Abstract
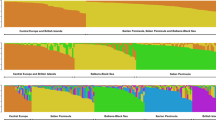
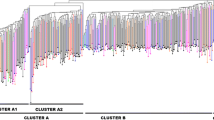
World hazelnut production is based primarily on selections from the wild. In this study, we used 21 pairs of simple sequence repeat (SSR) primers to investigate genetic diversity in 270 clonal accessions of European hazelnut (Corylus avellana) representing a wide geographic range. Of the 270 accessions, 198 had unique fingerprints while 72 were duplicates. Based on the 198 unique accessions, the number of detected alleles per locus averaged 9.81 and observed heterozygosity (Ho) averaged 0.67. Of the 206 total alleles amplified, 20 were unique to a single accession. A genetic similarity matrix was constructed and the resulting dendrogram revealed four major geographical groups: Central European, Black Sea, English and Spanish-Italian. SSR alleles indicated the parentage of 31 accessions. The fingerprints are publicly available through the Germplasm Resources Information Network (GRIN) database. The identification of duplicate and mislabeled accessions will improve management of hazelnut genebanks, and information on genetic variation in hazelnut will assist the international research community.





Similar content being viewed by others
References
Ahmad R, Ferguson L, Southwick SM (2003) Identification of pistachio (Pistacia vera L.) nuts with microsatellite markers. J Am Soc Hortic Sci 128:898–903
Aranzana MJ, Carbó J, Arus P (2002) Microsatellite variability in peach [Prunus persica (L.) Batsch.]: cultivar identification, marker mutation, pedigree inferences and population structure. Theor Appl Genet 106:1341–1352
Bassil NV, Botta R, Mehlenbacher SA (2005) Microsatellite markers in hazelnut: isolation, characterization and cross-species amplification. J Am Soc Hortic Sci 130:543–549
Boccacci P, Akkak A, Bassil NV, Mehlenbacher SA, Botta R (2005) Characterization and evaluation of microsatellite loci in European hazelnut (Corylus avellana L.) and their transferability to other Corylus species. Mol Ecol Notes 5:934–937. doi:10.1111/j.1471-8286.2005.01121.x
Boccacci P, Akkak A, Botta R (2006) DNA typing and genetic relations among European hazelnut (Corylus avellana L.) cultivars using microsatellite markers. Genome 49:598–611. doi:10.1139/G06-017
Botstein D, White RL, Skolnick MH, Davies RW (1980) Construction of genetic linkage map in man using restriction fragment length polymorphisms. Am J Hum Genet 32:314–331 Medline
Bowcock AM, Ruiz-Linares A, Tomforhde J, Minch E, Kidd JR, Cavalli-Sforza LL (1994) High resolution of human evolutionary trees with polymorphic microsatellites. Nature 368:455–457. doi:10.1038/368455a0
Brookfield JFY (1996) A simple new method for estimating null allele frequency from heterozygote deficiency. Mol Ecol 5:453–455. doi:10.1111/j.1365-294X.1996.tb00336.x
Brooks RM, Olmo HP (1997) Register of fruit & nut varieties, 3rd edn. ASHS Press, Alexandria, VA
Cabe PR, Baumgarten A, Onan K, Luby JJ, Bedford DS (2005) Using microsatellite analysis to verify breeding records: a study of ‘Honeycrisp’ and other cold-hardy apple cultivars. HortScience 40:15–17
Callen DF, Thompson AD, Shen Y, Philips HA, Richards RI, Mullen JC et al (1993) Incidence and origin of “null” alleles in the (AC)n microsatellite markers. Am J Hum Genet 52:922–927 Medline
Dangl GS, Mendum ML, Prins BH, Walker MA, Meredith CP, Simon CJ (2001) Simple sequence repeat analysis of a clonally propagated species: a tool for managing a grape germplasm collection. Genome 44:432–438. doi:10.1139/gen-44-3-432
FAO Production Yearbook (2004) faostat.fao.org (verified 19 May 2008)
Fatahi R, Ebadi A, Bassil NV, Mehlenbacher SA, Zamani Z (2003) Characterization of Iranian grapevine cultivars using microsatellite markers. Vitis 42:185–192
Gellatly JU (1964) Filazels. Ann Rept North Nut Growers Assn 55:153–155
Germain E (1994) The reproduction of hazelnut (Corylus avellana): a review. Acta Hortic 351:195–210
Goeschke F (1887) Die Haselnuss, ihre Arten und ihre Kultur. Paul Parey, Berlin
Guo X, Elston RC (1999) Linkage information content of polymorphic genetic markers. Hum Hered 49:112–118. doi:10.1159/000022855
Hampson CR, Azarenko AN, Soeldner A (1993) Pollen-stigma interactions following compatible and incompatible pollinations in hazelnut. J Am Soc Hortic Sci 118:814–819
Hokanson SC, Lamboy WF, Szewc-McFadden AK, McFerson JR (2001) Microsatellite (SSR) variation in a collection of Malus (apple) species and hybrids. Euphytica 118:281–294. doi:10.1023/A:1017591202215
Kalinowski ST, Taper ML, Marshall TC (2007) Revising how the computer program CERVUS accommodates genotyping error increases success in paternity assignment. Mol Ecol 16:1099–1106. doi:10.1111/j.1365-294X.2007.03089.x
Kasapligil B (1972) A bibliography on Corylus (Betulaceae) with annotations. Ann Rept North Nut Growers Assn 63:107–162
Kumar S, Tamura K, Nei M (2004) MEGA3: integrated software for molecular evolutionary genetics analysis and sequence alignment. Brief Bioinform 5:150–163. doi:10.1093/bib/5.2.150
Lagerstedt HB (1990) Filbert rootstock and cultivar introductions in Oregon. Ann Rept North Nut Growers Assn 81:60–63
Li Y, Karol AB, Fahima T, Beiles A, Nevo E (2002) Microsatellites: genomic distribution, putative functions, and mutational mechanisms: a review. Mol Ecol 11:2453–2464. doi:10.1046/j.1365-294X.2002.01643.x
Liu K, Muse S (2004) PowerMarker: new genetic data analysis software. Version 3.2 (http://statgen.ncsu.edu/powermarker/) (verified 19 May 2008)
Lunde CF, Mehlenbacher SA, Smith DC (2000) Survey of hazelnut cultivars for response to eastern filbert blight inoculation. HortScience 35:729–731
Lunde CF, Mehlenbacher SA, Smith DC (2006) Segregation for resistance to eastern filbert blight in progeny of ‘Zimmerman’ hazelnut. J Am Soc Hortic Sci 131:731–737
Manzo P, Tamponi G (1982) Monografia di cultivar di nocciuolo. Instituto Sperimentale per la Frutticoltura, Roma
Marshall TC, Slate J, Kruuk LEB, Pemberton JM (1998) Statistical confidence for likelihood-based paternity inference in natural populations. Mol Ecol 7:639–655. doi:10.1046/j.1365-294x.1998.00374.x
Martin JP, Borrego J, Cabello F, Ortiz JM (2003) Characterization of Spanish grapevine cultivar diversity using sequence-tagged microsatellite site markers. Genome 46:10–18. doi:10.1139/g02-098
Mehlenbacher SA (1997a) Revised dominance hierarchy for S-alleles in Corylus avellana L. Theor Appl Genet 94:360–366. doi:10.1007/s001220050424
Mehlenbacher SA (1997b) Testing compatibility of hazelnut crosses using fluorescence microscopy. Acta Hortic 445:167–171
Mehlenbacher SA, Miller AN (1989) ‘Barcelona’ hazelnut. Fruit Var J 43:90–95
Mehlenbacher SA, Brown RN, Nouhra ER, Gokirmak T, Bassil NV, Kubisiak TL (2006) A genetic linkage map for hazelnut (Corylus avellana L.) based on RAPD and SSR markers. Genome 49:122–133 Medline
Nei M (1973) Analysis of gene diversity in subdivided populations. Proc Natl Acad Sci USA 70:3321–3323. doi:10.1073/pnas.70.12.3321
Palmé AE, Vendramin GG (2002) Chloroplast DNA variation, postglacial recolonization and hybridization in hazel, Corylus avellana. Mol Ecol 11:1769–1779. doi:10.1046/j.1365-294X.2002.01581.x
Romero C, Pedryc A, Munoz V, Llacer G, Badenes ML (2003) Genetic diversity of different apricot geographical groups determined by SSR markers. Genome 46:244–252. doi:10.1139/g02-128
Summers K, Amos W (1997) Behavioral, ecological, and molecular genetic analyses of reproductive strategies in the Amazonian dart-poison frog, Dendrobates ventrimaculatus. Behav Ecol 8:260–267. doi:10.1093/beheco/8.3.260
Tasias Valls J (1975) El avellano en la provincia de Tarragona. EXCMA. Diputación Provincial de Tarragona, Tarragona, Spain, 363 pp
Testolin R, Marrazzo T, Cipriani G, Quarta R, Verde I, Dettori MT et al (2000) Microsatellite DNA in peach [Prunus persica (L.) Batsch)] and its use in fingerprinting and testing the genetic origin of cultivars. Genome 43:512–520. doi:10.1139/gen-43-3-512
Thompson MM, Lagerstedt HB, Mehlenbacher SA (1996) Hazelnuts. In: Janick J, Moore JN (eds) Fruit breeding. Nuts, vol 3. Wiley, New York, pp 125–184
Turcu E, Botu I (1997) Behaviour of some hazelnut varieties in the subcarpathian area of Oltenia-Romania. Acta Hortic 445:135–144
Weir BS (1996) Genetic data analysis II: methods for discrete population genetic data. Sinauer Associates, Sunderland, MA, pp 163–166
Zane L, Bargelloni L, Patarnello T (2002) Strategies for microsatellite isolation: a review. Mol Ecol 11:1–16. doi:10.1046/j.0962-1083.2001.01418.x
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Gökirmak, T., Mehlenbacher, S.A. & Bassil, N.V. Characterization of European hazelnut (Corylus avellana) cultivars using SSR markers. Genet Resour Crop Evol 56, 147–172 (2009). https://doi.org/10.1007/s10722-008-9352-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10722-008-9352-8