Abstract
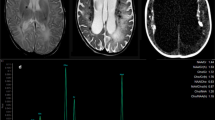
A novel multi-organ disease that is fatal in early childhood was identified in three patients from two non-consanguineous families. These children were born asymptomatic but at the age of 2 months they manifested progressive multi-organ symptoms resembling no previously known disease. The main clinical features included progressive cerebropulmonary symptoms, malabsorption, progressive growth failure, recurrent infections, chronic haemolytic anaemia and transient liver dysfunction. In the affected children, neuropathology revealed increased angiomatosis-like leptomeningeal, cortical and superficial white matter vascularisation and congestion, vacuolar degeneration and myelin loss in white matter, as well as neuronal degeneration. Interstitial fibrosis and previously undescribed granuloma-like lesions were observed in the lungs. Hepatomegaly, steatosis and collagen accumulation were detected in the liver. A whole-exome sequencing of the two unrelated families with the affected children revealed the transmission of two heterozygous variants in the NHL repeat-containing protein 2 (NHLRC2); an amino acid substitution p.Asp148Tyr and a frameshift 2-bp deletion p.Arg201GlyfsTer6. NHLRC2 is highly conserved and expressed in multiple organs and its function is unknown. It contains a thioredoxin-like domain; however, an insulin turbidity assay on human recombinant NHLRC2 showed no thioredoxin activity. In patient-derived fibroblasts, NHLRC2 levels were low, and only p.Asp148Tyr was expressed. Therefore, the allele with the frameshift deletion is likely non-functional. Development of the Nhlrc2 null mouse strain stalled before the morula stage. Morpholino knockdown of nhlrc2 in zebrafish embryos affected the integrity of cells in the midbrain region. This is the first description of a fatal, early-onset disease; we have named it FINCA disease based on the combination of pathological features that include fibrosis, neurodegeneration, and cerebral angiomatosis.






Similar content being viewed by others
References
Bancroft JD, Stevens A (1991) Theory and practice of histological techniques. Wiley, Churchill Livingstone, Edinburgh
Barabasi AL, Gulbahce N, Loscalzo J (2011) Network medicine: a network-based approach to human disease. Nat Rev Genet 12:56–68. https://doi.org/10.1038/nrg2918
Bossy-Wetzel E, Schwarzenbacher R, Lipton SA (2004) Molecular pathways to neurodegeneration. Nat Med 10(Suppl):S2–S9. https://doi.org/10.1038/nm1067
Diez-Roux G, Banfi S, Sultan M, Geffers L, Anand S, Rozado D et al (2011) A high-resolution anatomical atlas of the transcriptome in the mouse embryo. PLoS Biol 9:e1000582. https://doi.org/10.1371/journal.pbio.1000582
Edgar RC (2004) MUSCLE: multiple sequence alignment with high accuracy and high throughput. Nucleic Acids Res 32:1792–1797. https://doi.org/10.1093/nar/gkh340
Hamvas A, Deterding RR, Wert SE, White FV, Dishop MK, Alfano DN et al (2013) Heterogeneous pulmonary phenotypes associated with mutations in the thyroid transcription factor gene NKX2-1. Chest 144:794–804
Institute for Molecular Medicine Finland (FIMM), University of Helsinki, Finland (2016) Sequencing Initiative Suomi project (SISu). http://sisuproject.fi. Accessed 1/10 2017
Kaarteenaho-Wiik R, Sademies O, Paakko P, Risteli J, Soini Y (2007) Extracellular matrix proteins and myofibroblasts in granulomas of sarcoidosis, atypical mycobacteriosis, and tuberculosis of the lung. Hum Pathol 38:147–153
Karala AR, Ruddock LW (2010) Bacitracin is not a specific inhibitor of protein disulfide isomerase. FEBS J 277:2454–2462. https://doi.org/10.1111/j.1742-4658.2010.07660.x
Karjalainen MK, Huusko JM, Ulvila J, Sotkasiira J, Luukkonen A, Teramo K et al (2012) A potential novel spontaneous preterm birth gene, AR, identified by linkage and association analysis of X chromosomal markers. PLoS ONE 7:e51378. https://doi.org/10.1371/journal.pone.0051378
Kimmel CB, Ballard WW, Kimmel SR, Ullmann B, Schilling TF (1995) Stages of embryonic development of the zebrafish. Dev Dyn 203:253–310. https://doi.org/10.1002/aja.1002030302
Kleiner-Fisman G, Rogaeva E, Halliday W, Houle S, Kawarai T, Sato C et al (2003) Benign hereditary chorea: clinical, genetic, and pathological findings. Ann Neurol 54:244–247. https://doi.org/10.1002/ana.10637
Lek M, Karczewski KJ, Minikel EV, Samocha KE, Banks E, Fennell T et al (2016) Analysis of protein-coding genetic variation in 60,706 humans. Nature 536:285–291. https://doi.org/10.1038/nature19057
Long J, Pan G, Ifeachor E, Belshaw R, Li X (2016) Discovery of novel biomarkers for Alzheimer’s disease from blood. Dis Mark 2016:4250480. https://doi.org/10.1155/2016/4250480
Martin JL (1995) Thioredoxin–a fold for all reasons. Structure 3:245–250
Miller JA, Ding SL, Sunkin SM, Smith KA, Ng L, Szafer A et al (2014) Transcriptional landscape of the prenatal human brain. Nature 508:199–206. https://doi.org/10.1038/nature13185
Mitchell A, Chang HY, Daugherty L, Fraser M, Hunter S, Lopez R et al (2015) The InterPro protein families database: the classification resource after 15 years. Nucleic Acids Res 43:D213–D221. https://doi.org/10.1093/nar/gku1243
Nagy A, Gertsenstein M, Vintersten K, Behringer R (2003) Manipulating the mouse embryo, a laboratory manual, 3rd edn. Cold Spring Harbor Laboratory Press, New York, pp 198–200
Patel NJ, Jankovic J (2014) NKX2-1-related disorders. In: Adam MP, Ardinger HH, Pagon RA, Wallace SE, Bean LJH, Mefford HC et al (eds) GeneReviews(R). University of Washington, Seattle. GeneReviews is a registered trademark of the University of Washington, Seattle. All rights reserved, Seattle (WA)
Robu ME, Larson JD, Nasevicius A, Beiraghi S, Brenner C, Farber SA et al (2007) P53 activation by knockdown technologies. PLoS Genet 3:e78
Santiago JA, Potashkin JA (2013) Integrative network analysis unveils convergent molecular pathways in Parkinson’s disease and diabetes. PLoS ONE 8:e83940. https://doi.org/10.1371/journal.pone.0083940
Scavizzi F, Ryder E, Newman S, Raspa M, Gleeson D, Wardle-Jones H et al (2015) Blastocyst genotyping for quality control of mouse mutant archives: an ethical and economical approach. Transgenic Res 24:921–927. https://doi.org/10.1007/s11248-015-9897-1
Slack F, Ruvkun G (1998) Heterochronic genes in development and evolution. Biol Bull 195:375–376. https://doi.org/10.2307/1543152
Spagnolo P, Grunewald J, du Bois RM (2014) Genetic determinants of pulmonary fibrosis: evolving concepts. Lancet Respir Med 2:416–428. https://doi.org/10.1016/S2213-2600(14)70047-5
Sulonen AM, Ellonen P, Almusa H, Lepisto M, Eldfors S, Hannula S et al (2011) Comparison of solution-based exome capture methods for next generation sequencing. Genome Biol 12:R94. https://doi.org/10.1186/gb-2011-12-9-r94
Testa G, Schaft J, van der Hoeven F, Glaser S, Anastassiadis K, Zhang Y et al (2004) A reliable lacZ expression reporter cassette for multipurpose, knockout-first alleles. Genesis 38:151–158. https://doi.org/10.1002/gene.20012
Thorwarth A, Schnittert-Hubener S, Schrumpf P, Muller I, Jyrch S, Dame C et al (2014) Comprehensive genotyping and clinical characterisation reveal 27 novel NKX2-1 mutations and expand the phenotypic spectrum. J Med Genet 51:375–387. https://doi.org/10.1136/jmedgenet-2013-102248
Vakonakis I, Klewer DA, Williams SB, Golden SS, LiWang AC (2004) Structure of the N-terminal domain of the circadian clock-associated histidine kinase SasA. J Mol Biol 342:9–17. https://doi.org/10.1016/j.jmb.2004.07.010
Varilo T, Savukoski M, Norio R, Santavuori P, Peltonen L, Jarvela I (1996) The age of human mutation: genealogical and linkage disequilibrium analysis of the CLN5 mutation in the Finnish population. Am J Hum Genet 58:506–512
de Vries BB, Arts WF, Breedveld GJ, Hoogeboom JJ, Niermeijer MF, Heutink P (2000) Benign hereditary chorea of early onset maps to chromosome 14q. Am J Hum Genet 66:136–142
Witkowski L, Carrot-Zhang J, Albrecht S, Fahiminiya S, Hamel N, Tomiak E et al (2014) Germline and somatic SMARCA4 mutations characterize small cell carcinoma of the ovary, hypercalcemic type. Nat Genet 46:438–443. https://doi.org/10.1038/ng.2931
Zhang Y, Chen K, Sloan SA, Bennett ML, Scholze AR, O’Keeffe S et al (2014) An RNA-sequencing transcriptome and splicing database of glia, neurons, and vascular cells of the cerebral cortex. J Neurosci 34:11929–11947. https://doi.org/10.1523/JNEUROSCI.1860-14.2014
Acknowledgements
The authors would like to thank Professors Eric Shoubridge, Kalervo Hiltunen and Christer Betsholtz, Assistant Professor Michael Vanlandewijck, Adjunct Professor Siri Lehtonen and Dr. Riikka Pietilä for their expert advice and support and also Ms. Pirjo Keränen, Ms. Riitta Vuento, Ms. Maarit Haarala, Ms. Hanna Seppälä, Ms. Kirsi Säkkinen, the Transgenic Core Facility at Biocenter Oulu, and the Laboratory Animal Centre at the University of Oulu for their expert assistance. Biocenter Oulu Electron Microscopy core facility, a member of Biocenter Finland, is acknowledged for their help with EM analysis. The zebrafish work was carried out at University of Tampere core facility, supported by Biocenter Finland. The digital pathology scanner of Northern Finland Biobank Borealis was used in imaging the neuropathological findings. This work was conducted with support from the Research Council for Health of the Academy of Finland (JU, decision number 138566; RH, decision numbers 266498, 273790 and 303996; MH, decision number 1126662; LR, decision numbers 266457 and 272573); the Sigrid Juselius Foundation (JU, RH and MH); the Foundation for Paediatric Research, Finland (JU and MKK); the Alma and KA Snellman Foundation (JU and MKK); a Marie Curie International Outgoing Fellowship of the European Union’s Seventh Framework Programme (Grant agreement number 273669 [BioMit]) (RH); Foundation of the Finnish Anti-Tuberculosis Association (RK); the Jane and Aatos Erkko Foundation (MR); the Competitive State Research Financing of the Expert Responsibility Area of Tampere University Hospital (MR); Special State Grants for Health Research in the Department of Paediatrics and Adolescence at Oulu University Hospital, Finland (JU); the National Heart, Lung and Blood Institute of the US National Institutes of Health under award number HL-54703 (LMN) and the Eudowood Foundation (LMN).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Ethical approval
All procedures performed in the studies involving human participants were in accordance with the ethical standards of the institutional and national research committee and with the 1964 Helsinki Declaration and its later amendments or comparable ethical standards. Prior to the study, the guardians of the patients gave written informed consent to participate in the studies, and this was approved by the Ethics Committee of Oulu University Hospital (EETTMK 51/2008). Furthermore, the guardians of the patients in this manuscript have given written informed consent for the publication of their case details. All procedures performed in the studies involving animals were in accordance with the ethical standards of the institution or practice at which the studies were conducted. The National Animal Experiment Board of Finland approved the study protocol (ESAVI/5882/04.10.07/2014). Animal care and experimental procedures were conducted in accordance with the national legislation and EU Directive 2010/63/EU. Zebrafish housing and maintenance were done according to facility permission ESAVI/10079/04.10.06/2015. All applicable international, national and/or institutional guidelines for the care and use of animals were followed.
Conflict of interest
The authors declare that they have no conflict of interest.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Uusimaa, J., Kaarteenaho, R., Paakkola, T. et al. NHLRC2 variants identified in patients with fibrosis, neurodegeneration, and cerebral angiomatosis (FINCA): characterisation of a novel cerebropulmonary disease. Acta Neuropathol 135, 727–742 (2018). https://doi.org/10.1007/s00401-018-1817-z
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00401-018-1817-z