Abstract
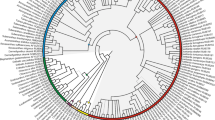
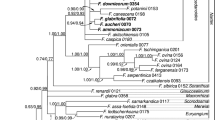
The phylogeny of Menispermaceae (Ranunculales) is reconstructed here using both morphological and molecular data from a broad sample of species. Morphological data include characters of leaves, wood, flowers, fruits, seeds, pollen and phytochemistry from 73 species representing the different subgroups recognized within the family. This dataset allowed us to study the morphological evolutionary trends in Menispermaceae. The molecular data focused on cpDNA sequences: rbcL and atpB. Maximum parsimony (MP) and Bayesian methods were used to reconstruct the phylogeny of Menispermaceae. The results obtained from the three datasets are partly incongruent. The morphology indicates that Menispermaceae can be divided into two major groups, one including the tribes Fibraureae and Tinosporeae, the other one including the tribes Anomospermeae, Pachygoneae and Menispermeae. Only Burasaia Thouars was not included in any of these groups. The characters of fruits and seeds appeared to be most useful to differentiate these groups whereas convergences were found for the androecium.






Similar content being viewed by others
References
Anderson CL, Bremer K, Friis EM (2005) Dating phylogenetically basal eudicots using rbcL sequences and multiple fossil reference points. Amer J Bot 92:1737–1748
APG II (2003) An update of the angiosperm phylogeny group classification for the orders and families of flowering plants: APG II. Bot J Linn Soc 141:399–436
Bhambie S (1972) Correlation between form, structure and habit in some lianas. Proc Indian Acad Sci B 75:246–256
Barbosa-Filho JM, Leitão da-Cunha EV, Gray AI (2000) Alkaloids of the Menispermaceae. The Alkaloids 54:1–190
Barneby RC (1970) Revision of neotropical Menispermaceae tribe Tinosporeae. Mem New York Bot Gard 20:81–158
Barneby RC (1972) New and notable Menispermaceae tribe Tinosporeae. Mem New York Bot Gard 22:137–151
Bremer K (1994) Branch support and tree stability. Cladistics 10:295–304
Chaw S-M, Parkinson CL, Cheng Y, Vincent TM, Palmer JD (2000) Seed plant phylogeny inferred from all three plant genomes: monophyly of extant Gymnosperms and origin of Gnetales from conifers. Proc Natl Acad Sci 97:4086–4091
Cummings MP, Otto SP, Wakeley J (1995) Sampling properties of DNA sequence data in phylogenetic analysis. Molec Biol Evol 12:814–822
Damerval C, Nadot S (2007) Evolution of perianth and stamen characteristics with respect to floral symmetry in Ranunculales. Ann Bot 100:631–640
Djauberg N, Andersen RA (1997) Phylogenetic analyses of the rbcL sequences of haptophytes and heterokont Algae suggest their chloroplasts are unrelated. Mol Biol Evol 14:1242–1251
Dekker AJFM (1983) A revision of the genera Penianthus Miers and Sphenocentrum Pierre (Menispermaceae) of West and Central Africa. Bull Jard Bot Natl Belg 53:17–66
Deleporte P (1993) Characters, attributes and tests of evolutionary scenarios. Cladistics 9:427–432
DePinna MCC (1991) Concepts and test of homology in the cladistic paradigm. Cladistics 7:367–394
Diels L (1910) Menispermaceae. In: Engler A (ed) Das Pflanzenreich IV, 94:1–345
Felsenstein J (1982) Numerical methods for inferring evolutionary trees. Quart Rev Biol 57:379–404
Felsenstein J (1985) Confidence limits on phylogenies: an approach using the bootstrap. Evolution 39:783–791
Ferguson IK (1975) Pollen morphology of the tribe Triclisieae of the Menispermaceae in relation to its taxonomy. Kew Bull 30:49–85
Ferguson IK (1978) Pollen morphology of the tribe Coscinieae of the Menipsermaceae in relation to its taxonomy. Kew Bull 32:339–349
Forman LL (1974) The endocarps of Cocculus (Menispermaceae). Kew Bull 29:477–481
Forman LL (1985) A revision of the tribe Fibraureae (Menispermaceae) in Asia. Kew Bull 40:539–551
Givnish TJ, Vermeij GJ (1976) Sizes and shapes of liana leaves. Amer Naturalist 110:743–778
Goldman N, Yang Z (1994) A codon-based model of nucleotide substitution for protein coding DNA sequences. Molec Biol Evol 11:725–736
Grandcolas P, Deleporte P, Desutter-Grandcolas L, Daugeron C (2001) Phylogenetics and ecology: as many characters as possible should be included in cladistic analysis. Clasdistics 17:104–110
Harley MM (1985) Pollen morphology and taxonomy of the tribe Fibraureae (Menispermaceae). Kew Bull 40:553–561
Harley MM, Ferguson IK (1982) Pollen morphology and taxonomy of the tribe Menispermeae (Menispermaceae). Kew Bull 37:353–366
Hickey LJ, Wolfe JA (1975) The bases of Angiosperm phylogeny: vegetative morphology. Ann Missouri Bot Gard 62:538–589
Hillis DM (1996) Inferring complex phylogenies. Nature 383:130–131
Hong Y-P, Chen Z-D, Lu A-M (2001a) Phylogeny of the tribe Menispermeae (Menispermaceae) reconstructed by ITS sequence data. Acta Phytotaxonomica Sinica 39:97–104 (with English summary)
Hong Y-P, Pan K-Y, Chen Z-D, Lu A-M (2001b) Characters of leaf epidermis and their systematic significance in Menispermaceae. Acta Bot Sin 43:615–623 (with English summary.)
Hoot SB, Culham A, Crane PR (1995) The utility of atpB gene sequences in resolving phylogenetic relationships: comparisons with rbcL and 18S ribosomal DNA sequences in the Lardizabalaceae. Ann Missouri Bot Gard 82:194–207
Hoot SB, Megallon-Puebla S, Crane PR (1999) Phylogeny of basal eudicots based on three molecular data sets: atpB, rbcL, and 18S nuclear ribosomal DNA sequences. Ann Missouri Bot Gard 86:1–32
Huelsenbeck JP, Ronquist F (2001) MRBAYES: Bayesian inference of phylogeny. Bioinformatics 17:754–755
Jacques FMB, De Franceschi D (2005) Endocarps of Menispermaceae from Le Quesnoy outcrop (Sparnacian facies, Lower Eocene, Paris Basin). Rev Palaebot Palynol 135:61–70
Jacques FMB, De Franceschi D (2007) Menispermaceae wood anatomy and cambial variants. IAWA J 28:139–172
Jacques FMB, Gallut C, Vignes-Lebbe R, Zaragüeta I, Bagils R (2007) Resolving phylogenetic reconstruction in Menispermaceae (Ranunculales) using fossils and a novel statistical test. Taxon 56:379–392
Källersjö M, Farris JS, Chase MW, Bremer B, Fay MF, Humphries CJ, Petersen G, Seberg O, Bremer K (1998) Simultaneous parsimony jackknife analysis of 2538 rbcL DNA sequences reveals support for major clades of green plants, land plants, seed plants and flowering plants. Pl Syst Evol 213:259–287
Kessler PJA (1993) Menispermaceae. In: Kubitzki K, Rohwer JG, Bittrich V (eds) The families and genera of vascular plants, 2. Springer, Berlin, pp 402–418
Knobloch E, Mai DH (1986) Monographie der Früchte und Samen in der Kreide Mitteleuropa. Rozpr Ústred úst geol Praha 47:1–279
Kopp A, True JR (2002) Phylogeny of the oriental Drosophila melanogaster species group: a multilocus reconstruction. Syst Biol 51:786–805
Lewis LA, Mishler BD, Vilgalys R (1997) Phylogenetic relationships of the liverworts (Hepaticeae), a basal embryophyte lineage, inferred from sequence nucleotide data of the chloroplast gene rbcL. Molec Phylogenet Evol 7:377–393
Lockart PJ, Larkum AWD, Steel MA, Waddell PJ, Penny D (1996) Evolution of chlorophyll and bacteriochlorophyll: the problem of invariant sites in sequence analysis. Proc Natl Acad Sci USA 93:1930–1934
Krukoff BA, Moldenke HN (1938) Studies of American Menispermaceae, with special reference to species used in preparation of arrow-poisons. Brittonia 3:1–74
Maddison WP, Maddison DR (2006) Mesquite: a modular system for evolutionary analysis. Version 1.1. http://mesquiteproject.org
Manchester SR (1994) Fruits seeds of Middle Eocene Nut Beds Flora, Clarno formation Oregon. Palaeontogr Am 58:1–205
Manhart JR (1994) Phylogenetic analysis of green plant rbcL sequences. Molec Phylogenet Evol 3:114–127
McDade LA (1995) Hybridization and phylogenetics. In: Hoch PC, Stephenson AG (eds) Experimental and molecular approaches to plant biosystematics. Missouri Botanical Garden, St Louis, pp 305–331
Mennega AMW (1982) Stem structure of the new world Menispermaceae. J Arnold Arbor 63:145–171
Miers J (1871) A complete monograph of the Menispermaceae. Contribution to Botany 3. Williams and Norgate, London
Muasya AM, Simpson DA, Chase MW, Culham A (1998) An assessment of suprageneric phylogeny in Cyperaceae using rbcL DNA sequences. Pl Syst Evol 211:257–271
Muse S, Gaut B (1994) A likelihood approach for comparing synonymous and non-synonymous substitution rates, with application to the chloroplast genome. Molec Biol Evol 11:715–724
Nickrent DL, Parkinson CL, Palmer JD, Duff RJ (2000) Multigene phylogeny of Land Plants with special reference to Bryophytes and the earliest land plants. Molec Biol Evol 17:1885–1895
Nixon KC (1999) The Parsimony Ratchet, a new method for rapid parsimony analysis. Cladistics 15:407–414
Obaton M (1960) Les lianes ligneuses à structure anormale des forêts denses d’Afrique occidentale. Ann Sci Nat Bot Biol Vég 1:1–220
Olmstead RG, Michaels HJ, Scott KM, Palmer JD (1992) Monophyly of the Asteridae and identification of their major lineages inferred from DNA sequences of rbcL. Ann Missouri Bot Gard 79:249–265
Ortiz RDC, Kellogg EA, Van der Werff H (2007) Molecular phylogeny of the moonseed family (Menispermaceae): implications for morphological diversification. Amer J Bot 94:1425–1438
Pimentel RA, Riggins R (1987) The nature of cladistics data. Cladistics 3:201–209
Pisani D, Benton MJ, Wilkinson M (2007) Congruence of morphological and molecular phylogenies. Acta Biotheor 55:269–281
Pollock DD, Zwickl DJ, McGuire JA, Hillis DM (2002) Increased taxon sampling is advantageous for phylogenetic inference. Syst Biol 51:664–671
Rannala B, Huelsenbeck JP, Yang Z, Nielsen R (1998) Taxon sampling and the accuracy of large phylogenies. Syt Biol 47:702–710
Rambaut A (1996) Se-Al: Sequence Alignment Editor. http://tree.bio.ed.ac.uk/software/seal/
Reid EM, Chandler MEJ (1933) The London clay. The British Museum (Natural History), London, p 561
Rieseberg LH, Soltis DE (1991) Phylogenetic consequences of cytoplasmic gene flow in plants. Evol Trends Plants 5:65–84
Rokas A, Carroll SB (2005) More genes or more taxa? The relative contribution of gene number and taxon number to phylogenetic accuracy. Molec Biol Evol 22:1337–1344
Rokas A, Williams BL, King N, Carroll SB (2003) Genome-scale approaches to resolving incongruence in molecular phylogenies. Nature 425:798–804
Ronquist F, Huelsenbeck JP (2003) MRBAYES 3: Bayesian inference under mixed models. Bioinformatics 19:1572–1574
Rosenberg MS, Kumar S (2001) Incomplete taxon sampling is not a problem for phylogenetic inference. Proc Natl Acad Sci 98:10751–10756
Sang T, Zhong Y (2000) Testing hybridization hypotheses based on incongruent gene trees. Syst Biol 49:422–434
Satta Y, Klein J, Takahata N (2000) DNA archives and our nearest relative: the trichotomy problem revisited. Molec Phylogent Evol 14:259–275
Savolainen V, Chase MW (2003) A decade of progress in plant molecular phylogenetics. Trends Genet 19:717–724
Savolainen V, Chase MW, Hoot SB, Morton CM, Soltis DE, Bayer C, Fay MF, De Bruijn AY, Sullivan S, Qiu Y-L (2000) Phylogenetics of flowering plants based on combined analysis of plastid atpB and rbcL gene sequences. Syst Biol 49:306–362
Schenk H (1893) Beiträge zur Anatomie der Lianen. II. Botanische Mittheilungen aus den Tropen 5:1–271
Sikes DS, Lewis PO (2001) Beta software, version 1. PAUPRat: PAUP* implementation of the parsimony ratchet. Distributed by the authors. Department of Ecology and Evolutionary Biology, University of Connecticut, Storrs
Simmons MP, Reeves A, Davis JI (2004) Character space versus rate of evolution in phylogenetic inference. Cladistics 20:191–204
Steel M, Huson D, Lockhart PJ (2000) Invariable sites models and their use in phylogeny reconstruction. Syst Biol 49:225–232
Strong EE, Lipscomb D (1999) Character coding and inapplicable data. Cladistics 15:363–371
Sugiyama M, Hara N (1988) Comparative study of early ontogeny of compound leaves in Lardizabalaceae. Amer J Bot 75:1598–1605
Swofford DL (1998) PAUP*. Phylogenetic analysis using Parsimony (* and other methods). Version 4. Sinauer Associates, Sunderland
Takhtajan A (1974) Magnolophyta fossilia URSS I. Magnoliaceae–Eucommiaceae. Nauka, Leninopoli, p 188; 124 pl
Tavaré S (1986) Some probabilistic and statistical problems on the analysis of DNA sequences. Lect Math Life Sci 17:57–86
Thanikaimoni G (1968) Morphologie des pollens des Ménispermacées. Institut français de Pondichéry, Pondicherry, p 57
Thanikaimoni G (1984) Ménispermacées: palynologie et systématique. Institut français de Pondichéry, Pondicherry, p 135
Troupin G (1962) Monographie des Menispermaceae africaines. Mémoires—Acad. R Sci o-m Cl Sci Nat et Méd Collect 8 13:1–313
Xia X (1998) The rate heterogeneity of nonsynonymous substitutions in mammalian mitochondrial genes. Molec Biol Evol 15:336–344
Xia X, Hafner MS, Sudman PD (1996) On transition bias in mitochondrial genes of pocket gophers. J Molec Evol 43:32–40
Wang W, Wang H-C, Chen Z-D (2007) Phylogeny and morphological evolution of tribe Menispermeae (Menispermaceae) inferred from chloroplast and nuclear sequences. Perspect Pl Ecol Evol Syst 8:141–154
Zwickl DJ, Hillis DM (2002) Increased taxon sampling greatly reduces phylogenetic error. Syst Biol 51:588–598
Acknowledgments
We thank the Bogor Botanic Garden, the Kepong Forest Reasearch Institute of Malaysia, the Queen Sirikit Botanical Garden and the National Botanic Garden of Belgium for their assistance in specimen collection. We thank Sophie Nadot, Dario De Franceschi and two anonymous reviewers for useful comments on former versions. This study was partly funded by LVMH.
Author information
Authors and Affiliations
Corresponding author
Appendix 1: character coding
Appendix 1: character coding
Habit. 1. Port: 0 = liana (all climbing habit), 1 = small tree. Stem. 2. Tubercle: 0 = absent, 1 = present; 3. Young stem pubescent: 0 = no, 1 = yes; 4. Long trichomes longs: 0 = no, 1 = yes; 5. Old stems pubescent: 0 = no, 1 = yes; 6. Old bark verrucose: 0 = no, 1 = yes; 7. Old bark lenticellate: 0 = no, 1 = yes; 8. Old bark parcheminate: 0 = no, 1 = yes; Leaf. 9. Petiole insertion cupular: 0 = no, 1 = yes; 10. Petiole pubescent: 0 = no, 1 = yes; 11. Petiole striation: 0 = no, 1 = yes; 12. Petiole sulcate: 0 = no, 1 = yes; 13. Petiole enlarged at base: 0 = no, 1 = yes; 14. Petiole swollen at base: 0 = no, 1 = yes; 15. Petiole geniculate at base: 0 = no, 1 = yes; 16. Petiole swollen at apex: 0 = no, 1 = yes; 17. Petiole geniculate at apex: 0 = no, 1 = yes; 18. Petiole length: 0 = <leaf length, 1 = ≥leaf length; 19. Compound leaf: 0 = no, 1 = yes; 20. Lobed leaf: 0 = no, 1 = yes; 21. Lamina shape: 0 = length/width < 1, 1 = length/width ≥1; 22. Leaf acuminate: 0 = no, 1 = yes; 23. Leaf apex mucronate: 0 = no, 1 = yes; 24. Lamina base: 0 = convex, 1 = straight, 2 = concave; 25. Leaf peltate: 0 = no, 1 = yes, petiole insertion < 10% of leaf length, 2 = yes, petiole insertion ≥ 10% of leaf length; 26. Venation type: 0 = palmate, 1 = pinnate; 27. If leaf pinnate, distal venation pinnate: 0 = no, 1 = yes; 28. Number of basal veins: 0 = 3, 1 = 5, 2 = >5; 29. Veins suprabasal: 0 = no, 1 = yes; 30. Tertiary external veins from the lateral basal veins: 0 = no, 1 = yes; 31. Veins fusing with margin: 0 = no, 1 = yes; 32. Margin type: 0 = entire, 1 = toothed, 2 = sinuous; 33. Veins prominent on abaxial face: 0 = no, 1 = yes; 34. Upper face pubescent: 0 = no, 1 = yes (even only few hairs); 35. Location of pubescence on upper face: 0 = everywhere, 1 = only on veins; 36. Lower face pubescent: 0 = no, 1 = yes (even only few hairs); 37. Location of pubescence on lower face: 0 = everywhere, 1 = only on veins; 38. Domatia: 0 = no, 1 = yes; 39. Leaf texture: 0 = membranous, 1 = chartaceous, 2 = coriaceous; 40. Shape of upper epidermis cells: 0 = polygonal, 1 = irregular; 41. Anticlinal wall of those cells: 0 = straight, 1 = sinuous, 2 = sinuolate; 42. Shape of lower epidermis cells: 0 = polygonal, 1 = irregular; 43. Anticlinal wall of those cells: 0 = straight, 1 = sinuous, 2 = sinuolate; 44. Stomata type: 0 = anomocytic, 1 = staurocytic, 2 = cyclocytic, 3 = actinocytic. Wood. 45. Wood with successive cambia: 0 = no, 1 = yes; 46. Wood rays lignified: 0 = no, 1 = yes. Inflorescence. The highly variability in inflorescence between species does not allow a simple coding of inflorescence type, the inflorescences characteristics were therefore separated in several characters. 47. Flower in inflorescence: 0 = no, 1 = yes; 48. Composition degree of inflorescence: 0 = 1, 1 = 2, 2 = >2; 49. Type of highest level of inflorescence: 0 = raceme, 1 = cyme; 50. Type of lowest level of inflorescence: 0 = raceme, 1 = cyme; 51. Inflorescences fasciculate: 0 = no, 1 = yes; 52. Inflorescence position: 0 = axillary, 1 = ramiflorous, 2 = terminal; 53. Inflorescence on old wood: 0 = no, 1 = yes; 54. Pauciflorous inflorescence: 0 = no, 1 = yes; 55. Inflorescence pubescent: 0 = no, 1 = yes; 56. Inflorescence length: 0 = <leaf length, 1 = ≥leaf length. Flower. As Menispermaceae species are all dioic and male and female flowers as sometimes different structures, all male and female characters were handled separately. 57. Flower type: 0 = hermaphrodite, 1 = monoecious, 2 = dioecious; 58. Merosity: 0 = 3, 1 = 2, 2 = 5. 59. Sepal number in male flower relatively to merosity: 0 = >3, 1 = 1, 2 = 2, 3 = 3; 60. Sepal number in female flower relatively to merosity: 0 = >3, 1 = 1, 2 = 2, 3 = 3; 61. Outer sepal pubescent: 0 = no, 1 = yes; 62. Inner sepals pubescent on outer face: 0 = no, 1 = yes; 63. Relative size of inner sepals: 0 = small, 1 = equal, 2 = large; 64. Sepals in tube: 0 = no, 1 = yes; 65. Sepals fleshy: 0 = no, 1 = yes; 66. Sepal type: 0 = valvate, 1 = imbricate; 67. Petal number in male flower relatively to merosity: 0 = 0, 1 = 1, 2 = 2, 3 = >3; 68. Petal number in female flower relatively to merosity: 0 = 0, 1 = 1, 2 = 2, 3 = ≥3, 4 = 1 petal; 69. Fused petal: 0 = no, 1 = yes; 70. Petal size relatively to inner sepals: 0 = minute, 1 = smaller than sepals, 2 = equal, 3 = larger. 71. Petals surrounding stamens: 0 = no, 1 = yes; 72. Petal texture: 0 = membranous, 1 = fleshy; 73. Stamen number relatively to merosity: 0 = >3, 1 = 1, 2 = 2, 3 = 3; 74. Stamen fusion: 0 = no, 1 = only inners, 2 = all; 75. Synandrium type: 0 = no synandrium, 1 = central column, vertical anthers, 2 = central column, horizontal anthers, 3 = stalked pyramid, 4 = sessile pyramid; 76. Connective extended over the anthers: 0 = no, 1 = yes; 77. Filament enlarging: 0 = no, 1 = yes; 78. Connective highly visible: 0 = no, 1 = yes; 79. Anther shape: 0 = globular, 1 = ellipsoid; 80. Orientation of outer anthers: 0 = introrse, 1 = extrorse; 81. Orientation of inner anthers: 0 = introrse, 1 = extrorse; 82. Orientation of dehiscence slit: 0 = transversal, 1 = longitudinal; 83. Pistillodes: 0 = no, 1 = yes. 84. Hairs in the center of male flower: 0 = no, 1 = yes; 85. Staminodes number relatively to merosity: 0 = 0, 1 = 1, 2 = 2, 3 = ≥3. 86. Anthers on the staminodes: 0 = no, 1 = yes. 87. Carpel number relatively to merosity: 0 = >2, 1 = 1, 2 = 2, 3 = 3, 4 = 1 carpel; 88. Ovule number per carpel: 0 = 1 or 2, 1 = >2; 89. Style: 0 = no, 1 = yes; 90. Stigma recurved: 0 = no, 1 = yes; 91. Stigma shape: 0 = entire, 1 = bifid, 2 = trifid; 92. Carpel pubescent: 0 = no, 1 = yes. Fruit. 93. Fruits apocarpous, 0 = no, 1 = yes; 94. Fruit shape: 0 = spheroid, 1 = ellipsoid; 95. Fruit colour: 0 = red, 1 = white, 2 = black; 96. Carpophore: 0 = no, 1 = yes; 97. Stylar scar position: 0 = basal, 1 = lateral, 2 = terminal; 98. Fruit pubescent: 0 = no, 1 = yes; 99. Mesocarp sclerified: 0 = no, 1 = yes; 100. Endocarp sclerified: 0 = no, 1 = yes; 101. Endocarp straight: 0 = no, 1 = yes; 102. Endocarp with excavated faces: 0 = no, 1 = yes; 103. If not straight, endocarp larger than wide: 0 = no, 1 = yes; 104. Endocarp with dorsal ridges: 0 = no, 1 = yes; 105. Number of lateral ridges on each face; 106. Ridges wing-shaped: 0 = no, 1 = yes; 107. Endocarp with lateral radial ridges: 0 = no, 1 = yes; 108. Endocarp surface reticulate: 0 = no, 1 = yes; 109. Type of reticulum: 0 = as hollow, 1 = as bump; 110. Endocarp surface with spines: 0 = no, 1 = yes; 111. Condyle present: 0 = no, 1 = yes; 112. Condyle outer conspicuous: 0 = no, 1 = yes; 113. Condyle type: 0 = single, 1 = double; 114. Condyle perforate: 0 = no, 1 = yes; 115. Condyle in prominent ventral chamber: 0 = no, 1 = yes; 116. If endocarp not straight, condyle parallel to symmetry plane: 0 = no, 1 = yes; 117. Embryo shape: 0 = straight, 1 = curved; 118. Endosperm: 0 = no, 1 = yes; 119. Endosperm ruminate: 0 = no, 1 = yes; 120. Cotyledon type: 0 = leafy, 1 = fleshy; 121. Cotyledons divaricate: 0 = no, 1 = yes; 122. Radicle length: 0 = small, 1 = equal. Pollen. 123. Pollen with colpus: 0 = no, 1 = yes; 124. Pollen with pore: 0 = no, 1 = yes; 125. Position of apertures: 0 = fosaperturate, 1 = angulaperturate, 2 = spiraperturate; 126. Pollen shape: 0 = P/E < 1, 1 = P/E = 1, 2 = P/E > 1; 127. Exine thickness: 0 = <1.5 nm, 1 = 1.5 nm, 2 = >1,5 nm; 128. Reticulation coarse: 0 = no, 1 = yes; 129. Wall section: 0 = cylindrical, 1 = triangular. 130. Endoaperture shape: 0 = elliptic, 1 = circular; 131. Lumina regular: 0 = no, 1 = yes; 132. Lumina hexagonal: 0 = no, 1 = yes. Phytochemistry. 133. Benzylisoquinoline: 0 = no, 1 = yes; 134. Bisbenzylisoquinoline: 0 = no, 1 = yes; 135. Proaporphine: 0 = no, 1 = yes; 136. Aporphine: 0 = no, 1 = yes; 137. Tropoloneisoquinoline: 0 = no, 1 = yes; 138. Azafluoranthene: 0 = no, 1 = yes; 139. Isooxaporphine: 0 = no, 1 = yes; 140. Protoberberine: 0 = no, 1 = yes; 141. Morphinane: 0 = no, 1 = yes; 142. Hasubanane: 0 = no, 1 = yes; 143. Acutimine: 0 = no, 1 = yes; 144. Isoquinoline: 0 = no, 1 = yes; 145. Phentylcinnamide: 0 = no, 1 = yes.
Rights and permissions
About this article
Cite this article
Jacques, F.M.B., Bertolino, P. Molecular and morphological phylogeny of Menispermaceae (Ranunculales). Plant Syst Evol 274, 83–97 (2008). https://doi.org/10.1007/s00606-008-0038-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00606-008-0038-7