Abstract
Purpose
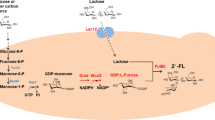
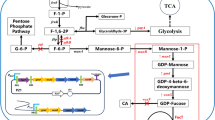
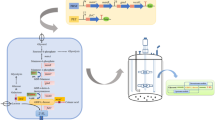
Pichia pastoris is well known for its ability to produce short and low-immunogenic humanized glycosyl chains onto recombinant glycoproteins, it was thus speculated to be applicable to synthesize oligosaccharides. In this study, generally recognized as safe (GRAS) microorganism Pichia pastoris GS115 was tested for its potential to be used as a new synthetic chassis to produce the most abundant human milk oligosaccharide 2′-fucosyllactose (2′-FL).
Methods
To enable the de novo synthesis of 2′-FL, lactose transporter lac12, two enzymes of gmd, gmer, and fucosyltransferases futC were integrated into the genome of P. pastoris, under the control of constitutive PGAP promoter.
Results
The resulting recombinant yeasts yielded up to 0.276 g/L through culture optimization in a 5 L bioreactor.
Conclusion
To our knowledge, this is the first report of 2′-FL production in engineered Pichia pastoris. This work is a good starting point to produce 2′-FL using Pichia pastoris as a viable chassis.
Graphical abstract











Similar content being viewed by others
References
Albermann C, Piepersberg W, Wehmeier UF (2001) Synthesis of the milk oligosaccharide 2’-fucosyllactose using recombinant bacterial enzymes. Carbohydr Res 334(2):97–103. https://doi.org/10.1016/s0008-6215(01)00177-x
Anderson A, Donald AS (1981) Improved method for the isolation of 2’ fucosyllactose from human milk. J Chromatogr 211(1):170–174. https://doi.org/10.1016/s0021-9673(00)81188-7
Araya-Garay JM, Feijoo-Siota L, Rosa-dos-Santos F, Veiga-Crespo P, Villa TG (2012) Construction of new Pichia pastoris X-33 strains for production of lycopene and β-carotene. Appl Microbiol Biotechnol 93(6):2483–2492. https://doi.org/10.1007/s00253-011-3764-7
Berger PK, Plows JF, Jones RB, Alderete TL, Yonemitsu C, Poulsen M, Goran MI (2020) Human milk oligosaccharide 2’-fucosyllactose links feedings at 1 month to cognitive development at 24 months in infants of normal and overweight mothers. PLoS ONE 15(2):0228323. https://doi.org/10.1371/journal.pone.0228323
Bhataya A, Schmidt-Dannert C, Lee PC (2009) Metabolic engineering of Pichia pastoris X-33 for lycopene production. Process Biochem 44(10):1095–1102. https://doi.org/10.1016/j.procbio.2009.05.012
Bode L (2012) Human milk oligosaccharides: every baby needs a sugar mama. Glycobiology 22(9):1147–1162. https://doi.org/10.1093/glycob/cws074
Bych K, Mikš MH, Johanson T, Hederos MJ, Vigsnæs LK, Becker P (2019) Production of HMOs using microbial hosts—from cell engineering to large scale production. Curr Opin Biotechnol 56:130–137
Çalık P, Bayraktar E, İnankur B, Soyaslan EŞ, Şahin M, Taşpınar H, Özdamar TH (2010) Influence of pH on recombinant human growth hormone production by Pichia pastoris. J Chem Technol Biotechnol 85(12):1628–1635. https://doi.org/10.1002/jctb.2474
Çelik E, Ozbay N, Oktar N, Çalık P (2008) Use of biodiesel byproduct crude glycerol as the carbon source for fermentation processes by recombinant Pichia pastoris. Ind Eng Chem Res 47(9):2985–2990
Deng J, Gu L, Chen T, Huang H, Yin X, Lv X, Li J (2019) Engineering the substrate transport and cofactor regeneration systems for enhancing 2′-fucosyllactose synthesis in Bacillus subtilis. ACS Synth Biol 8(10):2418–2427. https://doi.org/10.1021/acssynbio.9b00314
Deng J, Lv X, Li J, Du G, Chen J, Liu L (2020) Recent advances and challenges in microbial production of human milk oligosaccharides. Syst Microbiol Biomanuf 1:1–14
Döring F, Klapper M, Theis S, Daniel H (1998) Use of the glyceraldehyde-3-phosphate dehydrogenase promoter for production of functional mammalian membrane transport proteins in the yeast Pichia pastoris. Biochem Biophys Res Commun 250(2):531–535. https://doi.org/10.1006/bbrc.1998.9342
Facinelli B, Marini E, Magi G, Zampini L, Santoro L, Catassi C, Coppa GV (2019) Breast milk oligosaccharides: effects of 2’-fucosyllactose and 6’-sialyllactose on the adhesion of Escherichia coli and Salmonella fyris to Caco-2 cells. J Matern Fetal Neonatal Med 32(17):2950–2952. https://doi.org/10.1080/14767058.2018.1450864
Francisco G-R, Sergio A-S, Lorena G-R, Gabriela R-S, Mariano G-G, Alma C-G (2018) Synthesis of a fucosylated trisaccharide Via transglycosylation by α-L-fucosidase from thermotoga maritima. Appl Biochem Biotechnol 186(3):681–691
Han NS, Kim TJ, Park YC, Kim J, Seo JH (2012) Biotechnological production of human milk oligosaccharides. Biotechnol Adv 30(6):1268–1278. https://doi.org/10.1016/j.biotechadv.2011.11.003
Hollands K, Baron CM, Gibson KJ, Kelly KJ, Krasley EA, Laffend LA, Prasad JC (2019) Engineering two species of yeast as cell factories for 2′-fucosyllactose. Metab Eng 52:232–242
Holscher HD, Davis SR, Tappenden KA (2014) Human milk oligosaccharides influence maturation of human intestinal Caco-2Bbe and HT-29 cell lines. J Nutr 144(5):586–591. https://doi.org/10.3945/jn.113.189704
Huang D, Yang K, Liu J, Xu Y, Wang Y, Wang R, Feng L (2017) Metabolic engineering of Escherichia coli for the production of 2′-fucosyllactose and 3-fucosyllactose through modular pathway enhancement. Metab Eng 41:23–38
Izumi M, Tsuruta O, Harayama S, Hashimoto H (1997) Synthesis of 5-thio-L-fucose-containing disaccharides, as sequence specific inhibitors, and 2’-fucosyllactose, as a substrate of α- L fucosidases. Org Chem 62:992–998
Karbalaei M, Rezaee SA, Farsiani H (2020) Pichia pastoris: a highly successful expression system for optimal synthesis of heterologous proteins. J Cell Physiol 235(9):5867–5881. https://doi.org/10.1002/jcp.29583
Lee JW, Kwak S, Liu J-J, Yu S, Yun EJ, Kim DH, Jin Y-S (2020) Enhanced 2′-fucosyllactose production by engineered Saccharomyces cerevisiae using xylose as a co-substrate. Metab Eng 62:322–329
Lee JW, Kwak S, Liu J-J, Yun EJ, Jin Y-S (2021) 2’-Fucosyllactose production in engineered Escherichia coli with deletion of waaF and wcaJ and overexpression of FucT2. J Biotechnol 340:30–38
Lee H-J, Shin DJ, Han K, Chin Y-W, Park JP, Park K, Kim S-K (2022) Simultaneous production of 2′-fucosyllactose and difucosyllactose by engineered Escherichia coli with high secretion efficiency. Biotechnol J 17(3):2100629. https://doi.org/10.1002/biot.202100629
Li C, Wu M, Gao X, Zhu Z, Li Y, Lu F, Qin H-M (2020) Efficient biosynthesis of 2’-fucosyllactose using an in vitro multienzyme cascade. J Agric Food Chem. https://doi.org/10.1021/acs.jafc.0c04221
Lin-Cereghino J, Lin-Cereghino GP (2007) Vectors and strains for expression. Methods Mol Biol 389:11–26. https://doi.org/10.1007/978-1-59745-456-8_2
Liu J-J, Kwak S, Pathanibul P, Lee JW, Yu S, Yun EJ, Jin Y-S (2018) Biosynthesis of a functional human milk oligosaccharide, 2′-fucosyllactose, and l-fucose using engineered Saccharomyces cerevisiae. ACS Synth Biol 7(11):2529–2536. https://doi.org/10.1021/acssynbio.8b00134
Livak KJ, Schmittgen TD (2001) Analysis of relative gene expression data using real-time quantitative PCR and the 2−ΔΔCT method. Methods 25(4):402–408. https://doi.org/10.1006/meth.2001.1262
Marx H, Mattanovich D, Sauer M (2008) Overexpression of the riboflavin biosynthetic pathway in Pichia pastoris. Microb Cell Fact. https://doi.org/10.1186/1475-2859-7-23
Moriwaki K, Noda K, Nakagawa T, Asahi M, Yoshihara H, Taniguchi N, Miyoshi E (2007) A high expression of GDP-fucose transporter in hepatocellular carcinoma is a key factor for increases in fucosylation. Glycobiology 17(12):1311–1320
Nakayama K-I, Maeda Y, Jigami Y (2003) Interaction of GDP-4-keto-6-deoxymannose-3, 5-epimerase-4-reductase with GDP-mannose-4, 6-dehydratase stabilizes the enzyme activity for formation of GDP-fucose from GDP-mannose. Glycobiology 13(10):673–680
Park BS, Choi YH, Kim MW, Park BG, Kim E-J, Kim JY, Kim B-G (2022) Enhancing biosynthesis of 2’-Fucosyllactose in Escherichia coli through engineering lactose operon for lactose transport and α -1,2-Fucosyltransferase for solubility. Biotechnol Bioeng. https://doi.org/10.1002/bit.28048
Parschat K, Schreiber S, Wartenberg D, Engels B, Jennewein SM (2020) High-titer de novo biosynthesis of the predominant human milk oligosaccharide 2′ fucosyllactose from sucrose in Escherichia coli. ACS Synth Biol. https://doi.org/10.1021/acssynbio.0c00304
Parua PK, Ryan PM, Trang K, Young ET (2012) Pichia pastoris 14–3–3 regulates transcriptional activity of the methanol inducible transcription factor Mxr1 by direct interaction. Mol Microbiol 85(2):282–298. https://doi.org/10.1111/j.1365-2958.2012.08112.x
Prielhofer R, Cartwright SP, Graf AB, Valli M, Bill RM, Mattanovich D, Gasser B (2015) Pichia pastoris regulates its gene-specific response to different carbon sources at the transcriptional, rather than the translational, level. BMC Genomics 16(1):167–167. https://doi.org/10.1186/s12864-015-1393-8
Romanos MA, Scorer CA, Clare JJ (1992) Foreign gene expression in yeast: a review. Yeast 8(6):423–488. https://doi.org/10.1002/yea.320080602
Seo J, Chin Y, Jo H (2017) Method for producing 2′-fucosyllactose by using Corynebacterium glutamicum (WO2017188684). Google Scholar There is no corresponding record for this reference.
Seo J-H, Young-Wook C, Hae-Yong J (2020) Corynebacterium glutamicum for use in producing 2'-fucosyllactose. In: Google Patents.
Smilowitz JT, Lebrilla CB, Mills DA, German JB, Freeman SL (2014) Breast milk oligosaccharides: structure-function relationships in the neonate. Annu Rev Nutr 34:143–169. https://doi.org/10.1146/annurev-nutr-071813-105721
Spohner SC, Müller H, Quitmann H, Czermak P (2015) Expression of enzymes for the usage in food and feed industry with Pichia pastoris. J Biotechnol 202:118–134. https://doi.org/10.1016/j.jbiotec.2015.01.027
Tran A-M, Nguyen T-T, Nguyen C-T, Huynh-Thi X-M, Nguyen C-T, Trinh M-T, Tran-Van H (2017) Pichia pastoris versus Saccharomyces cerevisiae: a case study on the recombinant production of human granulocyte-macrophage colony-stimulating factor. BMC Res Notes 10(1):148–148. https://doi.org/10.1186/s13104-017-2471-6
Tu F-Z, Fu C-Y, Zhang T-Y, Luo J-X, Zhang A-L (2007) Constitutive expression of human angiostatin in Pichia pastoris using glycerol as only carbon source. Sheng Wu Gong Cheng Xue Bao = Chin J Biotechnol 23(5):902–906
Varela JA, Montini N, Scully D, Van der Ploeg R, Oreb M, Boles E, Morrissey JP (2017) Polymorphisms in the LAC12 gene explain lactose utilisation variability in Kluyveromyces marxianus strains. FEMS Yeast Res. https://doi.org/10.1093/femsyr/fox021
Várnai A, Tang C, Bengtsson O, Atterton A, Mathiesen G, Eijsink VGH (2014) Expression of endoglucanases in Pichia pastoris under control of the GAP promoter. Microb Cell Fact 13(1):57. https://doi.org/10.1186/1475-2859-13-57
Vazquez E, Barranco A, Ramirez M, Gruart A, Delgado-Garcia JM, Martinez-Lara E, Rueda R (2015) Effects of a human milk oligosaccharide, 2’-fucosyllactose, on hippocampal long-term potentiation and learning capabilities in rodents. J Nutr Biochem 26(5):455–465. https://doi.org/10.1016/j.jnutbio.2014.11.016
Vogl T, Glieder A (2013) Regulation of Pichia pastoris promoters and its consequences for protein production. New Biotechnol 30(4):385–404. https://doi.org/10.1016/j.nbt.2012.11.010
Wan L, Zhu Y, Chen G, Luo G, Zhang W, Mu W (2021) Efficient production of 2′-fucosyllactose from l-fucose via self-assembling multienzyme complexes in engineered Escherichia coli. ACS Synth Biol. https://doi.org/10.1021/acssynbio.1c00102
Waterham HR, Digan ME, Koutz PJ, Lair SV, Cregg JM (1997) Isolation of the Pichia pastoris glyceraldehyde-3-phosphate dehydrogenase gene and regulation and use of its promoter. Gene 186(1):37–44. https://doi.org/10.1016/s0378-1119(96)00675-0
Whittaker MM, Whittaker JW (2000) Expression of recombinant galactose oxidase by Pichia pastoris. Prot Expr Purif 20(1):105–111. https://doi.org/10.1006/prep.2000.1287
Wicinski M, Sawicka E, Gebalski J, Kubiak K, Malinowski B (2020) Human milk oligosaccharides: health benefits, potential applications in infant formulas, and pharmacology. Nutrients. https://doi.org/10.3390/nu12010266
Yang Z, Zhang Z (2018) Engineering strategies for enhanced production of protein and bio-products in Pichia pastoris: a review. Biotechnol Adv 36(1):182–195. https://doi.org/10.1016/j.biotechadv.2017.11.002
Yu S, Liu J-J, Yun EJ, Kwak S, Kim KH, Jin Y-S (2018) Production of a human milk oligosaccharide 2′-fucosyllactose by metabolically engineered Saccharomyces cerevisiae. Microb Cell Fact 17(1):1–10
Zabel BE, Gerdes S, Evans KC, Nedveck D, Singles SK, Volk B, Budinoff C (2020) Strain-specific strategies of 2’-fucosyllactose, 3-fucosyllactose, and difucosyllactose assimilation by Bifidobacterium longum subsp infantis Bi-26 and ATCC 15697. Sci Rep 10(1):15919. https://doi.org/10.1038/s41598-020-72792-z
Zhang G, Zhao J, Wen R, Zhu X, Liu L, Li C (2020) 2’-fucosyllactose promotes Bifidobacterium bifidum DNG6 adhesion to caco-2 cells. J Dairy Sci 103(11):9825–9834. https://doi.org/10.3168/jds.2020-18773
Zhang Q, Liu Z, Xia H, Huang Z, Zhu Y, Xu L, Liu L (2022a) Engineered Bacillus subtilis for the de novo production of 2′-fucosyllactose. Microbial Cell Fact 21(1):110. https://doi.org/10.1186/s12934-022-01838-w
Zhang X, Zhang C, Zhou M, Xia Q, Fan L, Zhao L (2022b) Enhanced bioproduction of chitin in engineered Pichia pastoris. Food Biosci 47:101606
Zhou W, Jiang H, Wang L, Liang X, Mao X (2021) Biotechnological production of 2′-fucosyllactose: a prevalent fucosylated human milk oligosaccharide. ACS Synth Biol 10(3):447–458
Acknowledgements
This research was supported by the National Key R&D Program of China (Grant Number 2019YFD0901800), Shanghai Post-doctoral Excellence Program (2020132), Natural Science Foundation of Shanghai (22ZR1418200) and the 111 Project (B18022).
Funding
This study was supported by National Key R& D Program of China, 2019YFD0901800, Shanghai Post-doctoral Excellence Program, 2020132, Natural Science Foundation of Shanghai, 22ZR1418200, 111 Project, B18022.
Author information
Authors and Affiliations
Contributions
LZ and LF conceived and supervised the project. DQ and MZ designed the experiments. DQ and CZ performed the experiments and wrote the manuscript.
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare no conflict of interest.
Ethical approval
This article does not contain any studies on humans or animals.
Informed consent
All relevant personnel in this study have obtained informed consent.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Qian, D., Zhang, C., Deng, C. et al. De novo biosynthesis of 2′-fucosyllactose in engineered Pichia pastoris. Biotechnol Lett 45, 521–536 (2023). https://doi.org/10.1007/s10529-023-03357-z
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10529-023-03357-z