Abstract
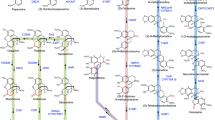
Since ancient times, opium poppy (Papaver somniferum L.) is known for its medicinal properties, related to its secondary metabolite content. Its most important secondary metabolites, called benzylisoquinoline alkaloids (BIAs), are still essential in pharmaceutical field. Few of them, like morphine, have specific clinical application but also effects on CNS. Not all poppy cultivars are able to biosynthesize morphine in high amount, making this plant useful for other purposes like food uses. For this reason it is crucial to deeply understand the origin of poppy, its possible use and have a deep knowledge of the BIA biosynthesis. These aspects are crucial for the final use of P. somniferum. This review aims to summarize the state-of-the-art on its taxonomy and origin beside its uses and BIA biosynthetic pathways, its most important metabolites. The review focuses on conflicting or unsolved questions about enzymatic localization, role of different plant organs in the biosynthesis, and storage and external conditions that influence the alkaloid production, highlighting the significant involvement of transcription factors. Behind this review, there is the firm belief that only a deep knowledge of alkaloid biosynthetic processes could lead to the characterization of undefined step and to the development of engineering cultivars optimizing the potential uses of P. somniferum. The goal is answer in more sustainable way to ever-increasing worldwide request of such products, in particular morphine and derivates, obtaining high morphine content cultivars useful for pharmaceutical market or no morphine producing cultivars appreciated as food. Devising cultivars with different BIA content could lead to decrease, or even avoid, illicit use and illegal extraction, confining only low alkaloid content cultivars to consumers market.






Similar content being viewed by others
Abbreviations
- 3′OHase:
-
3′-Hydroxylase
- 3′OMT:
-
3′-O-methyltransferase
- 3OHase:
-
Tyrosine/tyramine 3-hydroxylase
- 4′OMT:
-
3′-hydroxyl-N-methylcoclaurine 4′-O-methyltransferase
- 4HPAA:
-
4 Hydroxyphenylacetic acid
- 4HPPDC:
-
4-Hydroxyphenylpuruvate decarboxylase
- 6OMT:
-
Norcoclaurine 6-O-methyltransferase
- 7OMT:
-
Reticuline 7-O-methyltransferase
- AC:
-
Adenylyl cyclase
- AFLP:
-
Amplified fragment length polymorphism
- AMP:
-
Adenosine monophosphate
- AMPc:
-
Adenosine monophosphate cyclic
- BBE:
-
Berberine bridge enzyme
- BIAs:
-
Benzylisoquinoline alkaloids
- CAS:
-
Canadine synthase
- CFS:
-
Cheilanthifoline synthase
- CNMT:
-
Coclaurine N-methyltransferase
- CNS:
-
Central nervous system
- CODM:
-
Codeine O-demethylase
- CoOMT:
-
Columbamine O-methyltransferase
- COR:
-
Codeinone reductase
- CPVO:
-
Community Plant Variety Office
- CYP:
-
Cytochrome P450 enzyme family
- CYP82Y1:
-
N-methylcanadine 1-hydroxylase
- CYP80B:
-
(S)-N-methylcoclaurine 3′-hydroxylase isozyme
- Cys:
-
Cysteine
- DA:
-
Dopamine
- DBOX:
-
Dihydrosanguinarine oxidase
- EFSA:
-
European Food Safety Authority
- ER:
-
Endoplasmic reticulum
- EST:
-
Expressed sequence tag
- FT–ICR–MS:
-
Fourier-transform–ion-cyclotron resonance–mass spectrometry
- GC:
-
Gas chromatography
- GPCR:
-
G protein–coupled receptors
- HPLC:
-
High performance liquid chromatography
- MeJa:
-
Methyl jasmonate
- MIA:
-
Monoterpenoid indole alkaloid
- MLP15:
-
Major latex proteins
- MSH:
-
N-methylstylopine 14-hydroxylase
- N7OMT:
-
Norreticuline 7-O-methyltransferase
- NAc:
-
Nucleus accumbens
- NCS:
-
Norcoclaurine synthase
- NMCH:
-
N-methylcoclaurine 3′-hydroxylase
- NOS:
-
Noscapine synthase
- Nuclear factor-κB:
-
Nuclear factor kappa-light-chain-enhancer of activated B cells
- OR receptors:
-
Receptors for endogenous opiates
- ORL1:
-
Opioid receptor-like
- P6H:
-
Protopine 6-hydroxylase
- PKA:
-
Protein kinase A
- PKC:
-
Protein kinase C
- PPH:
-
Protopine-6-hydroxylase
- RAPD:
-
Random amplified polymorphic DNA
- RFLP:
-
Restriction fragment length polymorphism
- ROS:
-
Reactive oxygen species
- RSP:
-
Restriction site polymorphism
- RVM:
-
Rostral ventromedial medulla
- SalAT:
-
Salutaridinol 7-O-acetyltransferase
- SalR:
-
Salutaridine reductase
- SanR:
-
Sanguinarine reductase
- SAT:
-
(7S)-salutaridinol 7-O-acetyltransferase
- SNPs:
-
Single nucleotide polymorphism
- SOMT1:
-
Scoulerine 9-O-methyltransferase
- SOR:
-
7-Oxidoreductase
- SPS:
-
Stylopine synthase
- STOX:
-
(S)-tetrahydroxyprotoberberineoxidase
- STRs:
-
Short tandem repeat
- STS:
-
Salutaridine synthase
- StySyn:
-
Sstylopine synthase
- T6ODM:
-
Thebaine 6-O-demethylase
- TFBSs:
-
Transcription factor binding bites
- TFs:
-
Transcription factors
- TNMT:
-
Tetrahydroprotoberberine N-methyltransferase
- TYDC:
-
Tyrosine/DOPA decarboxylase
- UHPLC–ESI–QTOF–MS:
-
Ultra high performance liquid chromatography–electrospray ionization–quadrupole time-of-flight–mass spectrometry
- VNTRs:
-
Variable number of tandem repeat or minisatellites
References
Acheson RM, Harper JL, Mcnaughton IH (1956) Distribution of anthocyanin pigments in poppies. Nature 178:1283–1284
Acheson RM, Jenkins CL, Harper JL et al (2006) Floral pigments in papaver and their significance in the systematics of the genus. New Phytol 61(3):256–260
Agarwal PS, Lakhwani D et al (2016) Comparative analysis of transcription factor gene families from Papaver somniferum: identification of regulatory factors involved in benzylisoquinoline alkaloid biosynthesis. Protoplasma 253(3):857–871
Bajpai S, Gupta AP, Gupta MM et al (2000) Inter-relation between descriptors and morphine yield in Asian germplasm of opium poppy Papaver somniferum. Genet Resour Crop Evol 47(3):315–322
Batterham RL, Cowley MA, Small CJ et al (2004) Physiology: does gut hormone PYY3-36 decrease food intake in rodents? Nature 430(6996):413–414
Beaudoin GA, Facchini JP (2014) Benzylisoquinoline alkaloid biosynthesis in opium poppy. Planta 240(1):19–32
Bird DA, Franceschi VR, Facchini PJ (2003) A tale of three cell types: alkaloid biosynthesis is localized to sieve elements in opium poppy. The Plant Cell 15:2626–2635
Celik I, Gultekin V, Allmer J et al (2014) Development of genomic simple sequence repeat markers in opium poppy by next-generation sequencing. Mol Breed 34(2):323–334
Danert S (1958) Zur Systematik von Papaver somniferum L. Kulturpflanze 6:61–88
Dang TT, Onoyovwi A, Farrow SC et al (2012) Biochemical genomics for gene discovery in benzylisoquinoline alkaloid biosynthesis in opium poppy and related species. Methods Enzymol 515:231–266
Darokar MP, Dhawan SS, Shukla AK et al (2014) Assessment of genetic relatedness among selected cultivars of opium poppy (Papaver somniferum L.) through DNA profiling. Acta Hortic 1036:21–28
Desgagné-Penix I, Khan MF, Schremier DC et al (2010) Integration of deep transcriptome and proteome analyses reveals the components of alkaloid metabolism in opium poppy cell cultures. BMC Plant Biol 10(1):252
Desgagné-Penix I, Farrow SC, Cram D et al (2012) Integration of deep transcript and targeted metabolite profiles for eight cultivars of opium poppy. Plant Mol Biol 79(3):295–313
Dittbrenner A, Lohwasser U, Mock HP et al (2008) Molecular and phytochemical studies of Papaver somniferum in the context of infraspecific classification. Acta Hortic 799:81–88
Duarte DF (2005) Uma breve história do ópio e dos opióides. Rev Bras Anestesiol 55(1):135–146
Egan PA, Pendry CA, Shresth S (2011) Flora of Nepal Papaveraceae. Royal Botanic Gardens, Melbourne
El Nehir S, Karakaya S (2004) Radical scavenging and iron-chelating activities of some greens used as traditional dishes in mediterranean diet. Int J Food Sci Nutr 55(1):67–74
Eryilmaz T, Yesilyurt MK, Cesur C et al (2016) Biodiesel production potential from oil seeds in Turkey. Renew Sustain Energy Rev 58:842–851
European Food Safety Authority (EFSA) (2011) Scientific opinion on the risks for public health related to the presence of opium alkaloids in poppy seeds. EFSA J 9(11):1–150
Facchini PJ (2001) Alkaloid biosynthesis in plants: biochemistry, cell biology, molecular regulation, and metabolic engineering applications. Annu Rev Plant Biol 52(1):29–66
Facchini PJ, De Luca V (2008) Opium poppy and Madagascar periwinkle: model non-model systems to investigate alkaloid biosynthesis in plants. Plant Journal 54(4):763–784
Fernández EC, Rajchl A, Lachman J et al (2013) Impact of yacon landraces cultivated in the Czech Republic and their ploidy on the short- and long-chain fructooligosaccharides content in tuberous roots. LWT Food Sci Technol 54(1):80–86
Froede R (1972) Drugs of abuse: legal and illegal. Hum Pathol 3(1):23–36
Gurkok T, Turktas M, Parmaksiz I et al (2015) Transcriptome profiling of alkaloid biosynthesis in elicitor induced opium poppy. Plant Mol Biol Rep 33(3):673–688
Hagel JM, Facchini PJ (2013) Benzylisoquinoline alkaloid metabolism: a century of discovery and a brave new world. Plant Cell Physiol 54(5):647–672
Hammer K (1981) Problems of Papaver somniferum classification and some remarks on recently collected European poppy land-races. Genet Resour Crop Evol 29:287–296
Hao D, Xiao JG, Pei GX (2015) Medicinal plant: chemistry, biology and omics. Woodhead Publishing, Cambridge
Hofman PJ, Menary RC (1985) Alkaloid losses from the capsules of Papaver Somniferum L. during kiln drying and storage under commercial conditions in Tasmania. J Stored Prod Res 21(3):135–139
Inukai S, Hong Kock K, Bulyk ML (2017) Transcription factor-DNA binding: beyond binding site motifs. Curr Opin Genet Dev 43:110–119. https://doi.org/10.1016/j.gde.2017.02.007
Jablonická V, Mansfeld J, Heilmann I et al (2016) Identification of a secretory phospholipase A2 from Papaver somniferum L. that transforms membrane phospholipids. Phytochemistry 129:4–13
Jia L, Clegg MT, Jiang T (2004) Evolutionary dynamics of the DNA-binding domains in putative R2R3-MYB genes identified from rice subspecies indica and japonica genomes. Plant Physiol 134(2):575–585
Kadereit JW (1988) Sectional affinities and geographical distribution in the genus Papaver L. (Papaveraceae). Beitrage zur Biologie der Pflanzen 63:139–156
Kakeshpour T, Nayebi S, Rashidi Monfared S et al (2015) Identification and expression analyses of MYB and WRKY transcription factor genes in Papaver somniferum L. Physiol Mol Biol Plants 21(4):465–478
Kreek MJ (2007) Opioids, dopamine, stress, and the addictions. Dialog Clin Neurosci 9(4):363–378
Kutchan TM, Weid M, Ziegler J (2004) The roles of latex and the vascular bundle in morphine biosynthesis in the opium poppy, Papaver somniferum. Proc Natl Acad Sci USA 101(38):13957–13962
Li Q, Zhang H (2017) Research progress on the synthesis of morphine alkaloids. Chin J Organ Chem 37(7):1629–1652
Liscombe DK, Facchini PJ (2008) Evolutionary and cellular webs in benzylisoquinoline alkaloid biosynthesis. Curr Opin Biotechnol 19(2):173–180
Marciano MA, Panicker SX et al (2018) Development of a method to extract opium poppy (Papaver somniferum L.) DNA from heroin. Sci Rep 8:2590. https://doi.org/10.1038/s41598-018-20996-9
Meos A, Saks L, Raal A (2017) Content of alkaloids in ornamental Papaver somniferum L. cultivars growing in estonia. Proc Est Acad Sci 66(1):34–39
Mishra S, Triptahi V, Singh S, Phukan UJ, Gupta MM, Shanker K, Shukla RK (2013) Wound induced tanscriptional regulation of benzylisoquinoline pathway and characterization of wound inducible PsWRKY transcription factor from Papaver somniferum. PLoS ONE 8(1):e52784. https://doi.org/10.1371/journal.pone.0052784
Mitrová K, Svoboda P, Milella L et al (2018) Alliinase and cysteine synthase transcription in developing garlic (Allium sativum L.) over time. Food Chem 15(251):103–109. https://doi.org/10.1016/j.foodchem.2017.12.090
Moloney MG (2016) Natural products as a source for novel antibiotics. Trends Pharmacol Sci 37(8):689–701
Onoyovwe A, Hagel JM, Chen X et al (2013) Morphine biosynthesis in opium poppy involves two cell types: sieve elements and laticifers. Plant Cell 25(10):4110–4122
Ovesná J, Kučera L, Vaculová K et al (2013) Analysis of the genetic structure of a barley collection using DNA diversity array technology (DArT). Plant Mol Bio Rep 31(2):280–288
Pathak S, Lakhwani D, Gupta P et al (2013) Comparative transcriptome analysis using high papaverine mutant of Papaver Somniferum reveals pathway and uncharacterized steps of papaverine biosynthesis. PLoS ONE 8(5):e65622. https://doi.org/10.1371/journal.pone.0065622
Prajapati S, Bajpai S, Singh D et al (2002) Alkaloid profiles of the Indian land races of the opium poppy Papaver somniferum L. Genet Resour Crop Evol 49(2):183–188
Prescha A, Grajzer M, Dedyk M et al (2014) The Antioxidant activity and oxidative stability of cold-pressed oils. J Am Oil Chem Soc 91(8):1291–1301
Russo D, Valentão P, Andrade PB et al (2015) Evaluation of antioxidant, antidiabetic and anticholinesterase activities of Smallanthus sonchifolius landraces and correlation with their phytochemical profiles. Int J Mol Sci 16(8):17696–17718
Saunders JM, Pedroni MJ, Penrose LDJ et al (2000) AFLP analysis of opium poppy. Crop Sci 41(5):1596–1601
Şelale H, Çelik I, Visam Gü V et al (2013) Development of EST-SSR markers for diversity and breeding studies in opium poppy. Plant Breed 132(3):344–351
Singh M, Chaturvedi N, Shasany AK et al (2014) Impact of promising genotypes of Papaver somniferum L. developed for beneficial uses. Acta Hortic 1036:29–41
Sofo A, Milella L, Tataranni G (2010) Effects of Trichoderma harzianum strain T-22 on the growth of two Prunus rootstocks during the rooting phase. J Hortic Sci Biotechnol 85(6):497–502
Solanki G, Dodiya NS, Kunwar R et al (2017) Character Association and path coefficient analysis for seed yield and latex yield in opium poppy (Papaver somniferum L.). Int J Cur Microbiol Appl Sci 6(8):1116–1123
Stefano GB, Pilonis N, Ptacek R et al (2017) Reciprocal evolution of opiate science from medical and cultural perspectives. Med Sci Monit 23:2890–2896
Tavakkoli Z, Assadi M (2016) Evaluation of seed and leaf epidermis characters in the taxonomy of some annual species of the genus papaver (Papaveraceae). Nord J Bot 34(3):302–321
Tétényi P (1997) Opium botany and horticulture. In: Janick J (ed) Horticultural reviews, vol 19. Wiley, New York, pp 375–382
Wang H, Wang H, Shao H et al (2016) Recent advances in utilizing transcription factors to improve plant abiotic stress tolerance by transgenic technology. Front Plant Sci 7:1–13
Winzer T, Gazda V, Zhesi H et al (2012) A papaver somniferum 10-gene cluster for synthesis of the anticancer alkaloid noscapine. Science 336(6089):1704–1708
Ziegler J, Facchini PJ, Rene G et al (2009) Evolution of morphine biosynthesis in opium poppy. Phytochemistry 70(15–16):1696–1707
Acknowledgements
Authors would like to thank the Czech Minstry of Agriculture, Projects NAZV QK1720263 and RO0417 for the support.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Labanca, F., Ovesnà, J. & Milella, L. Papaver somniferum L. taxonomy, uses and new insight in poppy alkaloid pathways. Phytochem Rev 17, 853–871 (2018). https://doi.org/10.1007/s11101-018-9563-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11101-018-9563-3