Abstract
Background
Esophageal adenocarcinoma (EAC) has a very high case fatality rate and is one of the fastest rising cancers worldwide. At the same time, research into EAC has been hampered by a relative lack of pre-clinical models, including representative cell lines.
Aim
The purpose of this study was to establish and characterize a new EAC cell line.
Methods
Tumor cells were isolated from EAC tissue by enzymatic digestion. Origin of the cell line was confirmed by microsatellite-based genotyping. A panel of cancer-related genes was screened for mutations by targeted deep sequencing, Sanger sequencing and high resolution melting. CDKN2A promoter methylation was assessed by methylation specific high resolution melting. HER2 amplification was assessed by fluorescent in situ hybridization. Immunohistochemistry was used to assess expression of markers in xenografts grown in SCID mice.
Results
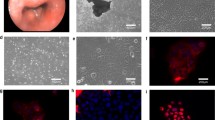
A novel EAC cell line, OANC1, was derived from a Barrett’s-associated EAC. Microsatellite-based genotyping of OANC1 and patient DNA confirmed the origin of the cell line. Sequencing of OANC1 DNA identified homozygous TP53 missense (c.856G>A, p.E286K) and SMAD4 nonsense (c.1333C>T, p.R445X) mutations. OANC1 are tumorigenic when injected sub-cutaneously into SCID mice and xenografts were positive for columnar, glandular and intestinal epithelial markers commonly expressed in EAC. Xenografts exhibited strong p53 expression, consistent with a TP53 mutation. Some proteins, including p16, EGFR and β-catenin, had heterogeneous expression patterns across xenograft cross-sections, indicative of tumor heterogeneity.
Conclusions
OANC1 represents a valuable addition to the limited range of pre-clinical models for EAC. This new cell line will be a useful model system for researchers studying both basic and translational aspects of this disease.






Similar content being viewed by others
References
Buas MF, Vaughan TL. Epidemiology and risk factors for gastroesophageal junction tumors: understanding the rising incidence of this disease. Semin Radiat Oncol. 2013;23:3–9.
Polednak AP. Trends in survival for both histologic types of esophageal cancer in US surveillance, epidemiology and end results areas. Int J Cancer. 2003;105:98–100.
Crane SJ, Locke GR 3rd, Harmsen WS, Zinsmeister AR, Romero Y, Talley NJ. Survival trends in patients with gastric and esophageal adenocarcinomas: a population-based study. Mayo Clin Proc. 2008;83:1087–1094.
Bouvier AM, Binquet C, Gagnaire A, Jouve JL, Faivre J, Bedenne L. Management and prognosis of esophageal cancers: has progress been made? Eur J Cancer. 2006;42:228–233.
Younes M, Henson DE, Ertan A, Miller CC. Incidence and survival trends of esophageal carcinoma in the United States: racial and gender differences by histological type. Scand J Gastroenterol. 2002;37:1359–1365.
Cook MB, Chow WH, Devesa SS. Oesophageal cancer incidence in the United States by race, sex, and histologic type, 1977–2005. Br J Cancer. 2009;101:855–859.
Pohl H, Welch HG. The role of overdiagnosis and reclassification in the marked increase of esophageal adenocarcinoma incidence. J Natl Cancer Inst. 2005;97:142–146.
Thrift AP, Whiteman DC. The incidence of esophageal adenocarcinoma continues to rise: analysis of period and birth cohort effects on recent trends. Ann Oncol Off J Eur Soc Med Oncol ESMO. 2012;23:3155–3162.
Boonstra JJ, Tilanus HW, Dinjens WNM. Translational research on esophageal adenocarcinoma: from cell line to clinic. Dis Esophagus. 2013. doi:10.1111/dote.12095.
Boonstra JJ, van Marion R, Beer DG, et al. Verification and unmasking of widely used human esophageal adenocarcinoma cell lines. J Natl Cancer Inst. 2010;102:271–274.
Dulak AM, Stojanov P, Peng S, et al. Exome and whole-genome sequencing of esophageal adenocarcinoma identifies recurrent driver events and mutational complexity. Nat Genet. 2013;45:478–486.
Krypuy M, Newnham GM, Thomas DM, Conron M, Dobrovic A. High resolution melting analysis for the rapid and sensitive detection of mutations in clinical samples: KRAS codon 12 and 13 mutations in non-small cell lung cancer. BMC Cancer. 2006;6:295.
Do H, Krypuy M, Mitchell PL, Fox SB, Dobrovic A. High resolution melting analysis for rapid and sensitive EGFR and KRAS mutation detection in formalin fixed paraffin embedded biopsies. BMC Cancer. 2008;8:142.
Krypuy M, Ahmed AA, Etemadmoghadam D, et al. High resolution melting for mutation scanning of TP53 exons 5–8. BMC Cancer. 2007;7:168.
Do H, Dobrovic A. Dramatic reduction of sequence artefacts from DNA isolated from formalin-fixed cancer biopsies by treatment with uracil- DNA glycosylase. Oncotarget. 2012;3:546–558.
Wojdacz TK, Dobrovic A. Methylation-sensitive high resolution melting (MS-HRM): a new approach for sensitive and high-throughput assessment of methylation. Nucleic Acids Res. 2007;35:e41.
Mandard AM, Dalibard F, Mandard JC, et al. Pathologic assessment of tumor regression after preoperative chemoradiotherapy of esophageal carcinoma. Clinicopathologic correlations. Cancer. 1994;73:2680–2686.
Dangles-Marie V, Pocard M, Richon S, et al. Establishment of human colon cancer cell lines from fresh tumors versus xenografts: comparison of success rate and cell line features. Cancer Res. 2007;67:398–407.
Graham TA, McDonald SA. Genetic diversity during the development of Barrett’s oesophagus-associated adenocarcinoma: how, when and why? Biochem Soc Trans. 2010;38:374–379.
Biankin AV, Waddell N, Kassahn KS, et al. Pancreatic cancer genomes reveal aberrations in axon guidance pathway genes. Nature. 2012;491:399–405.
Fleming NI, Jorissen RN, Mouradov D, et al. SMAD2, SMAD3 and SMAD4 mutations in colorectal cancer. Cancer Res. 2013;73:725–735.
Jones S, Zhang X, Parsons DW, et al. Core signaling pathways in human pancreatic cancers revealed by global genomic analyses. Science. 2008;321:1801–1806.
Agrawal N, Jiao Y, Bettegowda C, et al. Comparative genomic analysis of esophageal adenocarcinoma and squamous cell carcinoma. Cancer Discov. 2012;2:899–905.
Phillips WA, Russell SE, Ciavarella ML, et al. Mutation analysis of PIK3CA and PIK3CB in esophageal cancer and Barrett’s esophagus. Int J Cancer. 2006;118:2644–2646.
Vieth M, Schneider-Stock R, Rohrich K, et al. INK4a-ARF alterations in Barrett’s epithelium, intraepithelial neoplasia and Barrett’s adenocarcinoma. Virchows Arch. 2004;445:135–141.
Sommerer F, Vieth M, Markwarth A, et al. Mutations of BRAF and KRAS2 in the development of Barrett’s adenocarcinoma. Oncogene. 2004;23:554–558.
Lord RV, O’Grady R, Sheehan C, Field AF, Ward RL. K-ras codon 12 mutations in Barrett’s oesophagus and adenocarcinomas of the oesophagus and oesophagogastric junction. J Gastroenterol Hepatol. 2000;15:730–736.
Kwak EL, Jankowski J, Thayer SP, et al. Epidermal growth factor receptor kinase domain mutations in esophageal and pancreatic adenocarcinomas. Clin Cancer Res. 2006;12:4283–4287.
Clemons N, Phillips W, Lord RV. Signaling pathways in the molecular pathogenesis of adenocarcinomas of the esophagus and gastresophageal junction. Cancer Biol Ther. 2013. doi:10.4161/cbt.25362.
Barrett MT, Sanchez CA, Galipeau PC, Neshat K, Emond M, Reid BJ. Allelic loss of 9p21 and mutation of the CDKN2/p16 gene develop as early lesions during neoplastic progression in Barrett’s esophagus. Oncogene. 1996;13:1867–1873.
Bian YS, Osterheld MC, Fontolliet C, Bosman FT, Benhattar J. p16 inactivation by methylation of the CDKN2A promoter occurs early during neoplastic progression in Barrett’s esophagus. Gastroenterology. 2002;122:1113–1121.
Wong DJ, Barrett MT, Stoger R, Emond MJ, Reid BJ. p16INK4a promoter is hypermethylated at a high frequency in esophageal adenocarcinomas. Cancer Res. 1997;57:2619–2622.
Hardie LJ, Darnton SJ, Wallis YL, et al. p16 expression in Barrett’s esophagus and esophageal adenocarcinoma: association with genetic and epigenetic alterations. Cancer Lett. 2005;217:221–230.
Sarbia M, Geddert H, Klump B, Kiel S, Iskender E, Gabbert HE. Hypermethylation of tumor suppressor genes (p16INK4A, p14ARF and APC) in adenocarcinomas of the upper gastrointestinal tract. Int J Cancer. 2004;111:224–228.
Dahlberg PS, Jacobson BA, Dahal G, et al. ERBB2 amplifications in esophageal adenocarcinoma. Ann Thorac Surg. 2004;78:1790–1800.
Brien TP, Odze RD, Sheehan CE, McKenna BJ, Ross JS. HER-2/neu gene amplification by FISH predicts poor survival in Barrett’s esophagus-associated adenocarcinoma. Hum Pathol. 2000;31:35–39.
Rossi E, Villanacci V, Bassotti G, et al. Her-2/neu in barrett esophagus: a comparative study between histology, immunohistochemistry, and fluorescence in situ hybridization. Diagn Mol Pathol. 2006;15:125–130.
Reichelt U, Duesedau P, Tsourlakis M, et al. Frequent homogeneous HER-2 amplification in primary and metastatic adenocarcinoma of the esophagus. Mod Pathol. 2007;20:120–129.
Walch A, Specht K, Braselmann H, et al. Coamplification and coexpression of GRB7 and ERBB2 is found in high grade intraepithelial neoplasia and in invasive Barrett’s carcinoma. Int J Cancer. 2004;112:747–753.
Sehdev V, Peng D, Soutto M, et al. The aurora kinase A inhibitor MLN8237 enhances cisplatin-induced cell death in esophageal adenocarcinoma cells. Mol Cancer Ther. 2012;11:763–774.
Zhang K, Zhang S, Jiao X, et al. Slug regulates proliferation and invasiveness of esophageal adenocarcinoma cells in vitro and in vivo. Med Oncol. 2011;28:1089–1100.
Santander S, Cebrian C, Esquivias P et al. Cyclooxygenase inhibitors decrease the growth and induce regression. Int J Oncol. 2012;40(2):527–534.
Hasina R, Mollberg N, Kawada I, et al. Critical role for the receptor tyrosine kinase EPHB4 in esophageal cancers. Cancer Res. 2013;73:184–194.
Alvarez H, Koorstra JB, Hong SM, et al. Establishment and characterization of a bona fide Barrett esophagus-associated adenocarcinoma cell line. Cancer Biol Ther. 2008;7:1753–1755.
Acknowledgments
The authors would like to thank the Peter MacCallum Cancer Centre Department of Pathology for performing immunohistochemistry, FISH and sequencing, the Histology Core in the Research Division for preparation of xenograft tissue blocks, and Dr. James Eshleman for the provision of JH-EsoAd1 cells. The patient history and fresh and FFPE tumour tissue used in this project was provided by the Victorian Cancer Biobank with appropriate ethics approval. The Victorian Cancer Biobank is supported by the Victorian Government, Australia. NJC and WAP are supported by grants from the National Health and Medical Research Council, Australia. This work was supported by grants from the National Health and Medical Research Council, Australia.
Conflict of interest
None.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Clemons, N.J., Do, H., Fennell, C. et al. Characterization of a Novel Tumorigenic Esophageal Adenocarcinoma Cell Line: OANC1. Dig Dis Sci 59, 78–88 (2014). https://doi.org/10.1007/s10620-013-2882-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10620-013-2882-8