Abstract
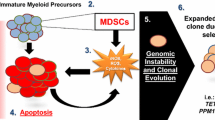
The transformation and progression of myelodysplastic syndromes (MDS) to secondary acute myeloid leukemia (sAML) involve genetic, epigenetic, and microenvironmental factors. Driver mutations have emerged as valuable markers for defining risk groups and as candidates for targeted treatment approaches in MDS. It is also evident that the risk of transformation to sAML is increased by evasion of adaptive immune surveillance. This study was designed to explore the immune microenvironment, immunogenic tumor-intrinsic mechanisms (HLA and PD-L1 expression), and tumor genetic features (somatic mutations and altered karyotypes) in MDS patients and to determine their influence on the progression of the disease. We detected major alterations of the immune microenvironment in MDS patients, with a reduced count of CD4+ T cells, a more frequent presence of markers related to T cell exhaustion, a more frequent presence of myeloid-derived suppressor cells (MDSCs), and changes in the functional phenotype of NK cells. HLA Class I (HLA-I) expression was normally expressed in CD34+ blasts and during myeloid differentiation. Only two out of thirty-six patients with homozygosity for HLA-C groups acquired complete copy-neutral loss of heterozygosity in the HLA region. PD-L1 expression on the leukemic clone was also increased in MDS patients. Finally, no interplay was observed between the anti-tumor immune microenvironment and mutational genomic features. In summary, extrinsic and intrinsic immunological factors might severely impair immune surveillance and contribute to clonal immune escape. Genomic alterations appear to make an independent contribution to the clonal evolution and progression of MDS.



Similar content being viewed by others
Abbreviations
- HLA-I:
-
HLA Class I
- HMR:
-
High molecular risk
- IPSS:
-
International prognostic scoring system
- IPSS-R:
-
International prognostic scoring system revised
- LOH HLA:
-
Loss of heterozygosity in the HLA region
- LOH:
-
Loss of heterozygosity
- MDS del(5q):
-
MDS with isolated del(5q)
- MDS EB:
-
MDS with excess blasts
- MDS:
-
Myelodysplastic syndromes
- MDSCs:
-
Myeloid derived suppressor cells
- MDS-MLD:
-
MDS with multilineage dysplasia
- MDS-RS:
-
MLD and ring sideroblasts
- MDS-SLD:
-
MDS with single lineage dysplasia
- MoAbs:
-
Monoclonal antibodies
- NGS:
-
Next-generation sequencing
- PB:
-
Peripheral blood
- sAML:
-
Secondary acute myeloid leukemia
References
Germing U, Kobbe G, Haas R, Gattermann N (2013) Myelodysplastic syndromes: diagnosis, prognosis, and treatment. Dtsch Arztebl Int 110:783–790. https://doi.org/10.3238/arztebl.2013.0783
Neukirchen J, Schoonen WM, Strupp C, Gattermann N, Aul C, Haas R, Germing U (2011) Incidence and prevalence of myelodysplastic syndromes: data from the Düsseldorf MDS-registry. Leuk Res 35:1591–1596. https://doi.org/10.1016/j.leukres.2011.06.001
Bennett JM (2016) Changes in the updated 2016: WHO Classification of the myelodysplastic syndromes and related myeloid neoplasms. Clin Lymphoma Myeloma Leuk 16:607–609. https://doi.org/10.1016/j.clml.2016.08.005
Greenberg P, Cox C, LeBeau MM, Fenaux P, Morel P, Sanz G, Sanz M, Vallespi T, Hamblin T, Oscier D, Ohyashiki K, Toyama K, Aul C, Mufti G, Bennett J (1997) International scoring system for evaluating prognosis in myelodysplastic syndromes. Blood 89:2079–2088
Greenberg PL, Tuechler H, Schanz J, Sanz G, Garcia-Manero G, Solé F, Bennett JM, Bowen D, Fenaux P, Dreyfus F, Kantarjian H, Kuendgen A, Levis A, Malcovati L, Cazzola M, Cermak J, Fonatsch C, Le Beau MM, Slovak ML, Krieger O, Luebbert M, Maciejewski J, Magalhaes SM, Miyazaki Y, Pfeilstöcker M, Sekeres M, Sperr WR, Stauder R, Tauro S, Valent P, Vallespi T, van de Loosdrecht AA, Germing U, Haase D (2012) Revised international prognostic scoring system for myelodysplastic syndromes. Blood 120:2454–2465. https://doi.org/10.1182/blood-2012-03-420489
Wall M (2017) Recurrent cytogenetic abnormalities in myelodysplastic syndromes. Methods Mol Biol 1541:209–222
Pellagatti A, Boultwood J (2015) The molecular pathogenesis of the myelodysplastic syndromes. Eur J Haematol 95:3–15. https://doi.org/10.1111/ejh.12515
Papaemmanuil E, Gerstung M, Malcovati L, Tauro S, Gundem G, Van Loo P, Yoon CJ, Ellis P, Wedge DC, Pellagatti A, Shlien A, Groves MJ, Forbes SA, Raine K, Hinton J, Mudie LJ, McLaren S, Hardy C, Latimer C, Della Porta MG, O’Meara S, Ambaglio I, Galli A, Butler AP, Walldin G, Teague JW, Quek L, Sternberg A, Gambacorti-Passerini C, Cross NC, Green AR, Boultwood J, Vyas P, Hellstrom-Lindberg E, Bowen D, Cazzola M, Stratton MR, Campbell PJ (2013) Clinical and biological implications of driver mutations in myelodysplastic syndromes. Blood 122:3616–3627. https://doi.org/10.1182/blood-2013-08-518886
Bejar R, Steensma DP (2014) Recent developments in myelodysplastic syndromes. Blood 124:2793–2803. https://doi.org/10.1182/blood-2014-04-522136
Wang C, Yang Y, Gao S, Chen J, Yu J, Zhang H, Li M, Zhan X, Li W (2018) Immune dysregulation in myelodysplastic syndrome: clinical features, pathogenesis and therapeutic strategies. Crit Rev Oncol Hematol 122:123–132. https://doi.org/10.1016/j.critrevonc.2017.12.013
Ivy KS, Brent Ferrell P Jr (2018) Disordered immune regulation and its therapeutic targeting in myelodysplastic syndromes. Curr Hematol Malig Rep 13:244–255. https://doi.org/10.1007/s11899-018-0463-9
Wolach O, Stone R (2016) Autoimmunity and inflammation in myelodysplastic syndromes. Acta Haematol 136:108–117. https://doi.org/10.1159/000446062
Kook H, Zeng W, Guibin C, Kirby M, Young NS, Maciejewski JP (2001) Increased cytotoxic T cells with effector phenotype in aplastic anemia and myelodysplasia. Exp Hematol 29:1270–1277
Bouchliou I, Miltiades P, Nakou E, Spanoudakis E, Goutzouvelidis A, Vakalopoulou S, Garypidou V, Kotoula V, Bourikas G, Tsatalas C, Kotsianidis I (2011) Th17 and Foxp3(+) T regulatory cell dynamics and distribution in myelodysplastic syndromes. Clin Immunol 139:350–359. https://doi.org/10.1016/j.clim.2011.03.001
Aggarwal S, van de Loosdrecht AA, Alhan C, Ossenkoppele GJ, Westers TM, Bontkes HJ (2011) Role of immune responses in the pathogenesis of low-risk MDS and high-risk MDS: implications for immunotherapy. Br J Haematol 153:568–581. https://doi.org/10.1111/j.1365-2141.2011.08683x
Epling-Burnette PK, Bai F, Painter JS, Rollison DE, Salih HR, Krusch M, Zou J, Ku E, Zhong B, Boulware D, Moscinski L, Wei S, Djeu JY, List AF (2007) Reduced natural killer (NK) function associated with high-risk myelodysplastic syndrome (MDS) and reduced expression of activating NK receptors. Blood 109:4816–4824
Yu H, Boyle TA, Zhou C, Rimm DL, Hirsch FR (2016) PD-L1 expression in lung cancer. J Thorac Oncol 11:964–975. https://doi.org/10.1016/j.jtho.2016.04.014
Jelinek T, Mihalyova J, Kascak M, Duras J, Hajek R (2017) PD-1/PD-L1 inhibitors in haematological malignancies: update 2017. Immunology 152:357–371. https://doi.org/10.1111/imm.12788Epub 2017 Aug 4
Montes P, Kerick M, Bernal M, Hernández F, Jiménez P, Garrido P, Márquez A, Jurado M, Martin J, Garrido F, Ruiz-Cabello F (2018) Genomic loss of HLA alleles may affect the clinical outcome in low-risk myelodysplastic syndrome patients. Oncotarget 9:36929–36944. https://doi.org/10.18632/oncotarget.26405
Del Mar Valenzuela-Membrives M, Perea-García F, Sanchez-Palencia A, Ruiz-Cabello F, Gómez-Morales M, Miranda-León MT, Galindo-Angel I, Fárez-Vidal ME (2016) Progressive changes in composition of lymphocytes in lung tissues from patients with non-small-cell lung cancer. Oncotarget 7:71608–71619. https://doi.org/10.18632/oncotarget.12264
Kittang AO, Kordasti S, Sand KE, Costantini B, Kramer AM, Perezabellan P, Seidl T, Rye KP, Hagen KM, Kulasekararaj A, Bruserud Ø, Mufti GJ (2015) Expansion of myeloid derived suppressor cells correlates with number of T regulatory cells and disease progression in myelodysplastic syndrome. Oncoimmunology 5:e1062208
Garrido F, Ruiz-Cabello F, Cabrera T, Pérez-Villar JJ, López-Botet M, Duggan-Keen M, Stern PL (1997) Implications for immunosurveillance of altered HLA Class I phenotypes in human tumours. Immunol Today 18:89–95
Seliger B, Cabrera T, Garrido F, Ferrone S (2002) HLA Class I antigen abnormalities and immune escape by malignant cells. Semin Cancer Biol 12:3–13. https://doi.org/10.1006/scbi.2001.0404
Glenthøj A, Ørskov AD, Hansen JW, Hadrup SR, O’Connell C, Grønbæk K (2016) Immune mechanisms in myelodysplastic syndrome. Int J Mol Sci. https://doi.org/10.3390/ijms17060944
Yu S, Liu C, Zhang L, Shan B, Tian T, Hu Y, Shao L, Sun Y, Ji C, Ma D (2014) Elevated Th22 cells correlated with Th17 cells in peripheral blood of patients with acute myeloid leukemia. Int J Mol Sci 15:1927–1945. https://doi.org/10.3390/ijms15021927
Tian T, Sun Y, Li M, He N, Yuan C, Yu S, Wang M, Ji C, Ma D (2013) Increased Th22 cells as well as Th17 cells in patients with adult T-cell acute lymphoblastic leukemia. Clin Chim Acta 426:108–113. https://doi.org/10.1016/j.cca.2013.09.014
Canale FP, Ramello MC, Núñez N, Araujo Furlan CL, Bossio SN, Gorosito Serrán M, Tosello Boari J, Del Castillo A, Ledesma M, Sedlik C, Piaggio E, Gruppi A, Acosta Rodríguez EA, Montes CL (2018) CD39 expression defines cell exhaustion in tumor-infiltrating CD8(+) T cells. Cancer Res 78:115–128. https://doi.org/10.1158/0008-5472.can-16-2684
Perry C, Hazan-Halevy I, Kay S, Cipok M, Grisaru D, Deutsch V, Polliack A, Naparstek E, Herishanu Y (2012) Increased CD39 expression on CD4(+) T lymphocytes has clinical and prognostic significance in chronic lymphocytic leukemia. Ann Hematol 91:1271–1279. https://doi.org/10.1007/s00277-012-1425-2
Cichocki F, Schlums H, Theorell J, Tesi B, Miller JS, Ljunggren HG, Bryceson YT (2016) Diversification and functional specialization of human NK cell subsets. Curr Top Microbiol 395:63–93
Costello RT, Fauriat C, Sivori S, Marcenaro E, Olive D (2004) NK cells: innate immunity against hematological malignancies? Trends Immunol 25:328–333
Pietra G, Vitale M, Moretta L, Mingari MC (2012) How melanoma cells inactivate NK cells. Oncoimmunology 1:974–975
Liu J, Zhou Y, Huang Q, Qiu L (2014) CD14(+) HLA-DR (low/−) expression: a novel prognostic factor in chronic lymphocytic leukemia. Oncol Lett 9:1167–1172
Yang H, Bueso-Ramos C, DiNardo C, Estecio MR, Davanlou M, Geng QR, Fang Z, Nguyen M, Pierce S, Wei Y, Parmar S, Cortes J, Kantarjian H, Garcia-Manero G (2014) Expression of PD-L1, PD-L2, PD-1 and CTLA4 in myelodysplastic syndromes is enhanced by treatment with hypomethylating agents. Leukemia 28:1280–1288. https://doi.org/10.1038/leu.2013.355
Perea F, Bernal M, Sánchez-Palencia A, Carretero J, Torres C, Bayarri C, Gómez-Morales M, Garrido F, Ruiz-Cabello F (2017) The absence of HLA Class I expression in non-small cell lung cancer correlates with the tumor tissue structure and the pattern of T cell infiltration. Int J Cancer 140:888–899. https://doi.org/10.1002/ijc.30489
McGranahan N, Rosenthal R, Hiley CT, Rowan AJ, Watkins TBK, Wilson GA, Birkbak NJ, Veeriah S, Van Loo P, Herrero J, Swanton C (2017) Allele-specific HLA loss and immune escape in lung cancer evolution. Cell 171:1259–1271. https://doi.org/10.1016/j.cell.2017.10.001
Carretero R, Romero JM, Ruiz-Cabello F, Maleno I, Rodriguez F, Camacho FM, Real LM, Garrido F, Cabrera T (2008) Analysis of HLA Class I expression in progressing and regressing metastatic melanoma lesions after immunotherapy. Immunogenetics 60:439–447. https://doi.org/10.1007/s00251-008-0303-5
Brouwer RE, van der Heiden P, Schreuder GM, Mulder A, Datema G, Anholts JD, Willemze R, Claas FH, Falkenburg JH (2002) Loss or downregulation of HLA Class I expression at the allelic level in acute leukemia is infrequent but functionally relevant and can be restored by interferon. Hum Immunol 63:200–210
Wetzler M, Baer MR, Stewart SJ, Donohue K, Ford L, Stewart CC, Repasky EA, Ferrone S (2001) HLA Class I antigen cell surface expression is preserved on acute myeloid leukemia blasts at diagnosis and at relapse. Leukemia 15:128–133
Abushok DV, Duke JL, Xie HM, Stanley N, Atienza J, Perdigones N, Nicholas P, Ferriola D, Li Y, Huang H, Ye W, Morrissette JJD, Kearns J, Porter DL, Podsakoff GM, Eisenlohr LC, Biegel JA, Chou ST, Monos DS, Bessler M, Olson TS (2017) Somatic HLA mutations expose the role of class I-mediated autoimmunity in aplastic anemia and its clonal complications. Blood Adv 1:1900–1910
Jordanova ES, Riemersma SA, Philippo K, Schuuring E, Kluin PM (2003) Beta2-microglobulin aberrations in diffuse large B-cell lymphoma of the testis and the central nervous system. Int J Cancer 103:393–398
Perea F, Sánchez-Palencia A, Gómez-Morales M, Bernal M, Concha Á, García MM, González-Ramírez AR, Kerick M, Martin J, Garrido F, Ruiz-Cabello F, Aptsiauri N (2017) HLA class I loss and PD-L1 expression in lung cancer: impact on T-cell infiltration and immune escape. Oncotarget 9:4120–4133. https://doi.org/10.18632/oncotarget.23469
Rosenthal R, Cadieux EL, Salgado R, Bakir MA, Moore DA, Hiley CT, Lund T, Tanić M, Reading JL, Joshi K, Henry JY, Ghorani E, Wilson GA, Birkbak NJ, Jamal-Hanjani M, Veeriah S, Szallasi Z, Loi S, Hellmann MD, Feber A, Chain B, Herrero J, Quezada SA, Demeulemeester J, Van Loo P, Beck S, McGranahan N, Swanton C (2019) Neoantigen-directed immune escape in lung cancer evolution. Nature 567:479–485. https://doi.org/10.1038/s41586-019-1032-7
Jordanova ES, Riemersma SA, Philippo K, Giphart-Gassler M, Schuuring E, Kluin PM (2002) Hemizygous deletions in the HLA region account for loss of heterozygosity in the majority of diffuse large B-cell lymphomas of the testis and the central nervous system. Genes Chromosom Cancer 35:38–48
Sebastián E, Alcoceba M, Martín-García D, Blanco Ó, Sanchez-Barba M, Balanzategui A, Marín L, Montes-Moreno S, González-Barca E, Pardal E, Jiménez C, García-Álvarez M, Clot G, Carracedo Á, Gutiérrez NC, Sarasquete ME, Chillón C, Corral R, Prieto-Conde MI, Caballero MD, Salaverria I, García-Sanz R, González M (2016) High-resolution copy number analysis of paired normal-tumor samples from diffuse large B cell lymphoma. Ann Hematol 95:253–262. https://doi.org/10.1007/s00277-015-2552-3
Albertsson PA, Basse PH, Hokland M, Goldfarb RH, Nagelkerke JF, Nannmark U, Kuppen PJ (2003) NK cells and the tumour microenvironment: implications for NK-cell function and anti-tumour activity. Trends Immunol 24:603–609
Acknowledgements
The authors thank Victoria Calvo and María Corzo for technical assistance.
Funding
This work was supported by Grants from the Instituto de Salud Carlos III co-financed by FEDER funds (European Union) (PI 16/00752, PI 17/00197) and Junta de Andalucía in Spain (Group CTS-143, PI09/0382). This study is part of the doctoral thesis of Paola Montes, whose pre-doctoral fellowship was partially financed by Abbott, Becton–Dickinson, Beckman Coulter, and the Spanish MDS group.
Author information
Authors and Affiliations
Contributions
PM and LNC contributed to the immunophenotypic analysis of the tumor microenvironment. PM and MB contributed to sequencing and data analysis. FH and PG contributed to the diagnosis and classification of patients based on their clinical and hematological characteristics. ARG-R carried out the statistical analyses. PM, MB, PJ, MJ, FG, and FR-C were involved with all aspects of the study’s design and contributed to the manuscript preparation.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Ethical approval and ethical standards
The procedures with human samples were performed in accordance with the Declaration of Helsinki and the ethical standards of the Research Ethics Committee of Virgen de las Nieves Hospital in Granada, Spain, which approved the project on June 28 2016 (PEIBA code 0713-N-16 and PROYECTO code 555).
Informed consent
Written informed consent was provided by all patients at the time of their diagnosis and by healthy donors at routine analyses during the first few months of the study.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Montes, P., Bernal, M., Campo, L.N. et al. Tumor genetic alterations and features of the immune microenvironment drive myelodysplastic syndrome escape and progression. Cancer Immunol Immunother 68, 2015–2027 (2019). https://doi.org/10.1007/s00262-019-02420-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00262-019-02420-x