Abstract
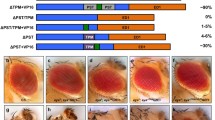
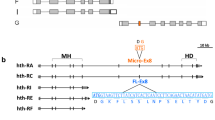
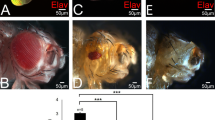
Physical and functional characteristics of the RUNX family of transcription factors are conserved between vertebrates and the Drosophila protein Lozenge. The runt-homology domain responsible for DNA binding and also the C-terminus are both nearly identical between the two proteins. The mammalian and fly proteins heterodimerize with a non-DNA binding partner protein to form a core binding factor essential for gene regulation during cell differentiation. The mammalian protein RUNX1 (AML1/PEBP2αB) interacts with the transcription factor Ets-1 to increase DNA binding and transactivation potential. Alternative splicing of the mammalian RUNX1 removes a domain required for this cooperative transactivation. In this work we determine the structure of the lozenge transcription unit and map 21 mutations. We show that the lozenge transcript is alternatively spliced during eye development to remove an Ets interaction domain. Emphasis is placed on Pointed the Drosophila homolog of the vertebrate Ets-1 protein; both Lozenge and Pointed proteins are needed for the activation of prospero expression. We use site-directed mutagenesis and yeast two-hybrid analysis to show that conserved amino acids within the alternate Lozenge exon are important for interaction with Pointed. Furthermore, the ectopic expression of Lozenge is sufficient to rescue Prospero expression in the presence of the Pointed competitor, YanACT. We show that both lozenge isoforms are expressed during eye development and that the relative ratio of the transcripts for the two isoforms is sensitive to changes in Ras activity. We suggest that during eye development, Lozenge isoforms function in divergent roles, either interacting with Pointed on downstream targets or by functioning independently to establish distinct cell fates.





Similar content being viewed by others
References
Albagli O, Klaes A, Ferreira E, Leprince D, Klambt C (1996) Function of ets genes is conserved between vertebrates and Drosophila. Mech Dev 59:29–40
Bae SC, Yamaguchiiwai Y, Ogawa E, Maruyama M, Inuzuka M, Kagoshima H, Shigesada K, Satake M, Ito Y (1993) Isolation of Pebp2-alpha-B cDNA representing the mouse homolog of human acute myeloid-leukemia gene, Aml1. Oncogene 8:809–814
Bae SC, Ogawa E, Maruyama M, Oka H, Satake M, Shigesada K, Jenkins NA, Gilbert DJ, Copeland NG, Ito Y (1994) Pebp2-alpha-B/mouse Aml1 consists of multiple isoforms that possess differential transactivation potentials. Mol Cell Biol 14:3242–3252
Batterham P, Crew JR, Sokac AM, Andrews JR, Pasquini GMF, Davies AG, Stocker R, Pollock JA (1996) Genetic analysis of the lozenge gene complex of Drosophila melanogaster: adult visual system phenotypes. J Neurogenet 10:193–220
Behan KJ, Nichols CD, Cheung TL, Farlow A, Hogan BM, Batterham P, Pollock JA (2002) Yan regulates Lozenge during Drosophila eye development. Dev Genes Evol 212:267–276
Berardi MJ, Sun CH, Zehr M, Abildgaard F, Peng J, Speck JN, Bushweller JH (1999) The Ig fold of the core binding factor alpha Runt domain is a member of a family of structurally and functionally related Ig-fold DNA-binding domains. Struct Fold Des 7:1247–1256
Bridges CB, Brehme KC (1944) The mutants of Drosophila melanogaster. Carnegie Inst Wash Publ 552:257
Brunner D, Ducker K, Oellers K, Hafen E, Scholz H, Klambt C (1994) The Ets domain protein pointed-P2 is a target of map kinase in the sevenless signal-transduction pathway. Nature 370:386–389
Canon J, Banerjee U (2000) Runt and Lozenge function in Drosophila development. Semin Cell Dev Biol 11:327–336
Crew JR, Batterham P, Pollock JA (1997) Developing compound eye in lozenge mutants of Drosophila: lozenge expression in the R7 equivalence group. Dev Genes Evol 206:481–493
Daga A, Karlovich CA, Dumstrei K, Banerjee U (1996) Patterning of cells in the Drosophila eye by lozenge, which shares homologous domains with AML1. Genes Dev 10:1194–1205
Dittmer J (2003) The biology of the Ets1 proto-oncogene. Mol Cancer 2(1):29
Erman B, Cortes M, Nikolajczyk BS, Speck NA, Sen R (1998) Ets-core binding factor: a common composite motif in antigen receptor gene enhancers. Mol Cell Biol 18:1322–1330
Feig LA, Cooper GM (1988) Inhibition of NIH-3T3 cell proliferation by a mutant Ras protein with preferential affinity for GDP. Mol Cell Biol 8:3235–3243
Fields S, Song O (1989) A novel genetic system to detect protein–protein interactions. Nature 340(6230):245–246
Flores GV, Daga A, Kalhor HR, Banerjee U (1998) Lozenge is expressed in pluripotent precursor cells and patterns multiple cell types in the Drosophila eye through the control of cell-specific transcription factors. Development 125:3681–3687
Flores GV, Duan H, Yan HJ, Nagaraj R, Fu WM, Zou Y, Noll M, Banerjee U (2000) Combinatorial signaling in the specification of unique cell fates. Cell 103:75–85
Ghozi MC, BernsteinY, Negreanu V, Levanon D, Groner Y (1996) Expression of the human acute myeloid leukemia gene AML1 is regulated by two promoter regions. Proc Natl Acad Sci U S A 93(5):1935–1940
Goetz TL, Gu TL, Speck NA, Graves BJ (2000) Auto-inhibition of Ets-1 is counteracted by DNA binding cooperativity with core-binding factor alpha2. Mol Cell Biol 20:81–90
Golling G, Li LH, Pepling M, Stebbins M, Gergen JP (1996) Drosophila homologs of the proto-oncogene product PEBP2/CBF beta regulate the DNA-binding properties of Runt. Mol Cell Biol 16:932–942
Goodrich JA, Schwartz ML, McClure WR (1990) Searching for and predicting the activity of sites for DNA-binding proteins—compilation and analysis of the binding-sites for Escherichia coli integration host factor (Ihf). Nucleic Acids Res 18:4993–5000
Green MM (1961) Phenogenetics of the lozenge loci in Drosophila melanogaster. II. Genetics of lozenge-krivshenko (lozengek). Genetics 46:1169–1176
Green MM (1990) The foundations of genetic fine structure: a retrospective from memory. Genetics 124:793–796
Green MM, Green KC (1949) Crossing over between alleles at the lozenge locus in Drosophila melanogaster. Proc Natl Acad Sci U S A 35:586–591
Green MM, Green KC (1956) A cytogenetic analysis of the lozenge psuedoalleles in Drosophila. Zeit Abs Verer 87:708–724
Gu TL, Goetz TL, Graves BJ, Speck NA (2000) Auto-inhibition and partner proteins, core-binding factor beta (CBFbeta) and Ets-1, modulate DNA binding by CBFalpha2 (AML1). Mol Cell Biol 20:91–103
Kalev-Zylinska ML, Horsfield JA, Flores MV, Postlethwait JH, Vitas MR, Baas AM, Crosier PS, Crosier KE (2002) RUNX1 is required for zebrafish blood and vessel development and expression of a human RUNX1-CBF2T1 transgene advances a model for studies of leukemogenesis. Development 129:2015–2030
Kaminker JS, Singh R, Lebestky T, Yan HJ, Banerjee U (2001) Redundant function of Runt domain binding partners, big brother and brother, during Drosophila development. Development 128:2639–2648
Kanno T, Kanno Y, Chen F, Ogawa E, Kim WY, Ito Y (1998) Intrinsic transcriptional activation-inhibition domains of the polyomavirus enhancer binding protein 2 core binding factor alpha subunit revealed in the presence of the beta subunit. Mol Cell Biol 18:2444–2454
Karim FD, Chang HC, Therrien M, Wassarman DA, Laverty T, Rubin GM (1996) A screen for genes that function downstream of Ras1 during Drosophila eye development. Genetics 143:315–329
Kauffmann RC, Li SH, Gallagher PA, Zhang JJ, Carthew RW (1996) Ras1 signaling and transcriptional competence in the R7 cell of Drosophila. Genes Dev 10:2167–2178
Kim WY, Sieweke M, Ogawa E, Wee HJ, Englmeier U, Graf T, Ito Y (1999) Mutual activation of Ets-1 and AML1 DNA binding by direct interaction of their autoinhibitory domains. EMBO J 18:1609–1620
Klambt C (1993) The Drosophila gene pointed encodes 2 Ets-like proteins which are involved in the development of the midline glial-cells. Development 117:163–176
Klein SL, Strausberg RL, Wagner L, Pontius J, Clifton SW, Richardson P (2002) Genetic and genomic tools for Xenopus research: the NIH Xenopus initiative. Dev Dyn 225(4):384–391
Kosman D, Small S, Reinitz J (1998) Rapid preparation of a panel of polyclonal antibodies to Drosophila segmentation proteins. Dev Genes Evol 208(5):290–294
Kozlova TY, Semeshin VF, Tretyakova IV, Kokoza EB, Pirrotta V, Grafodatskaya VE, Belyaeva ES, Zhimulev IF (1994) Molecular and cytogenetical characterization of the 10A1-2 band and adjoining region in the Drosophila melanogaster polytene X chromosome. Genetics 136:1063–1073
Lai ZC, Rubin GM (1992) Negative control of photoreceptor development in Drosophila by the product of the yan gene, an Ets domain protein. Cell 70:609–620
Lebestky T, Chang T, Hartenstein V, Banerjee U (2000) Specification of Drosophila hematopoietic lineage by conserved transcription factors. Science 288:146–149
Lee J, Ahnn J, Bae SC (2004) Homologs of RUNX and CBF beta/PEBP2 beta in C. elegans. Oncogene 23(24):4346–4352
Lefevre G (1976) A photographic representation and interpretation of the polytene chromosomes of Drosophila melanogaster salivary glands. In: Ashburner M, Novitsky E (eds) The genetics and biology of Drosophila, vol Ia. Academic, New York, pp 31–66
Li LH, Gergen JP (1999) Differential interactions between Brother proteins and Runt domain proteins in the Drosophila embryo and eye. Development 126:3313–3322
Matsuo T, Takahashi K, Kondo S, Kaibuchi K, Yamamoto D (1997) Regulation of cone cell formation by Canoe and Ras in the developing Drosophila eye. Development 124:2671–2680
Miyoshi H, Ohira M, Shimizu K, Mitani K, Hirai H, Imai T, Yokoyama K, Soeda E, Ohki M (1995) Alternative splicing and genomic structure of the Aml1 gene involved in acute myeloid-leukemia. Nucleic Acids Res 23:2762–2769
Morgan TH, Bridges CB, Sturtevant AH (1925) The genetics of Drosophila melanogaster. Biblphia Genet 2:1–262
Mount SM, Burks C, Hertz G, Stormo GD, White O, Fields C (1992) Splicing signals in Drosophila—intron size, information-content, and consensus sequences. Nucleic Acids Res 20:4255–4262
Nichols CD (1997) Molecular characterization of the lozenge locus of Drosophila melanogaster 1997. Department of Biological Sciences, Carnegie Mellon University, Pittsburgh PA
Oliver CP (1940) A reversion to wild type associated with crossing over in Drosophila melanogaster. Proc Natl Acad Sci U S A, 26:452–454
Pires-daSilva A, Sommer RJ (2003) The evolution of signalling pathways in animal development. Nat Rev Genet 4(1):39–49
Rebay I, Rubin GM (1995) Yan functions as a general inhibitor of differentiation and is negatively regulated by activation of the Ras1/MAPK pathway. Cell 81:857–866
Rizki TM, Rizki RM (1981) Alleles of lozenge as suppressors of the Bc-phene in Drosophila melanogaster. Genetics (Suppl.) 97:s90
Sambrook J, Fritsch EF, Maniatis T (1989) Molecular cloning: a laboratory manual. Cold Spring Harbor Laboratory Press, New York
Siddall N, Behan KJ, Crew JR, Cheung TL, Fair JA, Batterham P, Pollock JA (2003) Mutations in lozenge and D-Pax2 invoke ectopic patterned cell death in the developing Drosophila eye using distinct mechanisms. Dev Genes Evol 213:107–119
Spana EP, Doe CQ (1995) The Prospero transcription factor is asymmetrically localized to the cell cortex during neuroblast mitosis in Drosophila. Development 121:3187–3195
Tenen DG (2003) Disruption of differentiation in human cancer: AML shows the way. Nature Rev 3:89–101
Tracey WD, Pepling ME, Horb ME, Thomsen GH, Gergen JP (1998) A Xenopus homologue of aml-1 reveals unexpected patterning mechanisms leading to the formation of embryonic blood. Development 125:1371–1380
Wang H, McIntosh LP, Graves BJ (2002) Inhibitory module of Ets-1 allosterically regulates DNA binding through a dipole-facilitated phosphate contact. J Biol Chem 277:2225–2233
Wotton D, Ghysdael J, Wang SW, Speck NA, Owen MJ (1994) Cooperative binding of Ets-1 and Core Binding Factor to DNA. Mol Cell Biol 14:840–850
Xu CY, Kauffmann RC, Zhang JJ, Kladny S, Carthew RW (2000) Overlapping activators and repressors delimit transcriptional response to receptor tyrosine kinase signals in the Drosophila eye. Cell 103:87–97
Zhu X, Yeadon JE, Burden SJ (1994) AML1 is expressed in skeletal muscle and is regulated by innervation. Mol Cell Biol 14 (12):8051–8057
Acknowledgements
We thank Jana Van Patton and Claudia Almaguer for help with the Yeast Two-Hybrid system. We thank Ranga Rao and Joe Lepo for the use of their lab at UWF. We thank A. Javier Lopez, Richard Carthew and Andreas Nocker for discussion and advice. We thank Raghuram Selvaraju for critical review of the manuscript. We thank Sam Saba and Paul Keller for their help in optimizing PCR conditions, and James M. Burnette for assisting with MacTargSearch analysis. We thank Utpal Banerjee for flies and for the lozenge 3.5 clone, Gerald Rubin for the gift of flies, Richard Carthew for the PntP2 clone, and Chris Doe for the gift of Prospero antibody. Parts of this work were performed at Carnegie Mellon University and the University of West Florida. This work was supported in part by Duquesne University of the Holy Ghost and the following grants to J.A.P.: SURG/HHMI Awards to undergraduate researchers at Carnegie Mellon University; NIH grant EY09093; NSF/STC grant BIR-8920118; March of Dimes Birth Defects Foundation FY93-1010; Duquesne University Faculty Development Awards; Samuel and Emma Winters Foundation; and international collaborative support from the NSF INT-9605205 and the University of Melbourne.
Author information
Authors and Affiliations
Corresponding author
Additional information
Communicated by C. Desplan
K.J. Behan and J. Fair contributed equally to the data presented
Electronic Supplementary Material
Rights and permissions
About this article
Cite this article
Jackson Behan, K., Fair, J., Singh, S. et al. Alternative splicing removes an Ets interaction domain from Lozenge during Drosophila eye development. Dev Genes Evol 215, 423–435 (2005). https://doi.org/10.1007/s00427-005-0490-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00427-005-0490-0