrook JJ (2001) A single mutation in the NAD+-specific formate dehydrogenase from Candida methylica allows the enzyme to use NADP. Biotechnol Lett 23:283–287
Haberland J, Hummel W, Dausmann T, Liese A (2002) New continuous production process for enantiopure (2R, 5R)-hexanediol. Org Proc Res Dev 6:458–462
Ham W-H, Oh C-Y, Lee Y-S, Jeong J-H (2000) A new synthesis of a key intermediate of β-lactam antibiotics via diastereoselective alkylation of β-hydroxy ester. J Org Chem 65:8372–8374
Hanahan D (1983) Studies on transformation of Escherichia coli with plasmids. J Mol Biol 166:557–580
Hummel W (1997) New alcohol dehydrogenases for the synthesis of chiral compounds. Adv Biochem Eng Biotechnol 58:145–184
Hummel W (1999) Large-scale applications of NAD(P)-dependent oxidoreductases: recent developments. Trends Biotechnol 17:487–491
Hummel W, Riebel B (2002) (R)-Specific alcohol dehydrogenases from Lactobacillus with improved catalytic activity using an NAD+ substrate. US patent 6,413,750 B1
Kataoka M, Yamamoto K, Kawabata H, Wada M, Kita K, Yanase H, Shimizu S (1999) Stereoselective reduction of ethyl 4-chloro-3-oxobutanoate by Escherichia coli transformant cells coexpressing the aldehyde reductase and glucose dehydrogenase genes. Appl Microbiol Biotechnol 51:486–490
Kataoka M, Kita K, Wada M, Yasohara Y, Hasegawa J, Shimizu S (2003) Novel bioreduction system for the production of chiral alcohols. Appl Microbiol Biotechnol 62:437–445
Kaup B, Bringer-Meyer S, Sahm H (2004) Metabolic engineering of Escherichia coli: construction of an efficient biocatalyst for d-mannitol formation in a whole-cell biotransformation. Appl Microbiol Biotechnol 64:333–339
Kihumbu D, Stillger T, Hummel W, Liese A (2002) Enzymatic synthesis of all stereoisomers of 1-phenylpropane-1,2-diol. Tetrahedron Asymm 13:1069–1072
Kovach EM, Elzer PH, Hill DS, Robertson GT, Farris MA, Roop RM II, Peterson KM (1995) Four new derivatives of the broad-host-range cloning vector pBBR1MCS, carrying different antibiotic-resistance cassettes. Gene 166:175–176
Miller JH (1972) Experiments in molecular genetics. Cold Spring Harbor Laboratory, Cold Spring Harbor, N.Y., pp 352–355
Niefind K, Müller J, Riebel B, Hummel W, Schomburg D (2003) The crystal structure of R-specific alcohol dehydrogenase from Lactobacillus brevis suggests the structural basis of metal dependency. J Mol Biol 327:317–328
Okuda K, Urabe I, Okada H (1985) Coenzymic activity of NADP derivatives alkylated at 2′-phosphate and 6-amino groups. Eur J Biochem 147:249–253
Rabinowitz MH, Andrews RC, Becherer JD, Bickett DM, Bubacz DG, Conway JG, Cowan DJ, Gaul M, Glennon K, Lambert MH, Leesnitzer MA, McDougald DL, Moss ML, Musso DL, Rizzolio MC (2001) Design of selective and soluble inhibitors of tumor necrosis factor-alpha converting enzyme (TACE). J Med Chem 44:4252–4267
Reichmuth DS, Hittle JL, Blanch HW, Keasling JD (2000) Biodesulfurization of dibenzothiophene in Escherichia coli is enhanced by expression of a Vibrio harveyi oxidoreductase gene. Biotechnol Bioeng 67:72–79
Riebel B (1996) Biochemische und molekularbiologische Charakterisierung neuer mikrobieller NAD(P)-abhängiger Alkoholdehydrogenasen. PhD thesis, University of Düsseldorf, Düsseldorf
Sambrook J, Fritsch EF, Maniatis T (1989) Molecular cloning: a laboratory manual, 2nd edn. Cold Spring Harbor Laboratory, Cold Spring Harbor, N.Y.
Schoemaker HE, Mink D, Wubbolts MG (2003) Dispelling the myths—biocatalysis in industrial synthesis. Science 299:16941697
Schubert T, Hummel W, Kula MR, Müller M (2001) Enantioselective synthesis of both enantiomers of various propargylic alcohols by use of two oxidoreductases. Eur J Org Chem 4181–4187
Schubert T, Hummel W, Müller M (2002) Highly enantioselective preparation of multifunctionalized propargylic building blocks. Angew Chem Int Edn 41:634–637
Schütte H, Flossdorf J, Sahm H, Kula M-R (1976) Purification and properties of formaldehyde dehydrogenase and formate dehydrogenase from Candida boidinii. Eur J Biochem 62:151–160
Serov AE, Popova AS, Fedorchuk VV, Tishkov VI (2002) Engineering of coenzyme specificity of formate dehydrogenase from Saccharomyces cerevisiae. Biochem J 367:841–847
Shaked Z, Whitesides GM (1980) Enzyme-catalyzed organic synthesis: NADH regeneration by using formate dehydrogenase. J Am Chem Soc 102:7105–7107
Tchieu JH, Norris V, Edwards JS, Saier MH Jr (2001) The complete phosphotransferase system in Escherichia coli. J Mol Microbiol Biotechnol 3:329–346
Tischer W, Tiemeyer W, Simon H (1980) Stereospecific hydrogenations with immobilized microbial cells or enzymes. Biochimie 62:331–339
Tishkov VI, Galkin AG, Fedorchuk VV, Savitzky PA, Rojkova AM, Gieren H, Kula, M-R (1999) Pilot scale production and isolation of recombinant NAD+- and NADP+-specific formate dehydrogenases. Biotechnol Bioeng 64:187–193
Walton AZ, Stewart JD (2002) An efficient enzymatic Baeyer–Villiger oxidation by engineered Escherichia coli cells under non-growing conditions. Biotechnol Prog 18:262–268
Wolberg M, Hummel W, Wandrey C, Müller M (2000) Highly regio and enantioselective reduction of 3,5-dioxocarboxylates. Angew Chem Int Edn 39:4306–4308
Wolberg M, Hummel W, Müller M (2001) Biocatalytic reduction of β,δ-diketo esters: a highly stereoselective approach to all four stereoisomers of a chlorinated β,δ-dihydroxy hexanoate. Chem Eur J 7:4562–4571
Acknowledgements
We wish to thank N. Esaki for plasmid pMcFDH and thank W. Hummel and A. Weckbecker for valuable discussions and GC analyses. We also thank the Fonds der Chemischen Industrie. This work was supported by the Institut für Technologie der Kohlenhydrate-Zuckerinstitut e. V.-Braunschweig, Germany. All experiments complied with current German laws.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Ernst, M., Kaup, B., Müller, M. et al. Enantioselective reduction of carbonyl compounds by whole-cell biotransformation, combining a formate dehydrogenase and a (R)-specific alcohol dehydrogenase. Appl Microbiol Biotechnol 66, 629–634 (2005). https://doi.org/10.1007/s00253-004-1765-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00253-004-1765-5