Abstract
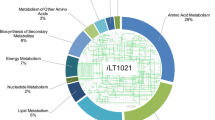
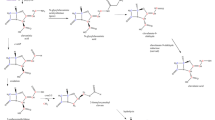
In the present work, a comprehensive metabolic network of Streptomyces roseosporus LC-54-20 was proposed for daptomycin production. The analysis of extracellular metabolites throughout the batch fermentation was evaluated in addition to daptomycin and biomass production. Metabolic flux distributions were based on stoichiometrical reaction as well as the extracellular metabolites fluxes. Experimental and calculated values for both the specific growth rate and daptomycin production rate indicated that the in silico model proved a powerful tool to analyze the metabolic behaviors based on the analysis under different initial glucose concentrations throughout the fermentation. Through manipulating different pH values, the production rates of various extracellular metabolites were also presented in this paper. Flux distribution variations revealed that the daptomycin production could be significantly influenced by the branch points of glucose 6-phosphate, 3-phosphoglycerate, phosphoenolpyruvate, pyruvate, and oxaloacetate. The five principal metabolites were certified as the flexible nodes and could form potential bottlenecks for a further enhancement of daptomycin production. Furthermore, various concentrations of the five precursors were added into the batch fermentation and led to the enhancement of daptomycin concentration and production rate.


Similar content being viewed by others
References
Eliopoulos, G. M. (2009). Microbiology of drugs for treating multiply drug-resistant Gram-positive bacteria. Journal of Infection, 59, s17–s24.
Nailor, M. D., & Sobel, J. D. (2009). Antibiotics for gram-positive bacterial infections: Vancomycin, teicoplanin, quinupristin/dalfopristin, oxazolidinones, daptomycin, dalbavancin, and telavancin. Infectious Disease Clinics of North America, 23, 965–982.
Hojati, Z., Milne, C., Harvey, B., Gordon, L., Borg, M., Flett, F., et al. (2002). Structure, biosynthetic origin, and engineered biosynthesis of calcium-dependent antibiotics from Streptomyces coelicolor. Chemistry & Biology, 9, 1175–1187.
Vertesy, L., Ehlers, E., Kogler, H., Kurz, M., Meiwes, J., Seibert, G., et al. (2000). Friulimicins: novel lipopeptide antibiotics with peptidoglycan synthesis inhibiting activity from Actinoplanes friuliensis sp. nov. II. Isolation and structural characterization. The Journal of Antibiotics, 53, 816–827.
Miao, V., Brost, R., Chapple, J., She, K., Gal, M. F., & Baltz, R. H. (2006). The lipopeptide antibiotic A54145 biosynthetic gene cluster from Streptomyces fradiae. Journal of Industrial Microbiology and Biotechnology, 33, 129–140.
Borders, D. B., Leese, R. A., Jarolmen, H., Francis, N. D., Fantini, A. A., Falla, T., et al. (2007). Laspartomycin, an acidic lipopeptide antibiotic with a unique peptide core. Journal of Natural Products, 70, 443–446.
Baltz, R. H. (2008). Biosynthesis and genetic engineering of lipopeptide antibiotics related to daptomycin. Current Topics in Medicinal Chemistry, 8, 618–638.
Alper, H., Jin, Y. S., Moxley, J. F., & Stephanopoulos, G. (2005). Identifying gene targets for the metabolic engineering of lycopene biosynthesis in Escherichia coli. Metabolic Engineering, 7, 155–164.
David, H., Ozçelik, I. S., Hofmann, G., & Nielsen, J. (2008). Analysis of Aspergillus nidulans metabolism at the genome-scale. BMC Genomics, 9, 163.
Boyle, N. R., & Morgan, J. A. (2009). Flux balance analysis of primary metabolism in Chlamydomonas reinhardtii. BMC Systems Biology, 3, 4.
Lee, S. J., Lee, D. Y., Kim, T. Y., Kim, B. H., Lee, J., & Lee, S. Y. (2005). Metabolic engineering of Escherichia coli for enhanced production of succinic acid, based on genome comparison and in silico gene knockout simulation. Applied and Environmental Microbiology, 71, 7880–7887.
Mukhopadhyay, A., Redding, A. M., Rutherford, B. J., & Keasling, J. D. (2008). Importance of systems biology in engineering microbes for biofuel production. Current Opinion in Biotechnology, 19, 228–234.
Gunnarsson, N., Eliasson, A., & Nielsen, J. (2004). Control of fluxes towards antibiotics and the role of primary metabolism in production of antibiotics. Advances in Biochemical Engineering/Biotechnology, 88, 137–178.
Borodina, I., Siebring, J., Zhang, J., Smith, C. P., van Keulen, G., Dijkhuizen, L., et al. (2008). Antibiotic overproduction in Streptomyces coelicolor A3(2) mediated by phosphofructokinase deletion. The Journal of Biological Chemistry, 283, 25186–25199.
Huber, F. M., Pieper, R. L., & Tietz, A. J. (1988). The formation of daptomycin by supplying decanoic acid to Streptomyces roseosporus cultures producing the antibiotic complex A21978C. Journal of Biotechnology, 7, 283–292.
Vallino, J. J., & Stephanopoulos, G. (1993). Metabolic flux distributions in Corynebacterium glutamicum during growth and lysine overproduction. Biotechnology and Bioengineering, 41, 633–646.
Vallino, J. J., & Stephanopoulos, G. (1994). Carbon flux distributions at the pyruvate branch point in Corynebacterium glutamicum during lysine overproduction. Biotechnology Progress, 10, 320–326.
Kim, H. B., Smith, C. P., Micklefield, J., & Mavituna, F. (2004). Metabolic flux analysis for calcium dependent antibiotic (CDA) production in Streptomyces coelicolor. Metabolic Engineering, 6, 313–325.
Blank, L. M., & Kuepfer, L. (2010). Metabolic flux distributions: Genetic information, computational predictions, and experimental validation. Applied Microbiology and Biotechnology, 86, 1243–1255.
Melzer, G., Dalpiaz, A., Grote, A., Kucklick, M., Göcke, Y., Jonas, R., et al. (2007). Metabolic flux analysis using stoichiometric models for Aspergillus niger: Comparison under glucoamylase-producing and non-producing conditions. Journal of Biotechnology, 132, 405–417.
Xu, H., Dou, W. F., Xu, H. Y., Zhang, X. M., Rao, Z. M., Shi, Z. P., et al. (2009). A two-stage oxygen supply strategy for enhanced l-arginine production by Corynebacterium crenatum based on metabolic fluxes analysis. Biochemical Engineering Journal, 43, 41–51.
Celik, E., Calik, P., & Oliver, S. G. (2010). Metabolic flux analysis for recombinant protein production by Pichia pastoris using dual carbon sources: Effects of methanol feeding rate. Biotechnology and Bioengineering, 105, 317–329.
Sauer, U., & Eikmanns, B. J. (2005). The PEP-pyruvate-oxaloacetate node as the switch point for carbon flux distribution in bacteria. FEMS Microbiology Reviews, 29, 765–794.
Gao, H. J., Du, G. C., & Chen, J. (2006). Analysis of metabolic fluxes for hyaluronic acid (HA) production by Streptococcus zooepidemicus. World Journal of Microbiology and Biotechnology, 22, 399–408.
Nie, Z. K., Ji, X. J., Huang, H., Du, J., Li, Z. Y., Qu, L., et al. (2011). An effective and simplified fed-batch strategy for improved 2,3-butanediol production by Klebsiella oxytoca. Applied Biochemistry and Biotechnology, 163, 946–953.
Ji, X. J., Huang, H., Li, S., Du, J., & Lian, M. (2008). Enhanced 2,3-butanediol production by altering the mixed acid fermentation pathway in Klebsiella oxytoca. Biotechnology Letters, 30, 731–734.
Miao, V., Coëffet-Legal, M. F., Brian, P., Brost, R., Penn, J., Whiting, A., et al. (2005). Daptomycin biosynthesis in Streptomyces roseosporus: Cloning and analysis of the gene cluster and revision of peptide stereochemistry. Microbiology, 151, 1507–1523.
Yu, G., Jia, X., Wen, J., Lu, W., Wang, G., Caiyin, Q., et al. (2011). Strain improvement of Streptomyces roseosporus for daptomycin production by rational screening of He-Ne laser and NTG induced mutants and kinetic modeling. Applied Biochemistry and Biotechnology, 163, 729–743.
Varma, A., & Palsson, B. O. (1994). Stoichiometric flux balance models quantitatively predict growth and metabolic by-product secretion in wild-type Escherichia coli W3110. Applied and Environmental Microbiology, 60, 3724–3731.
Gheshlaghi, R., Scharer, J. M., Moo-Young, M., & Douglas, P. L. (2007). Metabolic flux analysis for optimizing the specific growth rate of recombinant Aspergillus niger. Bioprocess and Biosystems Engineering, 30, 397–418.
Madden, T., Ward, J. M., & Ison, A. P. (1996). Organic acid excretion by Streptomyces lividans TK24 during growth on defined carbon and nitrogen sources. Microbiology, 142, 3181–3185.
Naeimpoor, F., & Mavituna, F. (2000). Metabolic flux analysis in Streptomyces coelicolor under various nutrient limitations. Metabolic Engineering, 2, 140–148.
Pons, A., Dussap, C. G., Péquignot, C., & Gros, J. B. (1996). Metabolic flux distribution in Corynebacterium melassecola ATCC 17965 for various carbon sources. Biotechnology and Bioengineering, 51, 177–189.
Fürch, T., Wittmann, C., Wang, W., Franco-Lara, E., Jahn, D., & Deckwer, W. D. (2007). Effect of different carbon sources on central metabolic fluxes and the recombinant production of a hydrolase from Thermobifida fusca in Bacillus megaterium. Journal of Biotechnology, 132, 385–394.
Rokem, J. S., Lantz, A. E., & Nielsen, J. (2007). Systems biology of antibiotic production by microorganisms. Natural Product Reports, 24, 1262–1287.
van Gulik, W. M., de Laat, W. T., Vinke, J. L., & Heijnen, J. J. (2000). Application of metabolic flux analysis for the identification of metabolic bottlenecks in the biosynthesis of penicillin-G. Biotechnology and Bioengineering, 68, 602–618.
Acknowledgments
The authors wish to acknowledge the financial support provided by the National 973 Project of China (No. 2011CB710800), the Key Program of National Natural Science Foundation of China (Grant No. 20936002), National Natural Science Foundation of China (No. 21076022).
Author information
Authors and Affiliations
Corresponding author
Electronic Supplementary Material
Below is the link to the electronic supplementary material.
ESM 1
(DOC 162 kb)
Rights and permissions
About this article
Cite this article
Huang, D., Jia, X., Wen, J. et al. Metabolic Flux Analysis and Principal Nodes Identification for Daptomycin Production Improvement by Streptomyces roseosporus . Appl Biochem Biotechnol 165, 1725–1739 (2011). https://doi.org/10.1007/s12010-011-9390-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12010-011-9390-0