Abstract
Cytochrome P450 monooxygenases (CYP450s) are abundant in eukaryotes, specifically in plants and fungi where they play important roles in the synthesis and degradation of secondary metabolites. In eukaryotes, the best studied “self-sufficient” CYP450s, with a fused redox partner, belong to the CYP505 family. Members of the CYP505 family are generally considered sub-terminal fatty acid hydroxylases. CYP505E3 from Aspergillus terreus, however, gives remarkable in-chain hydroxylation at the ω-7 position of C10 to C16 alkanes and C12 and C14 fatty alcohols. Because CYP505E3 is a promising catalyst for the synthesis of δ-dodecalactone, we set out to delineate the unique ω-7 hydroxylase activity of CYP505E3. CYP505E3 and six additional CYP505Es as well as four closely related CYP505s from four different subfamilies were expressed in Pichia pastoris. Only the CYP505Es, sharing more than 70% amino acid identity, displayed significant ω-7 hydroxylase activity toward 1-dodecanol, dodecanoic acid, and tetradecanoic acid giving products that can readily be converted to δ-dodecalactone. Concentrations of δ-dodecalactone, directly extracted from dodecanoic acid biotransformations, were higher than previously obtained with E. coli. Searches of the UniProt and NCBI databases yielded a total of only 23 unique CYP505Es, all from the Aspergillaceae. Given that CYP505Es with this remarkable activity occur in only a few Aspergillus and Penicillium spp., we further explored the genetic environments in which they occur. These were found to be very distinct environments which include a specific ABC transporter but could not be linked to apparent secondary metabolite gene clusters.
Key points
• Identified CYP505Es share > 70% amino acid identity.
• CYP505Es hydroxylate 1-dodecanol, dodecanoic, and tetradecanoic acid at ω-7 position.
• CYP505E genes occur in Aspergillus and Penicillium spp. near an ABC transporter.
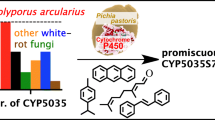
Graphical Abstract






Similar content being viewed by others
Data availability
The data generated and/or analyzed during the current study are available from the corresponding author on reasonable request.
References
Aschenbrenner JC, Ebrecht AC, Tolmie C, Smit MS, Opperman DJ (2021) Structure of the fungal hydroxylase, CYP505A30, and rational transfer of mutation data from CYP102A1 to alter regioselectivity. Catal Sci Technol 11:7359–7367. https://doi.org/10.1039/d1cy01348c
Baker GJ, Girvan HM, Matthews S, McLean KJ, Golovanova M, Waltham TN, Rigby SEJ, Nelson DR, Blankley RT, Munro AW (2017) Expression, purification, and biochemical characterization of the flavocytochrome P450 CYP505A30 from Myceliophthora thermophila. ACS Omega 2:4705–4724. https://doi.org/10.1021/acsomega.7b00450
Blin K, Shaw S, Kloosterman AM, Charlop-Powers Z, Van Wezel GP, Medema MH, Weber T (2021) AntiSMASH 6.0: improving cluster detection and comparison capabilities. Nucleic Acids Res 49:W29–W35. https://doi.org/10.1093/nar/gkab335
Cai P, Wu X, Deng J, Gao L, Shen Y, Yao L, Zhou YJ (2022) Methanol biotransformation toward high-level production of fatty acid derivatives by engineering the industrial yeast Pichia pastoris. Proc Natl Acad Sci 119:1–9. https://doi.org/10.1073/pnas.2201711119
Durairaj P, Hur J-S, Yun H (2016) Versatile biocatalysis of fungal cytochrome P450 monooxygenases. Microb Cell Fact 15:125. https://doi.org/10.1186/s12934-016-0523-6
Ebrecht AC, Aschenbrenner JC, Smit MS, Opperman DJ (2021) Biocatalytic synthesis of non-vicinal aliphatic diols. Org Biomol Chem 19:439–445. https://doi.org/10.1039/d0ob02086a
Edgar RC (2004) MUSCLE: multiple sequence alignment with high accuracy and high throughput. Nucleic Acid Res 32:1792–1797. https://doi.org/10.1093/nar/gkh340
Eser BE, Zhang Y, Zong L, Guo Z (2021) Self-sufficient Cytochrome P450s and their potential applications in biotechnology. Chinese J Chem Eng 30:121–135. https://doi.org/10.1016/j.cjche.2020.12.002
Fessner ND, Srdič M, Weber H, Schmid C, Schönauer D, Schwaneberg U, Glieder A (2020) Preparative-scale production of testosterone metabolites by human liver cytochrome P450 enzyme 3A4. Adv Synth Catal 362:2725–2738. https://doi.org/10.1002/adsc.202000251
Fessner ND, Nelson DR, Glieder A (2021) Evolution and enrichment of CYP5035 in polyporales: functionality of an understudied P450 family. Appl Microbiol Biotechnol 105:6779–6792. https://doi.org/10.1007/s00253-021-11444-2
Gudiminchi RK, Geier M, Glieder A, Camattari A (2013) Screening for cytochrome P450 expression in Pichia pastoris whole cells by P450-carbon monoxide complex determination. Biotechnol J 8:146–152. https://doi.org/10.1002/biot.201200185
Johnston WA, Huang W, De Voss JJ, Hayes MA, Gillam EMJ (2008) Quantitative whole-cell cytochrome P450 measurement suitable for high-throughput application. J Biomol Screen 13:135–141. https://doi.org/10.1177/1087057107312780
Kelly SL, Kelly DE, B PTRS, Kelly SL, Kelly DE, Kelly SL, Kelly DE (2013) Microbial cytochromes P450: biodiversity and biotechnology. Where do cytochromes P450 come from, what do they do and what can they do for us?. Philos Trans R Soc B Biol Sci 368. https://doi.org/10.1098/rstb.2012.0476
Kitazume T, Takaya N, Nakayama N, Shoun H (2000) Fusarium oxysporum fatty-acid subterminal hydroxylase (CYP505) is a membrane-bound eukaryotic counterpart of Bacillus megaterium cytochrome P450BM3. J Biol Chem 275:39734–39740. https://doi.org/10.1074/jbc.M005617200
Li Z, Jiang Y, Guengerich FP, Ma L, Li S, Zhang W (2020) Engineering cytochrome P450 enzyme systems for biomedical and biotechnological applications. J Biol Chem 295:833–849. https://doi.org/10.1016/s0021-9258(17)49939-x
Marella ER, Dahlin J, Dam MI, ter Horst J, Christensen HB, Sudarsan S, Wang G, Holkenbrink C, Borodina I (2020) A single-host fermentation process for the production of flavor lactones from non-hydroxylated fatty acids. Metab Eng 61:427–436. https://doi.org/10.1016/j.ymben.2019.08.009
Maseme MJ, Pennec A, van Marwijk J, Opperman DJ, Smit MS (2020) CYP505E3: a novel self-sufficient ω-7 in-chain hydroxylase. Angew Chemie - Int Ed 59:10359–10362. https://doi.org/10.1002/anie.202001055
Minerdi D, Sadeghi SJ, Pautasso L, Morra S, Aigotti R, Medana C, Gilardi G, Gullino ML, Gilardi G (2020) Expression and role of CYP505A1 in pathogenicity of Fusarium oxysporum f. sp. lactucae. Biochim Biophys Acta - Proteins Proteomics 1868:140268. https://doi.org/10.1016/j.bbapap.2019.140268
Montevecchi G, Vasile Simone G, Masino F, Bignami C, Antonelli A (2012) Physical and chemical characterization of Pescabivona, a Sicilian white flesh peach cultivar [Prunus persica (L.) Batsch]. Food Res Int 45:123–131. https://doi.org/10.1016/j.foodres.2011.10.019
Nakayama N, Takemae A, Shoun H (1996) Cytochrome P450foxy, a catalytically self-sufficient fatty acid hydroxylase of the fungus Fusarium oxysporum. J Biochem 119:435–440. https://doi.org/10.1093/oxfordjournals.jbchem.a021260
Naranjo-Ortiz MA, Gabaldón T (2019) Fungal evolution: diversity, taxonomy and phylogeny of the Fungi. Biol Rev 94:2101–2137. https://doi.org/10.1111/brv.12550
Nelson DR (2006) Cytochrome P450 nomenclature, 2004. Methods Mol Biol 320:1–10. https://doi.org/10.1385/1-59259-998-2:1
Nelson DR (2018) Cytochrome P450 diversity in the tree of life. Biochim Biophys Acta - Proteins Proteomics 1866:141–154. https://doi.org/10.1016/j.bbapap.2017.05.003
Omura T, Sato R (1964) The carbon monoxide-binding pigment of liver microsomes. J Biol Chem 239:2370–2378. https://doi.org/10.1016/S0021-9258(20)82244-3
Pandian BA, Sathishraj R, Djanaguiraman M, Prasad PVV, Jugulam M (2020) Role of cytochrome P450 enzymes in plant stress response. Antioxidants 9:1–15. https://doi.org/10.3390/antiox9050454
Sakai K, Matsuzaki F, Wise L, Sakai Y, Jindou S, Ichinose H, Takaya N, Kato M, Wariishi H, Shimizu M (2018) Biochemical characterization of CYP505D6, a self-sufficient cytochrome P450 from the white-rot fungus Phanerochaete chrysosporium. Appl Environ Microbiol 84. https://doi.org/10.1128/aem.01091-18
Schenkman JB, Jansson I (2006) Spectral analyses of cytochromes P450. Methods Mol Biol 320:11–18. https://doi.org/10.1385/1-59259-998-2:11
Shin JY, Bui D-C, Lee Y, Nam H, Jung S, Fang M, Kim J-C, Lee T, Kim H, Choi GJ, Son H, Lee Y-W (2017) Functional characterization of cytochrome P450 monooxygenases in the cereal head blight fungus Fusarium graminearum. Environ Microbiol 19:2053–2067. https://doi.org/10.1111/1462-2920.13730
Shin J, Kim J-E, Lee Y-W, Son H (2018) Fungal Cytochrome P450s and the P450 complement (CYPome) of Fusarium graminearum. Toxins (basel) 10:112. https://doi.org/10.3390/toxins10030112
Steenwyk JL, Shen X-X, Lind AL, Goldman GH, Rokas A (2019) A robust phylogenomic time tree for biotechnologically and medically important fungi in the genera Aspergillus and Penicillium. MBio 10. https://doi.org/10.1128/mBio.00925-19
Tamura K, Stecher G, Kumar S (2021) MEGA11: molecular evolutionary genetics analysis version 11. Mol Biol Evol 38:3022–3027. https://doi.org/10.1093/molbev/msab120
Uhlig S, Busman M, Shane DS, Rønning H, Rise F, Proctor R (2012) Identification of early fumonisin biosynthetic intermediates by inactivation of the FUM6 gene in Fusarium verticillioides. J Agric Food Chem 60:10293–10301. https://doi.org/10.1021/jf302967b
Waché Y, Aguedo M, LeDall MT, Nicaud JM, Belin JM (2002) Optimization of Yarrowia lipolytica’s β-oxidation pathway for γ-decalactone production. J Mol Catal B Enzym 19–20:347–351. https://doi.org/10.1016/S1381-1177(02)00185-6
Whitehouse CJC, Bell SG, Wong L-L (2011) P450(BM3) (CYP102A1): connecting the dots. Chem Soc Rev 1218–1260. https://doi.org/10.1039/c1cs15192d
Yan Y, Wu J, Hu G, Gao C, Guo L, Chen X, Liu L, Song W (2022) Current state and future perspectives of cytochrome P450 enzymes for C-H and C=C oxygenation. Synth Syst Biotechnol 7:887–899. https://doi.org/10.1016/j.synbio.2022.04.009
Yeny I, Darwo, Nuroniah HS (2020) Sustainability of masoyi (Cryptocarya Massoy (Oken) Kosterm) for essential oil industry materials. IOP Conf Ser Mater Sci Eng 935:012071. https://doi.org/10.1088/1757-899X/935/1/012071
Yoshinaga K, Tago A, Yoshinaga-Kiriake A, Nagai T, Yoshida A, Gotoh N (2019) Effects of heat treatment on lactone content of butter and margarine. J Oleo Sci 68:1295–1301. https://doi.org/10.5650/jos.ess19234
Zhang C, Ma Y, Miao H, Tang X, Xu B, Wu Q, Mu Y, Huang Z (2020) Transcriptomic analysis of Pichia pastoris (Komagataella phaffii) GS115 during heterologous protein production using a high-cell-density fed-batch cultivation strategy. Front Microbiol 11:1–17. https://doi.org/10.3389/fmicb.2020.00463
Zheng X, Li P, Lu X (2019) Research advances in cytochrome P450-catalysed pharmaceutical terpenoid biosynthesis in plants. J Exp Bot 70:4619–4630. https://doi.org/10.1093/jxb/erz203
Zhu GY, Xiao ZB, Zhou RJ, Zhu YL, Niu YW (2013) Study on development of a fresh peach flavor. Adv Mater Res 781–784:1570–1573. https://doi.org/10.4028/www.scientific.net/AMR.781-784.1570
Acknowledgements
Financial support by the South African Department of Science and Technology/National Research Foundation Centre of Excellence in Catalysis is gratefully acknowledged. We are very grateful to Prof David Nelson for naming all previously unclassified CYP505s and for supplying additional relevant sequences. We also thank Sarel Marais for technical assistance with GC analyses.
Funding
This study was funded by the National Research Foundation, South Africa (grant number 132500) and Department of Science and Technology/ National Research Foundation Centre of Excellence in Catalysis, South Africa (grant number PAR02).
Author information
Authors and Affiliations
Contributions
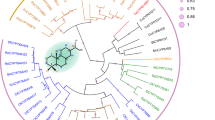
MS and DO conceived and designed the study. JvM cloned the genes and JvM and MM did the experiments. MS analyzed the data, collected the CYP505 sequences, and wrote the original draft. JA did the phylogenetic analysis. All authors read and approved the manuscript.
Corresponding author
Ethics declarations
Ethics approval
This article does not contain any studies with human participants or animals performed by any of the authors.
Conflict of interest
The authors declare no conflict of interest.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Smit, M.S., Maseme, M.J., van Marwijk, J. et al. Delineation of the CYP505E subfamily of fungal self-sufficient in-chain hydroxylating cytochrome P450 monooxygenases. Appl Microbiol Biotechnol 107, 735–747 (2023). https://doi.org/10.1007/s00253-022-12329-8
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00253-022-12329-8