Abstract
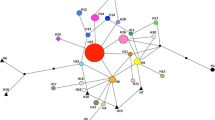
Population genetic surveys can reveal the patterns of genetic diversity and provide insights into the details of invasion history. Plantago virginica L. (Plantaginaceae) is a North America species that was first reported in China in 1951 and has rapidly spread into the south-eastern areas of China. Although previous work has examined phenotypic variation during this invasion, the genetic variation and structure of P. virginica remains unclear. We used two chloroplast and seven microsatellite loci to analyse genetic diversity and population differentiation in 17 populations of P. virginica from the invaded range (China, n = 688) and 10 populations from its native range (USA, n = 625). Chloroplast haplotypes from all populations were clustered into two lineages (I and II) and invasive populations had 28 haplotypes, whereas native populations had only 10 haplotypes. DIYABC simulations based on microsatellite data also suggest multiple invasions into China. Invasive populations displayed high genetic diversity that was similar to that of the native range. However, there was markedly less genetic differentiation in Chinese populations (FST = 0.051) than in NA populations (FST = 0.143), suggesting increased gene flow within China. STRUCTURE analysis also revealed significant gene flow and lineage admixture among Chinese populations and MIGRATE analysis suggested high levels of gene flow to inland China from both the coast of China and populations in the United States. These results suggest that multiple introductions and genetic admixture likely play important roles in facilitating the invasion and geographic expansion of P. virginica into China.




Similar content being viewed by others
Data availability
Data from the current study are available from the corresponding author upon reasonable request, and the NCBI GenBank accession numbers are MW722451–MW722520.
References
Ahlstrand NI, Verstraete B, Hassemer G, Dunbar-Co S, Hoggard R, Meudt HM, Rønsted N (2019) Ancestral range reconstruction of remote oceanic island species of Plantago (Plantaginaceae) reveals differing scales and modes of dispersal. J Biogeogr 46:706–722
Allendorf FW, Lundquist LL (2003) Introduction: population biology, evolution, and control of invasive species. Conserv Biol 17:24–30
Amos W, Hoffman JI, Frodsham A, Zhang L, Best S, Hill AVS (2010) Automated binning of microsatellite alleles: problems and solutions. Mol Ecol Resour 7:10–14
Bandelt HJ, Forster P, Rohl A (1999) Median-joining networks for inferring intraspecific phylogenies. Mol Biol Evol 16:37–48
Barrett SCH, Husband BC (1990) Plant population genetics, breeding, and genetic resources. Sinauer, Sunderland, pp. 254–278
Beerli P, Palczewski M (2010) Unified framework to evaluate panmixia and migration direction among multiple sampling locations. Genetics 185:313–326
Bossdorf O, Auge H, Lafuma L, Rogers WE, Siemann E, Prati D (2005) Phenotypic and genetic differentiation between native and introduced plant populations. Oecologia 144:1–11
Brown AHD, Marshall DR (1981) Evolutionary changes accompanying colonization in plants. Hunt Institute, Pittsburgh, pp. 351–363
Cornuet JM, Pudlo P, Veyssier J, Dehne-Garcia A, Gautier M, Leblois R, Marin JM, Estoup A (2014) DIYABC v2.0: a software to make approximate Bayesian computation inferences about population history using single nucleotide polymorphism. DNA Seq Microsatell Data Bioinfo 30:187–1189
Ding J, Mack RN, Lu L, Ren M, Huang H (2008) China’s booming economy is sparking and accelerating biological invasions. Bioscience 58:317–324
Earl DA, von Holdt BM (2012) STRUCTURE HARVESTER: a website and program for visualizing STRUCTURE output and implementing the Evanno method. Conserv Genet Resour 4:359–361
Ellstrand NC, Schierenbeck KA (2000) Hybridization as a stimulus for the evolution of invasiveness in plants? P Natl Acad Sci USA 97:7043–7050
Evanno G, Regnaut S, Goudet J (2005) Detecting the number of clusters of individuals using the software STRUCTURE: a simulation study. Mol Ecol 14:2611–2620
Excoffier L, Laval G, Schneider S (2007) Arlequin (version 3.0): an integrated software package for population genetics data analysis. Evol Bioinform Online 1:47–50
Fang F, Guo S, Huang H, Cao A (2004) Influences of population density of Plantago virginica on its morphological characters of underground and aboveground organs. J Trop Subtrop Bot 12:419–424
Felker-Quinn E, Bailey JK, Schweitzer JA (2011) Soil biota drive expression of genetic variation and development of population-specific feedbacks in an invasive plant. Ecology 92:1208–1214
Guo S, Gu D, Liu P, Hu Y (1996) Biological and ecological characteristics of Plantago virginica L. Acta Ecol Sin 16:302–307
Guo SL, Fang F, Qiang S (2004) Studies on the reproduction and photosynthetic ecophysiology of the exotic invasive plant, Plantago virginica. Acta Phytoecol Sin 28:787–793
Havrdová A, Douda J, Krak K, Vít P, Hadincová V, Zákravský P, Mandák B (2015) Higher genetic diversity in recolonized areas than in refugia of Alnus glutinosa triggered by continent-wide lineage admixture. Mol Ecol 24:4759–4777
Hulme PE, Bacher S, Kenis M, Klotz S, Kühn I, Minchin D (2008) Grasping at the routes of biological invasions: a framework for integrating pathways into policy. J Appl Ecol 45:403–414
Kalinowski ST, Taper ML, Marshall TC (2010) Revising how the computer program CERVUS accommodates genotyping error increases success in paternity assignment. Mol Ecol 16:1099–1106
Larkin MA, Blackshields G, Brown NP, Chenna RM, Higgins DG (2007) ClustalW and ClustalX version 2. Bioinformatics 23:2947–2948
Librado P, Rozas J (2009) DnaSP v5: a software for comprehensive analysis of DNA polymorphism data. Bioinformatics 25:1451–1452
Luo X, Xu X, Zheng Y, Guo H, Hu S (2019) The role of phenotypic plasticity and rapid adaptation in determining invasion success of Plantago virginica. Biol Invasions 21:2679–2692
Maebara Y, Tamaoki M, Iguchi Y, Nakahama N, Hayasaka D (2020) Genetic diversity of invasive Spartina alterniflora Loisel. (Poaceae) introduced unintentionally into japan and its invasion pathway. Front Plant Sci 11:1–12
Peakall R, Smouse PE (2012) GenAlEx 6.5: genetic analysis in Excel. Population genetic software for teaching and research—an update. Bioinformatics 28:2537–2539
Pritchard JK, Stephens M, Donnelly P (2000) Inference of population structure using multilocus genotype data. Genetics 155:945–959
Qiao HM, Liu WW, Zhang YH, Zhang YY, Li QQ (2019) Genetic admixture accelerates invasion via provisioning rapid adaptive evolution. Mol Ecol 28:4012–4027
Shaw J, Lickey EB, Beck JT, Farmer SB, Liu W, Miller J (2005) The tortoise and the hare: relative utility of 21 noncoding chloroplast DNA sequences for phylogenetic analysis. Am J Bot 92:142–166
Sudhir K, Glen S, Koichiro T (2016) MEGA7: molecular evolutionary genetics analysis version 7.0 for bigger datasets. Mol Biol Evol 33:1870–1884
Sun Y, Bossdorf O, Grados RD, Liao ZY, Müller-Schrer H (2020) Rapid genomic and phenotypic change in response to climate warming in a widespread plant invader. Glob Change Biol 26:6511–6522
Tang JS, Ma M (2020) Genetic diversity and genetic differentiation of invasive weed Xanthium italicum in China. CR Biol 343:63–72
Vicente S, Cristina M, Trindade H (2018) Genetic diversity and differentiation of invasive Acacia longifolia in Portugal. Web Ecol 18:91–103
Wang HT, Zhou YM, Chen Y (2015) Allelopathic potential of invasive Plantago virginica on four lawn species. PLoS ONE 10:1–12
Wang J, Gaughan S, Lamer JD, Deng C, Hu WT (2020) Resolving the genetic paradox of invasions: Preadapted genomes and post introduction hybridization of bigheaded carps in the Mississippi River Basin. Evol Appl 13:263–277
Wang R, Wan FH (2010) Prediction of the potential survival area of Xanthium italicum in China. Acta Pratac Sin 6:222–230
Wang XY, Shen DW, Jiao J, Xu NN, Yu S, Zhou XF (2012) Genotypic diversity enhances invasive ability of Spartina alterniflora. Mol Ecol 21:2542–2551
White DM, Huang JP, Jara-Muoz OA, Madrián S, Mason-Gamer RJ (2021) The origins of coca: museum genomics reveals multiple independent domestications from progenitor Erythroxylum gracilipes. Syst Biol 70:1–13
Xiao YA, Zhou B, Hu WH (2012) Differences in the selectivity of competitive strategy between Plantago virginica and P. asiatica in natural habitats. J JINGANGSHAN U (Nat Sci) 33:96–103
Xu X, Luo X, Wang X, Guo H, Hu S (2017) Microsatellite primers in Plantago virginica (Plantaginaceae), an invasive species with both cleistogamous and chasmogamous flowers. Genes Genet Syst 92:293–297
Xu X, Wolfe L, Diez J, Zheng Y, Guo H, Hu S (2019) Differential germination strategies of native and introduced populations of the invasive species Plantago virginica. NeoBiota 43:101–118
Young ND, Healy J (2003) GapCoder automates the use of indel characters in phylogenetic analysis. BMC Bioinfo 4:6–12
Zhang DX, Liu LM, Guo SL (2010) Analysis of genetic diversity in Plantago virginica population. Weed Sci 2:14–17
Zhang Y, Tang JS, Ren G, Zhao KX, Wang XF (2021) Global potential distribution prediction of Xanthium italicum based on Maxent model. Sci Rep 11:16545
Acknowledgements
This work was supported by the National Natural Science Foundation of China (No. 31100298, U20A2080, 31622015) and Sichuan University (Fundamental Research Funds for the Central Universities, SCU2020D003, SCU2021D006). We thank Dr. Lei Shang, Meng Lu, Qiang Yang, Yazhu Liu and other colleagues for thier help and insightful discussions during the experimental processes. We are grateful to Prof. Zhiping Song for his very helpful suggestions.
Funding
The authors have not disclosed any funding.
Author information
Authors and Affiliations
Contributions
JT and KM wrote the manuscript. XX and XX performed the experiments. JT and HZ performed the data analysis. HG and BL conceived and designed the experiments. KM and HG finalized the manuscript, all authors read and revised the manuscript.
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that there is no conflict of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Tang, J., Mao, K., Zhang, H. et al. Multiple introductions and genetic admixture facilitate the successful invasion of Plantago virginica into China. Biol Invasions 24, 2261–2272 (2022). https://doi.org/10.1007/s10530-022-02773-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10530-022-02773-y