Abstract
Purpose
We aimed to characterize genetic correlations and causal associations between circulating C-reactive protein (CRP) levels and the risk of lung cancer (LC).
Methods
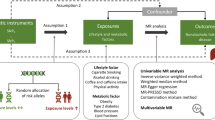
Leveraging summary statistics from genome-wide association studies of circulating CRP levels among 575,531 individuals of European ancestry, and LC risk among 29,266 cases and 56,450 controls, we investigated genetic associations of circulating CRP levels with the risk of overall lung cancer and its histological subtypes, by using linkage disequilibrium score (LDSC) regression and Mendelian randomization (MR) analyses.
Results
Significant positive genetic correlations between circulating CRP levels and the risk of LC and its histological subtypes were identified from LDSC regression, with correlation coefficients ranging from 0.12 to 0.26, and all false discovery adjusted p < 0.05. Univariable MR demonstrated a nominal association between CRP levels and an increased risk of lung squamous cell carcinoma (SCC) (inverse variance-weighted OR = 1.15, 95% CI 1.01–1.30). However, this association disappeared when multivariable MR included cigarettes per day and/or body mass index. By using our recently developed constrained maximum likelihood-based MR method, we identified significant associations of CRP levels with the risk of overall LC (OR 1.06, 95% CI 1.03–1.09), SCC (OR 1.06, 95% CI 1.02–1.09), and small cell lung cancer (SCLC, OR 1.09, 95% CI 1.03–1.15). Moreover, most univariable and multivariable MR analyses also revealed consistent CRP-SCLC associations.
Conclusion
There may be a genetic and causal association between circulating CRP levels and the risk of SCLC, which is in line with previous population-based observational studies.




Similar content being viewed by others
Data availability
Summary statistics of GWAS for C-reactive protein and body mass index can be obtained from GWAS catalog (https://www.ebi.ac.uk/gwas) that for cigarettes per day and drinks per week from the Data Repository for University of Minnesota (DRUM) website(https://conservancy.umn.edu/drum), and those lung cancer GWAS data can be obtained from dbGaP (https://www.ncbi.nlm.nih.gov/gap/) under accession phs001273.v1.p1.
References
Torre LA, Bray F, Siegel RL et al (2015) Global cancer statistics, 2012. CA Cancer J Clin 65:87–108. https://doi.org/10.3322/caac.21262
Bray F, Ferlay J, Soerjomataram I et al (2018) Global cancer statistics 2018: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J Clin 68:394–424. https://doi.org/10.3322/caac.21492
Malhotra J, Malvezzi M, Negri E et al (2016) Risk factors for lung cancer worldwide. Eur Respir J 48:889–902. https://doi.org/10.1183/13993003.00359-2016
Yang IA, Holloway JW, Fong KM (2013) Genetic susceptibility to lung cancer and co-morbidities. J Thorac Dis 5(5):S454-462. https://doi.org/10.3978/j.issn.2072-1439.2013.08.06
Schwartz AG, Cote ML (2016) Epidemiology of lung cancer. Adv Exp Med Biol 893:21–41. https://doi.org/10.1007/978-3-319-24223-1_2
Bossé Y, Amos CI (2018) A decade of GWAS results in lung cancer. Cancer Epidemiol Biomarkers Prev 27:363–379. https://doi.org/10.1158/1055-9965.EPI-16-0794
Grivennikov SI, Greten FR, Karin M (2010) Immunity, inflammation, and cancer. Cell 140:883–899. https://doi.org/10.1016/j.cell.2010.01.025
Elinav E, Nowarski R, Thaiss CA et al (2013) Inflammation-induced cancer: crosstalk between tumours, immune cells and microorganisms. Nat Rev Cancer 13:759–771. https://doi.org/10.1038/nrc3611
Shalapour S, Karin M (2015) Immunity, inflammation, and cancer: an eternal fight between good and evil. J Clin Invest 125:3347–3355. https://doi.org/10.1172/JCI80007
Greten FR, Grivennikov SI (2019) Inflammation and cancer: triggers, mechanisms, and consequences. Immunity 51:27–41. https://doi.org/10.1016/j.immuni.2019.06.025
Karki R, Kanneganti TD (2019) Diverging inflammasome signals in tumorigenesis and potential targeting. Nat Rev Cancer 19:197–214. https://doi.org/10.1038/s41568-019-0123-y
Cho WC, Kwan CK, Yau S et al (2011) The role of inflammation in the pathogenesis of lung cancer. Expert Opin Ther Targets 15:1127–1137. https://doi.org/10.1517/14728222.2011.599801
Gomes M, Teixeira AL, Coelho A et al (2014) The role of inflammation in lung cancer. Adv Exp Med Biol 816:1–23. https://doi.org/10.1007/978-3-0348-0837-8_1
Budisan L, Zanoaga O, Braicu C et al (2021) Links between infections, lung cancer, and the immune system. Int J Mol Sci 22:9394. https://doi.org/10.3390/ijms22179394
Agassandian M, Shurin GV, Ma Y, Shurin MR (2014) C-reactive protein and lung diseases. Int J Biochem Cell Biol 53:77–88. https://doi.org/10.1016/j.biocel.2014.05.016
Chaturvedi AK, Caporaso NE, Katki HA et al (2010) C-reactive protein and risk of lung cancer. J Clin Oncol Off J Am Soc Clin Oncol 28:2719–2726. https://doi.org/10.1200/JCO.2009.27.0454
Zhou B, Liu J, Wang Z-M, Xi T (2012) C-reactive protein, interleukin 6 and lung cancer risk: a meta-analysis. PLoS ONE 7:e43075. https://doi.org/10.1371/journal.pone.0043075
Ji M, Du L, Ma Z et al (2022) Circulating C-reactive protein increases lung cancer risk: results from a prospective cohort of UK Biobank. Int J Cancer 150:47–55. https://doi.org/10.1002/ijc.33780
Edwards JK, Cole SR, Westreich D (2015) All your data are always missing: incorporating bias due to measurement error into the potential outcomes framework. Int J Epidemiol 44:1452–1459. https://doi.org/10.1093/ije/dyu272
Hammerton G, Munafò MR (2021) Causal inference with observational data: the need for triangulation of evidence. Psychol Med 51:563–578. https://doi.org/10.1017/S0033291720005127
Yarmolinsky J, Wade KH, Richmond RC et al (2018) Causal inference in cancer epidemiology: what is the role of Mendelian randomization? Cancer Epidemiol Biomark Prev 27:995–1010. https://doi.org/10.1158/1055-9965.EPI-17-1177
Bulik-Sullivan BK, Loh P-R, Finucane HK et al (2015) LD Score regression distinguishes confounding from polygenicity in genome-wide association studies. Nat Genet 47:291–295. https://doi.org/10.1038/ng.3211
Bulik-Sullivan B, Finucane HK, Anttila V et al (2015) An atlas of genetic correlations across human diseases and traits. Nat Genet 47:1236–1241. https://doi.org/10.1038/ng.3406
Davey Smith G, Hemani G (2014) Mendelian randomization: genetic anchors for causal inference in epidemiological studies. Hum Mol Genet 23:R89–R98. https://doi.org/10.1093/hmg/ddu328
Davies NM, Holmes MV, Davey Smith G (2018) Reading Mendelian randomisation studies: a guide, glossary, and checklist for clinicians. BMJ. https://doi.org/10.1136/bmj.k601
Pingault JB, O’Reilly PF, Schoeler T et al (2018) Using genetic data to strengthen causal inference in observational research. Nat Rev Genet 19:566–580. https://doi.org/10.1038/s41576-018-0020-3
Han X, Ong JS, An J et al (2020) Using Mendelian randomization to evaluate the causal relationship between serum C-reactive protein levels and age-related macular degeneration. Eur J Epidemiol 35:139–146. https://doi.org/10.1007/s10654-019-00598-z
Sabatti C, Service SK, Hartikainen AL et al (2009) Genome-wide association analysis of metabolic traits in a birth cohort from a founder population. Nat Genet 41:35–46
Sanderson E, Davey Smith G, Windmeijer F, Bowden J (2019) An examination of multivariable Mendelian randomization in the single-sample and two-sample summary data settings. Int J Epidemiol 48:713–727. https://doi.org/10.1093/ije/dyy262
Xue H, Shen X, Pan W (2021) Constrained maximum likelihood-based Mendelian randomization robust to both correlated and uncorrelated pleiotropic effects. Am J Hum Genet 108:1251–1269. https://doi.org/10.1016/j.ajhg.2021.05.014
Said S, Pazoki R, Karhunen V et al (2022) Genetic analysis of over half a million people characterises C-reactive protein loci. Nat Commun 13:2198. https://doi.org/10.1038/s41467-022-29650-5
Saunders GRB, Wang X, Chen F et al (2022) Genetic diversity fuels gene discovery for tobacco and alcohol use. Nature 612:720–724. https://doi.org/10.1038/s41586-022-05477-4
Pulit SL, Stoneman C, Morris AP et al (2019) Meta-analysis of genome-wide association studies for body fat distribution in 694 649 individuals of European ancestry. Hum Mol Genet 28:166–174. https://doi.org/10.1093/hmg/ddy327
McKay JD, Hung RJ, Han Y et al (2017) Large-scale association analysis identifies new lung cancer susceptibility loci and heterogeneity in genetic susceptibility across histological subtypes. Nat Genet 49:1126–1132. https://doi.org/10.1038/ng.3892
Korthauer K, Kimes PK, Duvallet C et al (2019) A practical guide to methods controlling false discoveries in computational biology. Genome Biol 20:118. https://doi.org/10.1186/s13059-019-1716-1
Bowden J, Del Greco MF, Minelli C et al (2017) A framework for the investigation of pleiotropy in two-sample summary data Mendelian randomization. Stat Med 36:1783–1802. https://doi.org/10.1002/sim.7221
Bowden J, Davey Smith G, Haycock PC, Burgess S (2016) Consistent estimation in Mendelian randomization with some invalid instruments using a weighted median estimator. Genet Epidemiol 40:304–314. https://doi.org/10.1002/gepi.21965
Bowden J, Davey Smith G, Burgess S (2015) Mendelian randomization with invalid instruments: effect estimation and bias detection through Egger regression. Int J Epidemiol 44:512–525. https://doi.org/10.1093/ije/dyv080
Verbanck M, Chen CY, Neale B, Do R (2018) Detection of widespread horizontal pleiotropy in causal relationships inferred from Mendelian randomization between complex traits and diseases. Nat Genet 50:693–698. https://doi.org/10.1038/s41588-018-0099-7
Burgess S, Thompson SG (2015) Multivariable Mendelian randomization: the use of pleiotropic genetic variants to estimate causal effects. Am J Epidemiol 181:251–260. https://doi.org/10.1093/aje/kwu283
Bowden J, Holmes MV (2019) Meta-analysis and Mendelian randomization: a review. Res Synth Methods 10:486–496. https://doi.org/10.1002/jrsm.1346
Brion M-JA, Shakhbazov K, Visscher PM (2013) Calculating statistical power in Mendelian randomization studies. Int J Epidemiol 42:1497–1501. https://doi.org/10.1093/ije/dyt179
Hemani G, Zheng J, Elsworth B et al (2018) The MR-base platform supports systematic causal inference across the human phenome. Elife. https://doi.org/10.7554/eLife.34408
Sanderson E, Spiller W, Bowden J (2021) Testing and correcting for weak and pleiotropic instruments in two-sample multivariable Mendelian randomization. Stat Med 40:5434–5452. https://doi.org/10.1002/sim.9133
Muller DC, Larose TL, Hodge A et al (2019) Circulating high sensitivity C reactive protein concentrations and risk of lung cancer: nested case-control study within lung cancer cohort consortium. BMJ 364:k4981. https://doi.org/10.1136/bmj.k4981
Siemes C, Visser LE, Coebergh JWW et al (2006) C-reactive protein levels, variation in the C-reactive protein gene, and cancer risk: the Rotterdam Study. J Clin Oncol Off J Am Soc Clin Oncol 24:5216–5222. https://doi.org/10.1200/JCO.2006.07.1381
Hamilton G, Rath B (2015) Smoking, inflammation and small cell lung cancer: recent developments. Wien Med Wochenschr 165:379–386. https://doi.org/10.1007/s10354-015-0381-6
Çolak Y, Afzal S, Lange P, Nordestgaard BG (2019) Smoking, systemic inflammation, and airflow limitation: a Mendelian randomization analysis of 98 085 individuals from the general population. Nicotine Tob Res Off J Soc Res Nicotine Tob 21:1036–1044. https://doi.org/10.1093/ntr/nty077
Galan D, Perry BI, Warrier V et al (2022) Applying Mendelian randomization to appraise causality in relationships between smoking, depression and inflammation. Sci Rep 12:15041. https://doi.org/10.1038/s41598-022-19214-4
Zhou W, Liu G, Hung RJ et al (2021) Causal relationships between body mass index, smoking and lung cancer: univariable and multivariable Mendelian randomization. Int J Cancer 148:1077–1086. https://doi.org/10.1002/ijc.33292
Markozannes G, Kanellopoulou A, Dimopoulou O et al (2022) Systematic review of Mendelian randomization studies on risk of cancer. BMC Med 20:41. https://doi.org/10.1186/s12916-022-02246-y
Koskeridis F, Evangelou E, Said S et al (2022) Pleiotropic genetic architecture and novel loci for C-reactive protein levels. Nat Commun 13:6939. https://doi.org/10.1038/s41467-022-34688-6
Goto T (2022) Microbiota and lung cancer. Semin Cancer Biol 86:1–10. https://doi.org/10.1016/j.semcancer.2022.07.006
Knaapen AM, Borm PJA, Albrecht C, Schins RPF (2004) Inhaled particles and lung cancer. Part A: Mechanisms Int J Cancer 109:799–809. https://doi.org/10.1002/ijc.11708
Acknowledgements
We thank the TRICL-ILCCO, UK Biobank, GSCAN, GIANT, and CHARGE for providing valuable data resources for this study.
Funding
This work was supported in part by a grant from the National Cancer Institute R01CA249863 (to Qiuyin Cai and Jirong Long).
Author information
Authors and Affiliations
Contributions
JS, WW, and QC conceived the study. JS and HX analyzed the data. JS, WW, HX, RT, and WP contributed to methodology. JS, JL, YY, and QC obtained resources. HX and WP contributed to software development. WW, XS, and QC supervised the study. JS drafted the manuscript. All authors edited and approved the final draft.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Ethical approval
Our study is based on publicly available summary statistics from genome-wide association studies that had already obtained ethical approval from respective review boards or ethics committees. Therefore, obtaining a separate ethical approval for this study was not required.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Shi, J., Wen, W., Long, J. et al. Genetic correlation and causal associations between circulating C-reactive protein levels and lung cancer risk. Cancer Causes Control (2024). https://doi.org/10.1007/s10552-024-01855-7
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s10552-024-01855-7