Abstract
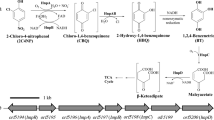
A pure bacterial culture was isolated by its ability to utilize 3-nitrotoluene (3NT) as the sole source of carbon, nitrogen, and energy for growth. Analysis of its 16S rRNA gene showed that the organism (strain ZWL3NT) belongs to the genus Rhodococcus. A rapid disappearance of 3NT with concomitant release of nitrite was observed when strain ZWL3NT was grown on 3NT. The isolate also grew on 2-nitrotoluene, 3-methylcatechol and catechol. Two metabolites, 3-methylcatechol and 2-methyl-cis,cis-muconate, in the reaction mixture were detected after incubation of cells of strain ZWL3NT with 3NT. Enzyme assays showed the presence of both catechol 1,2-dioxygenase and catechol 2,3-dioxygenase in strain ZWL3NT. In addition, a catechol degradation gene cluster (catRABC cluster) for catechol ortho-cleavage pathway was cloned from this strain and cell extracts of Escherichia coli expressing CatA and CatB exhibited catechol 1,2-dioxygenase activity and cis,cis-muconate cycloisomerase activity, respectively. These experimental evidences suggest a novel pathway for 3NT degradation with 3-methylcatechol as a key metabolite by Rhodococcus sp. strain ZWL3NT.


Similar content being viewed by others
References
Alisadat S, Mohan KS, Walia SK (1995) A novel pathway for the biodegradation of 3-nitrotoluene in Pseudomonas putida. FEMS Microbiol Ecol 17(3):169–176. doi:10.1111/j.1574-6941.1995.tb00140.x
An DM, Gibson DT, Spain JC (1994) Oxidative release of nitrite from 2-nitrotoluene by a 3-component enzyme system from Pseudomonas sp strain JS42. J Bacteriol 176(24):7462–7467
Bradford MM (1976) A rapid and sensitive method for the quantitation of microgram quantities of protein utilizing the principle of protein-dye binding. Anal Biochem 72:248–254
Bruning T, Chronz C, Thier R, Havelka J, Ko Y, Bolt HM (1999) Occurrence of urinary tract tumors in miners highly exposed to dinitrotoluene. J Occup Environ Med 41(3):144–149
Dorn E, Knackmuss HJ (1978) Chemical-structure and biodegradability of halogenated aromatic-compounds — substituent effects on 1,2-dioxygenation of catechol. Biochem J 174(1):85–94
Fuchs K, Schreiner A, Lingens F (1991) Degradation of 2-methylaniline and chlorinated isomers of 2-methylaniline by Rhodococcus rhodochrous strain CTM. J Gen Microbiol 137:2033–2039
Gibson DT, Koch JR, Kallio RE (1968) Oxidative degradation of aromatic hydrocarbons by microorganisms: I. Enzymatic formation of catechol from benzene. Biochemistry 7(7):2653–2662. doi:10.1021/bi00847a031
Haigler BE, Spain JC (1993) Biodegradation of 4-nitrotoluene by pseudomonas sp strain 4NT. Appl Environ Microbiol 59(7):2239–2243
Haigler BE, Wallace WH, Spain JC (1994) Biodegradation of 2-nitrotoluene by pseudomonas sp strain JS42. Appl Environ Microbiol 60(9):3466–3469
Hughes MA, Williams PA (2001) Cloning and characterization of the pnb genes, encoding enzymes for 4-nitrobenzoate catabolism in Pseudomonas putida TW3. J Bacteriol 183(4):1225–1232. doi:10.1128/jb.183.4.1225-1232.2001
James KD, Hughes MA, Williams PA (2000) Cloning and expression of ntnD, encoding a novel NAD(P)(+)-independent 4-nitrobenzyl alcohol dehydrogenase from Pseudomonas sp strain TW3. J Bacteriol 182(11):3136–3141. doi:10.1128/jb.182.11.3136-3141.2000
Ju KS, Parales RE (2010) Nitroaromatic compounds, from synthesis to biodegradation. Microbiol Mol Biol Rev 74(2):250–272. doi:10.1128/mmbr.00006-10
Ju KS, Parales RE (2011) Evolution of a new bacterial pathway for 4-nitrotoluene degradation. Mol Microbiol 82(2):355–364. doi:10.1111/j.1365-2958.2011.07817.x
Kim Y, Choi B, Lee J, Chang H, Rak Min K (1992) Characterization of catechol 2,3-dioxygenases. Biochem Biophys Res Commun 183(1):77–82. doi:10.1016/0006-291x(92)91611-s
Lessner DJ, Johnson GR, Parales RE, Spain JC, Gibson DT (2002) Molecular characterization and substrate specificity of nitrobenzene dioxygenase from Comamonas sp strain JS765. Appl Environ Microbiol 68(2):634–641. doi:10.1128/aem.68.2.634-641.2002
Liu H, Wang SJ, Zhou NY (2005) A new isolate of Pseudomonas stutzeri that degrades 2-chloronitrobenzene. Biotechnol Lett 27(4):275–278. doi:10.1007/s10529-004-8293-3
Liu H, Wang SJ, Zhang JJ, Dai H, Tang HR, Zhou NY (2011) Patchwork assembly of nag-like nitroarene dioxygenase genes and the 3-chlorocatechol degradation cluster for evolution of the 2-chloronitrobenzene catabolism pathway in Pseudomonas stutzeri ZWLR2-1. Appl Environ Microbiol 77(13):4547–4552. doi:10.1128/aem.02543-10
Matsumura E, Sakai M, Hayashi K, Murakami S, Takenaka S, Aoki K (2006) Constitutive expression of catABC genes in the aniline-assimilating bacterium Rhodococcus species AN-22: production, purification, characterization and gene analysis of CatA, CatB and CatC. Biochem J 393:219–226. doi:10.1042/bj20050740
Meyer D, Witholt B, Schmid A (2005) Suitability of recombinant Escherichia coli and Pseudomonas putida strains for selective biotransformation of m-nitrotoluene by xylene monooxygenase. Appl Environ Microbiol 71(11):6624–6632. doi:10.1128/aem.71.11.6624-6632.2005
Mulla SI, Hoskeri RS, Shouche YS, Ninnekar HZ (2011) Biodegradation of 2-nitrotoluene by Micrococcus sp strain SMN-1. Biodegradation 22(1):95–102. doi:10.1007/s10532-010-9379-3
Murakami S, Takemoto J, Takashima A, Shinke R, Aoki K (1998) Purification and characterization of two muconate cycloisomerase isozymes from aniline-assimilating Frateuria species ANA-18. Biosci Biotechnol Biochem 62(6):1129–1133. doi:10.1271/bbb.62.1129
Nishino SF, Spain JC (1995) Oxidative pathway for the biodegradation of nitrobenzene by Comamonas sp strain JS765. Appl Environ Microbiol 61(6):2308–2313
Parales JV, Kumar A, Parales RE, Gibson DT (1996) Cloning and sequencing of the genes encoding 2-nitrotoluene dioxygenase from Pseudomonas sp. JS42. Gene 181((1–2):57–61. doi:10.1016/s0378-1119(96)00462-3
Parales JV, Parales RE, Resnick SM, Gibson DT (1998) Enzyme specificity of 2-nitrotoluene 2,3-dioxygenase from Pseudomonas sp. strain JS42 is determined by the C-terminal region of the alpha subunit of the oxygenase component. J Bacteriol 180(5):1194–1199
Rhyswilliams W, Taylor SC, Williams PA (1993) A novel pathway for the catabolism of 4-nitrotoluene by pseudomonas. J Gen Microbiol 139:1967–1972
Robertson JB, Spain JC, Haddock JD, Gibson DT (1992) Oxidation of nitrotoluenes by toluene dioxygenase: evidence for a monooxygenase reaction. Appl Environ Microbiol 58(8):2643–2648
Sambrook J, Russell WD (2001) Molecular cloning — a laboratory manual, 3rd edn. Cold Spring Harbor Laboratory Press, Cold Spring Harbor
Schäfer A, Tauch A, Jäger W, Kalinowski J, Thierbach G, Pühler A (1994) Small mobilizable multi-purpose cloning vectors derived from the Escherichia coli plasmids pK18 and pK19: selection of defined deletions in the chromosome of Corynebacterium glutamicum. Gene 145(1):69–73
Siebert PD, Chenchik A, Kellogg DE, Lukyanov KA, Lukyanov SA (1995) An improved PCR method for walking in uncloned genomic DNA. Nucleic Acids Res 23(6):1087–1088. doi:10.1093/nar/23.6.1087
Spiess T, Desiere F, Fischer P, Spain JC, Knackmuss HJ, Lenke H (1998) A new 4-nitrotoluene degradation pathway in a Mycobacterium strain. Appl Environ Microbiol 64(2):446–452
Struijs J, Stoltenkamp J (1986) Ultimate biodegradation of 2nitrotoluene, 3nitrotoluene and 4-nitrotoluene. Sci Total Environ 57:161–170. doi:10.1016/0048-9697(86)90020-3
Watanabe C, Egami T, Midorikawa K, Hiraku Y, Oikawa S, Kawanishi S, Murata M (2010) DNA damage and estrogenic activity induced by the environmental pollutant 2-nitrotoluene and its metabolite. Environ Health Prev Med 15(5):319–326. doi:10.1007/s12199-010-0146-1
Zylstra GJ, Gibson DT (1989) Toluene degradation by Pseudomonas putida Fl. Nucleotide sequence of the todClC2BADE genes and their expression in Escherichia coli. J Biol Chem 264(25):14940–14946
Acknowledgments
This work was supported by grants from the National Natural Science Foundation of China (30970031) and the National Basic Research Program of China (973 Program) (2012CB721003).
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Tian, XJ., Liu, XY., Liu, H. et al. Biodegradation of 3-nitrotoluene by Rhodococcus sp. strain ZWL3NT. Appl Microbiol Biotechnol 97, 9217–9223 (2013). https://doi.org/10.1007/s00253-012-4619-6
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00253-012-4619-6