Abstract
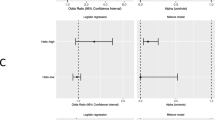
Sequence-based testing of disease-susceptibility genes has identified many variants of unknown significance (VUSs) whose pathogenicity is unknown at the time of their measurement. Female breast cancer cases aged 20–49 years at diagnosis and who have VUSs in BRCA1 and no mutations in BRCA2 have previously been identified through the population-based Los Angeles County Cancer Surveillance Program. These nominal BRCA1 VUSs have been classified as “low,” “medium,” and “high” risk by four classification methods: Align-GVGD, Polyphen, Grantham matrix scores, and sequence conservation in mammalian species. Average hazard ratios (HRs) for classes of variants, i.e., the age-specific incidences of cancer for carriers of such variants divided by the population incidences, were estimated from the cancer family histories of first- and second-degree relatives of the index cases using modified segregation analysis. The study sample comprised 270 index cases and 4,543 of their relatives. There was weak evidence that the risk of breast cancer increases with the degree of sequence conservation (P = 0.03) and that missense variants at highly conserved sites are associated with a 5.6-fold (95 % confidence interval 1.4–22.2; P = 0.05) increased incidence of breast cancer. An upper bound of 2.3 is given for the average breast cancer HRs corresponding to variants classified as “low risk” by any of the four VUS classification methods. In summary, we have given a method to estimate cancer risks for groups of VUSs by combining existing classification methods with traditional penetrance analyses. This analysis suggests that classification methods for BRCA1 variants based on sequence conservation might be useful in a clinical setting. We have shown in principle that our method can be used to classify VUSs into clinically useful risk categories, but our specific findings should not be put into clinical practice unless confirmed by larger studies.
Similar content being viewed by others
References
Woods MO, Williams P, Careen A, Edwards L, Bartlett S, McLaughlin JR, Younghusband HB (2007) A new variant database for mismatch repair genes associated with Lynch syndrome. Hum Mutat 28(7):669–673. doi:10.1002/humu.20502
Szabo C, Masiello A, Ryan JF, Brody LC (2000) The breast cancer information core: database design, structure, and scope. Hum Mutat 16(2):123–131. doi:10.1002/1098-1004(200008)16:2<123:AID-HUMU4>3.0.CO;2-Y
Adzhubei IA, Schmidt S, Peshkin L, Ramensky VE, Gerasimova A, Bork P, Kondrashov AS, Sunyaev SR (2010) A method and server for predicting damaging missense mutations. Nat Methods 7(4):248–249. doi:10.1038/nmeth0410-248
Arnold S, Buchanan DD, Barker M, Jaskowski L, Walsh MD, Birney G, Woods MO, Hopper JL, Jenkins MA, Brown MA, Tavtigian SV, Goldgar DE, Young JP, Spurdle AB (2009) Classifying MLH1 and MSH2 variants using bioinformatic prediction, splicing assays, segregation, and tumor characteristics. Hum Mutat 30(5):757–770. doi:10.1002/humu.20936
Goldgar DE, Healey S, Dowty JG, Da Silva L, Chen X, Spurdle AB, Terry MB, Daly MJ, Buys SM, Southey MC, Andrulis I, John EM, Khanna KK, Hopper JL, Oefner PJ, Lakhani S, Chenevix-Trench G (2011) Rare variants in the ATM gene and risk of breast cancer. Breast Cancer Res 13(4):R73. doi:10.1186/bcr2919
Grantham R (1974) Amino acid difference formula to help explain protein evolution. Science 185(4154):862–864
Ramensky V, Bork P, Sunyaev S (2002) Human non-synonymous SNPs: server and survey. Nucleic Acids Res 30(17):3894–3900
Tavtigian SV, Deffenbaugh AM, Yin L, Judkins T, Scholl T, Samollow PB, de Silva D, Zharkikh A, Thomas A (2006) Comprehensive statistical study of 452 BRCA1 missense substitutions with classification of eight recurrent substitutions as neutral. J Med Genet 43(4):295–305. doi:10.1136/jmg.2005.033878
Lee E, McKean-Cowdin R, Ma H, Chen Z, Van Den Berg D, Henderson BE, Bernstein L, Ursin G (2008) Evaluation of unclassified variants in the breast cancer susceptibility genes BRCA1 and BRCA2 using five methods: results from a population-based study of young breast cancer patients. Breast Cancer Res 10(1):R19. doi:10.1186/bcr1865
Ma H, Bernstein L, Ross RK, Ursin G (2006) Hormone-related risk factors for breast cancer in women under age 50 years by estrogen and progesterone receptor status: results from a case–control and a case–case comparison. Breast Cancer Res 8(4):R39. doi:10.1186/bcr1514
Marchbanks PA, McDonald JA, Wilson HG, Folger SG, Mandel MG, Daling JR, Bernstein L, Malone KE, Ursin G, Strom BL, Norman SA, Wingo PA, Burkman RT, Berlin JA, Simon MS, Spirtas R, Weiss LK (2002) Oral contraceptives and the risk of breast cancer. N Engl J Med 346(26):2025–2032. doi:10.1056/NEJMoa013202
McKean-Cowdin R, Spencer Feigelson H, Xia LY, Pearce CL, Thomas DC, Stram DO, Henderson BE (2005) BRCA1 variants in a family study of African-American and Latina women. Hum Genet 116(6):497–506. doi:10.1007/s00439-004-1240-5
Freedman ML, Penney KL, Stram DO, Le Marchand L, Hirschhorn JN, Kolonel LN, Altshuler D, Henderson BE, Haiman CA (2004) Common variation in BRCA2 and breast cancer risk: a haplotype-based analysis in the Multiethnic Cohort. Hum Mol Genet 13(20):2431–2441. doi:10.1093/hmg/ddh270
Abkevich V, Zharkikh A, Deffenbaugh AM, Frank D, Chen Y, Shattuck D, Skolnick MH, Gutin A, Tavtigian SV (2004) Analysis of missense variation in human BRCA1 in the context of interspecific sequence variation. J Med Genet 41(7):492–507
Antoniou AC, Pharoah PD, McMullan G, Day NE, Ponder BA, Easton D (2001) Evidence for further breast cancer susceptibility genes in addition to BRCA1 and BRCA2 in a population-based study. Genet Epidemiol 21(1):1–18. doi:10.1002/gepi.1014
Lange K (2002) Mathematical and statistical methods for genetic analysis. Statistics for biology and health, 2nd edn. Springer, New York
Kraft P, Thomas DC (2000) Bias and efficiency in family-based gene-characterization studies: conditional, prospective, retrospective, and joint likelihoods. Am J Hum Genet 66(3):1119–1131. doi:10.1086/302808
Gong G, Hannon N, Whittemore AS (2010) Estimating gene penetrance from family data. Genet Epidemiol 34(4):373–381. doi:10.1002/gepi.20493
Cannings C, Thompson E, Skolnick M (1978) Probability functions on complex pedigrees. Adv Appl Prob 10(1):26–61
Antoniou AC, Pharoah PD, McMullan G, Day NE, Stratton MR, Peto J, Ponder BJ, Easton DF (2002) A comprehensive model for familial breast cancer incorporating BRCA1, BRCA2 and other genes. Br J Cancer 86(1):76–83. doi:10.1038/sj.bjc.6600008
Ries LAGKC, Hankey BF, Harras A, Miller BA, Edwards BK (1996) SEER cancer statistics review, 1973–1993: tables and graphs. National Cancer Institute, Bethesda
Lange K, Weeks D, Boehnke M (1988) Programs for Pedigree Analysis: MENDEL, FISHER, and dGENE. Genet Epidemiol 5(6):471–472. doi:10.1002/gepi.1370050611
Team RDC (2010) R: A language and environment for statistical computing. R Foundation for Statistical Computing, Vienna
Akbari MR, Zhang S, Fan I, Royer R, Li S, Risch H, McLaughlin J, Rosen B, Sun P, Narod SA (2011) Clinical impact of unclassified variants of the BRCA1 and BRCA2 genes. J Med Genet 48(11):783–786. doi:10.1136/jmedgenet-2011-100305
Acknowledgments
The authors would like to thank Sean V. Tavtigian for his insightful comments and for his help with Align-GVGD. The authors are also very grateful to the women who participated in this study, the interviewers who collected the data, the phlebotomists who collected the blood samples, and Ms. Juliana Bamrick who managed all study activities. This study was supported by grants CA17054 and CA74847 from the National Cancer Institute, National Institutes of Health, by 4 PB-0092 from the California Breast Cancer Research Program of the University of California, and in part through contract number N01-PC-35139 and T32 ES-013678 from the National Institute of Environmental Health Sciences, National Institute of Health. The collection of cancer incidence data used in this publication was supported by the California Department of Health Services as part of the statewide cancer reporting program mandated by California Health and Safety Code Section 103885.
Conflict of interest
The authors declare that they have no conflicts of interest.
Disclaimer
Authors had full responsibility for the design of the study, the collection of the data, the analysis and interpretation of the data, the decision to submit the manuscript for publication, and the writing of the manuscript. The ideas and opinions expressed herein are those of the authors, and no endorsement by the State of California, Department of Health Services is intended or should be inferred.
Ethics standard
The Women’s Learning the Influence of Family and Environment Study was approved by the Institutional Review Board of the University of Southern California. All participants provided written informed consent. All experiments complied with the current laws of the US where they were performed.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Dowty, J.G., Lee, E., McKean-Cowdin, R. et al. Estimating risks for variants of unknown significance according to their predicted pathogenicity classes with application to BRCA1 . Breast Cancer Res Treat 144, 171–177 (2014). https://doi.org/10.1007/s10549-014-2845-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10549-014-2845-6