Abstract
Purpose
Probiotics have been shown to exert beneficial effects in IBD although their exact mechanisms are not completely understood. The aim of the present study was to assess the intestinal anti-inflammatory activity of different probiotics (Lactobacillus fermentum CECT5716, Lactobacillus salivarius CECT5713, Escherichia coli Nissle 1917, Saccharomyces boulardii CNCMI-745 in the dinitrobenzene sulfonic acid (DNBS) model of mouse colitis and correlate it with the modifications of the gut microbiota and the immune response, focusing on miRNA expression.
Methods
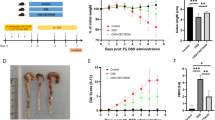
The probiotics were daily administered orally for 25 days. On day 19 colitis was induced by rectal installation of DNBS. At the end of the treatment, mice were sacrificed and the colonic damage was assessed biochemically by analysing the expression of different markers involved in the immune response, including miRNAs; and the colonic microbiota by pyrosequencing. Probiotics properties were also evaluated in vitro in different immune cell types (CMT-93 intestinal epithelial cells and bone marrow-derived macrophages), where the expression of different mRNAs and miRNAs was examined.
Results
All the probiotics displayed intestinal anti-inflammatory effects but slightly different, especially regarding miRNAs expression. Likewise, the probiotics ameliorated the colitis-associated dysbiosis, although showing differences in the main bacterial groups affected.
Conclusion
Among the probiotics assayed, Lactobacillus fermentum CECT5716 and Escherichia coli Nissle 1917 appear to present the best intestinal anti-inflammatory effects, being the latter one of the few probiotics with reputed efficacy in human IBD. Therefore, Lactobacillus fermentum CECT5716 could be considered as a complementary nutritional strategy for IBD treatment.





Similar content being viewed by others
Abbreviations
- ANOVA:
-
Analysis of variance
- AUC:
-
Area under the curve
- BMDM:
-
Bone marrow-derived macrophages
- DAI:
-
Disease activity index
- DMEM:
-
Dulbecco's Modified Eagle Medium
- DNBS:
-
Dinitrobenzene sulfonic acid
- emPCR:
-
Emulsion-based clonal amplification
- Gapdh :
-
Glyceraldehyde-3-phosphate dehydrogenase
- IBD:
-
Inflammatory bowel disease
- iNos :
-
Inducible nitric oxide synthase
- miRNAs:
-
MicroRNAs
- NO:
-
Nitric oxide
- Ocln :
-
Occluding
- RDP:
-
Ribosomal database project
- Snord95:
-
Small nucleolar RNA, C/D box 95
- STAMP:
-
Statistical analysis of metagenomic profiles
References
Strober W, Fuss I, Mannon P (2007) The fundamental basis of inflammatory bowel disease. J Clin Investig 117(3):514–521. https://doi.org/10.1172/JCI30587
Frank DN, St Amand AL, Feldman RA, Boedeker EC, Harpaz N, Pace NR (2007) Molecular-phylogenetic characterization of microbial community imbalances in human inflammatory bowel diseases. Proc Natl Acad Sci USA 104(34):13780–13785. https://doi.org/10.1073/pnas.0706625104
Wang C, Chen J (2019) microRNAs as therapeutic targets in intestinal diseases. ExRNA 1(1):23
Behrouzi A, Ashrafian F, Mazaheri H, Lari A, Nouri M, Riazi Rad F, Hoseini Tavassol Z, Siadat SD (2020) The importance of interaction between MicroRNAs and gut microbiota in several pathways. Microb Pathog 144:104200. https://doi.org/10.1016/j.micpath.2020.104200
Rodriguez-Nogales A, Algieri F, Garrido-Mesa J, Vezza T, Utrilla MP, Chueca N, Garcia F, Olivares M, Rodriguez-Cabezas ME, Galvez J (2017) Differential intestinal anti-inflammatory effects of Lactobacillus fermentum and Lactobacillus salivarius in DSS mouse colitis: impact on microRNAs expression and microbiota composition. Mol Nutr Food Res. https://doi.org/10.1002/mnfr.201700144
Rodriguez-Nogales A, Algieri F, Garrido-Mesa J, Vezza T, Utrilla MP, Chueca N, Garcia F, Rodriguez-Cabezas ME, Galvez J (2018) Intestinal anti-inflammatory effect of the probiotic Saccharomyces boulardii in DSS-induced colitis in mice: Impact on microRNAs expression and gut microbiota composition. J Nutr Biochem 61:129–139. https://doi.org/10.1016/j.jnutbio.2018.08.005
Basso PJ, Camara NOS, Sales-Campos H (2018) Microbial-based therapies in the treatment of inflammatory bowel disease—an overview of human studies. Front Pharmacol 9:1571. https://doi.org/10.3389/fphar.2018.01571
Plaza-Diaz J, Ruiz-Ojeda FJ, Gil-Campos M, Gil A (2019) Mechanisms of action of probiotics. Adv Nutr 10(suppl_1):S49–S66. https://doi.org/10.1093/advances/nmy063
Reid G (2016) Probiotics: definition, scope and mechanisms of action. Best Pract Res Clin Gastroenterol 30(1):17–25. https://doi.org/10.1016/j.bpg.2015.12.001
O’Hara AM, Shanahan F (2007) Mechanisms of action of probiotics in intestinal diseases. Sci World J 7:31–46. https://doi.org/10.1100/tsw.2007.26
Rodriguez-Nogales A, Algieri F, Garrido-Mesa J, Vezza T, Utrilla MP, Chueca N, Fernandez-Caballero JA, Garcia F, Rodriguez-Cabezas ME, Galvez J (2018) The Administration of Escherichia coli Nissle 1917 ameliorates development of DSS-induced colitis in mice. Front Pharmacol 9:468. https://doi.org/10.3389/fphar.2018.00468
Randhawa PK, Singh K, Singh N, Jaggi AS (2014) A review on chemical-induced inflammatory bowel disease models in rodents. Korean J Physiol Pharmacol 18(4):279–288. https://doi.org/10.4196/kjpp.2014.18.4.279
Xaus J, Mirabet M, Lloberas J, Soler C, Lluis C, Franco R, Celada A (1999) IFN-gamma up-regulates the A2B adenosine receptor expression in macrophages: a mechanism of macrophage deactivation. J Immunol 162(6):3607–3614
Qiu BS, Vallance BA, Blennerhassett PA, Collins SM (1999) The role of CD4+ lymphocytes in the susceptibility of mice to stress-induced reactivation of experimental colitis. Nat Med 5(10):1178–1182. https://doi.org/10.1038/13503
Cole JR, Wang Q, Cardenas E, Fish J, Chai B, Farris RJ, Kulam-Syed-Mohideen AS, McGarrell DM, Marsh T, Garrity GM, Tiedje JM (2009) The ribosomal database project: improved alignments and new tools for rRNA analysis. Nucleic Acids Res 37(Database issue):D141-145. https://doi.org/10.1093/nar/gkn879
Parks DH, Tyson GW, Hugenholtz P, Beiko RG (2014) STAMP: statistical analysis of taxonomic and functional profiles. Bioinformatics 30(21):3123–3124. https://doi.org/10.1093/bioinformatics/btu494
Dalal SR, Chang EB (2014) The microbial basis of inflammatory bowel diseases. J Clin Investig 124(10):4190–4196. https://doi.org/10.1172/JCI72330
Derwa Y, Gracie DJ, Hamlin PJ, Ford AC (2017) Systematic review with meta-analysis: the efficacy of probiotics in inflammatory bowel disease. Aliment Pharmacol Ther 46(4):389–400. https://doi.org/10.1111/apt.14203
Baugh MD, Perry MJ, Hollander AP, Davies DR, Cross SS, Lobo AJ, Taylor CJ, Evans GS (1999) Matrix metalloproteinase levels are elevated in inflammatory bowel disease. Gastroenterology 117(4):814–822. https://doi.org/10.1016/s0016-5085(99)70339-2
Lanas A (2008) Role of nitric oxide in the gastrointestinal tract. Arthritis Res Ther 10(Suppl 2):S4. https://doi.org/10.1186/ar2465
Guihot G, Guimbaud R, Bertrand V, Narcy-Lambare B, Couturier D, Duee PH, Chaussade S, Blachier F (2000) Inducible nitric oxide synthase activity in colon biopsies from inflammatory areas: correlation with inflammation intensity in patients with ulcerative colitis but not with Crohn’s disease. Amino Acids 18(3):229–237. https://doi.org/10.1007/s007260050020
Johansson ME (2014) Mucus layers in inflammatory bowel disease. Inflamm Bowel Dis 20(11):2124–2131. https://doi.org/10.1097/MIB.0000000000000117
Tufekci KU, Meuwissen RL, Genc S (2014) The role of microRNAs in biological processes. Methods Mol Biol 1107:15–31. https://doi.org/10.1007/978-1-62703-748-8_2
Griffiths-Jones S, Grocock RJ, van Dongen S, Bateman A, Enright AJ (2006) miRBase: microRNA sequences, targets and gene nomenclature. Nucleic Acids Res 34(Database issue):D140-144. https://doi.org/10.1093/nar/gkj112
Archanioti P, Gazouli M, Theodoropoulos G, Vaiopoulou A, Nikiteas N (2011) Micro-RNAs as regulators and possible diagnostic bio-markers in inflammatory bowel disease. J Crohn’s Colitis 5(6):520–524. https://doi.org/10.1016/j.crohns.2011.05.007
O’Connell RM, Rao DS, Chaudhuri AA, Baltimore D (2010) Physiological and pathological roles for microRNAs in the immune system. Nat Rev Immunol 10(2):111–122. https://doi.org/10.1038/nri2708
Haneklaus M, Gerlic M, Kurowska-Stolarska M, Rainey AA, Pich D, McInnes IB, Hammerschmidt W, O’Neill LA, Masters SL (2012) Cutting edge: miR-223 and EBV miR-BART15 regulate the NLRP3 inflammasome and IL-1beta production. J Immunol 189(8):3795–3799. https://doi.org/10.4049/jimmunol.1200312
Starczynowski DT, Kuchenbauer F, Argiropoulos B, Sung S, Morin R, Muranyi A, Hirst M, Hogge D, Marra M, Wells RA, Buckstein R, Lam W, Humphries RK, Karsan A (2010) Identification of miR-145 and miR-146a as mediators of the 5q- syndrome phenotype. Nat Med 16(1):49–58. https://doi.org/10.1038/nm.2054
Chivukula RR, Shi G, Acharya A, Mills EW, Zeitels LR, Anandam JL, Abdelnaby AA, Balch GC, Mansour JC, Yopp AC, Maitra A, Mendell JT (2014) An essential mesenchymal function for miR-143/145 in intestinal epithelial regeneration. Cell 157(5):1104–1116. https://doi.org/10.1016/j.cell.2014.03.055
Chen CZ, Li L, Lodish HF, Bartel DP (2004) MicroRNAs modulate hematopoietic lineage differentiation. Science 303(5654):83–86. https://doi.org/10.1126/science.1091903
Goto Y, Kiyono H (2011) Epithelial cell microRNAs in gut immunity. Nat Immunol 12(3):195–197. https://doi.org/10.1038/ni0311-195
Biton M, Levin A, Slyper M, Alkalay I, Horwitz E, Mor H, Kredo-Russo S, Avnit-Sagi T, Cojocaru G, Zreik F, Bentwich Z, Poy MN, Artis D, Walker MD, Hornstein E, Pikarsky E, Ben-Neriah Y (2011) Epithelial microRNAs regulate gut mucosal immunity via epithelium-T cell crosstalk. Nat Immunol 12(3):239–246. https://doi.org/10.1038/ni.1994
Garrido-Mesa J, Algieri F, Rodriguez-Nogales A, Vezza T, Utrilla MP, Garcia F, Chueca N, Rodriguez-Cabezas ME, Garrido-Mesa N, Galvez J (2018) Immunomodulatory tetracyclines ameliorate DNBS-colitis: Impact on microRNA expression and microbiota composition. Biochem Pharmacol 155:524–536. https://doi.org/10.1016/j.bcp.2018.07.044
Schaefer JS, Montufar-Solis D, Vigneswaran N, Klein JR (2011) Selective upregulation of microRNA expression in peripheral blood leukocytes in IL-10-/- mice precedes expression in the colon. J Immunol 187(11):5834–5841. https://doi.org/10.4049/jimmunol.1100922
Giahi L, Aumueller E, Elmadfa I, Haslberger AG (2012) Regulation of TLR4, p38 MAPkinase, IkappaB and miRNAs by inactivated strains of lactobacilli in human dendritic cells. Benef Microbes 3(2):91–98. https://doi.org/10.3920/BM2011.0052
Knox NC, Forbes JD, Peterson CL, Van Domselaar G, Bernstein CN (2019) The gut microbiome in inflammatory bowel disease: lessons learned from other immune-mediated inflammatory diseases. Am J Gastroenterol 114(7):1051–1070. https://doi.org/10.14309/ajg.0000000000000305
Yu LC (2018) Microbiota dysbiosis and barrier dysfunction in inflammatory bowel disease and colorectal cancers: exploring a common ground hypothesis. J Biomed Sci 25(1):79. https://doi.org/10.1186/s12929-018-0483-8
Gophna U, Sommerfeld K, Gophna S, Doolittle WF, Veldhuyzen van Zanten SJ (2006) Differences between tissue-associated intestinal microfloras of patients with Crohn’s disease and ulcerative colitis. J Clin Microbiol 44(11):4136–4141. https://doi.org/10.1128/JCM.01004-06
Andoh A, Kuzuoka H, Tsujikawa T, Nakamura S, Hirai F, Suzuki Y, Matsui T, Fujiyama Y, Matsumoto T (2012) Multicenter analysis of fecal microbiota profiles in Japanese patients with Crohn’s disease. J Gastroenterol 47(12):1298–1307. https://doi.org/10.1007/s00535-012-0605-0
Atarashi K, Umesaki Y, Honda K (2011) Microbiotal influence on T cell subset development. Semin Immunol 23(2):146–153. https://doi.org/10.1016/j.smim.2011.01.010
Kim YS, Ho SB (2010) Intestinal goblet cells and mucins in health and disease: recent insights and progress. Curr Gastroenterol Rep 12(5):319–330. https://doi.org/10.1007/s11894-010-0131-2
Kim YS, Milner JA (2007) Dietary modulation of colon cancer risk. J Nutr 137(11 Suppl):2576S-2579S. https://doi.org/10.1093/jn/137.11.2576S
Stephani J, Radulovic K, Niess JH (2011) Gut microbiota, probiotics and inflammatory bowel disease. Arch Immunol Ther Exp 59(3):161–177. https://doi.org/10.1007/s00005-011-0122-5
Acknowledgements
This work was supported by the Junta de Andalucía (CTS 164) and by the Spanish Ministry of Economy and Competitiveness (SAF2011-29648 and AGL2015-67995-C3-3-R) with funds from the European Union. A. Rodriguez-Nogales is a postdoctoral fellow of Instituto de Salud Carlos III (Miguel Servet Program); R. Moron is a postdoctoral fellow of Instituto de Salud Carlos III (Rio Hortega Program); T. Vezza is a postdoctoral fellow from Instituto de Investigación Biosanitaria de Granada. The CIBER-EHD and the “Red de Investigación en SIDA” are funded by the Instituto de Salud Carlos III.
Author information
Authors and Affiliations
Contributions
Conceptualization: GG, AR-N and RM; Methodology: FA, JG-M, MJR-S, MER-C, MO and TV; Formal analysis and investigation: FA, JG-M, MJR-S, MER-C, MO and TV; Writing—original draft preparation: RM and AR-N; Writing—review and editing: TV, RM and AR-N; Funding acquisition: JG; Supervision: JG, RM and AR-N.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they do not have any competing interests. The funders had no role in the study design, data collection, and analysis.
Ethical approval
All studies were carried out following the ‘Guide for the Care and Use of Laboratory Animals’ as promulgated by the National Institute of Health and the protocols approved by the Ethic Committee of Laboratory Animals of the University of Granada (Spain) (Ref. No. CEEA-2010–286). They were therefore performed in accordance with the ethical standards laid down in the 1964 Declaration of Helsinki and its later amendments.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Algieri, F., Garrido-Mesa, J., Vezza, T. et al. Intestinal anti-inflammatory effects of probiotics in DNBS-colitis via modulation of gut microbiota and microRNAs. Eur J Nutr 60, 2537–2551 (2021). https://doi.org/10.1007/s00394-020-02441-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00394-020-02441-8