Abstract
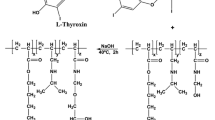
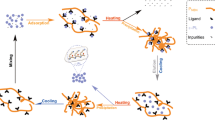
A microbial transglutaminase (MTG) was efficiently purified by using pH-responsive affinity precipitation with Crocein orange G (COG) as the affinity ligand. The docking method was used to identify the appropriate ligand. The molecular simulation results were compared with the label-free detection data analyzed by ForteBio’s Octet. A pH-responsive polymer, PMMDN, was polymerized and subsequently coupled with COG as the ligand. The isoelectric point (pI) and recovery of PMMDN and PMMDN–COG were 4.51, 99.8,% and 4.78, 98.1 %, respectively. The optimal adsorption conditions were found to be a ligand density of 60.0 μmol/g, pH 7.0, and 0.2 mol/L NaCl. The adsorption isotherm showed that the maximum adsorption capacity was 91.32 mg/g polymer and the dissociation constant was 0.021 mg/mL. Interaction information between PMMDN–COG and MTG in the whole process of affinity precipitation were obtained by Fourier transform infrared spectroscopy (FT-IR) and scanning electron microscopy (SEM). PB (0.02 mol/L pH 7.0) in 20.0 % glycol was used to elute the binding MTG from PMMDN–COG. Under these conditions, electrophoretically pure MTG was obtained in only one step with elution recoveries of 98.96 % (protein) and 95.09 % (activity).







Similar content being viewed by others
References
Ando, H., Adachi, M., Umeda, K., Matsuura, A., Nonaka, M., Uchio, R., Tanaka, H., & Motoki, M. (1989). Purification and characteristics of a novel transglutaminase derived from microorganisms. Agricultural and Biological Chemistry, 53, 2613–2617.
Bourneow, C. and Benjakul, S. (2011) Purification and characterization of microbial transglutaminase from Enterobacter sp. C2361. Thai Journal of Agricultural Science, 496–504. Agricultural Science Society of Thailand.
Brozell, S., Mukherjee, S., Balius, T., Roe, D., Case, D., & Rizzo, R. (2012). Evaluation of DOCK 6 as a pose generation and database enrichment tool. J Computer Aided Molecular Design, 26, 749–773.
Chang, R. (2000) Physical chemistry for the chemical and biological sciences.ed. University Science Books.
Costioli, M. D., Fisch, I., Garret-Flaudy, F., Hilbrig, F., & Freitag, R. (2003). DNA purification by triple-helix affinity precipitation. Biotechnology and Bioengineering, 81, 535–545.
Cuatrecasas, P., & Anfinsen, C. B. (1971). Affinity chromatography. Annual Review of Biochemistry, 40, 259–278.
Denizli, A., & Pişkin, E. (2001). Dye-ligand affinity systems. Journal of Biochemical Biophysical Methods, 49, 391–416.
Ding, Z., & Cao, X. (2013). Affinity precipitation of cellulase using pH-response polymer with Cibacron Blue F3GA. Separation and Purification Technology, 102, 136–141.
Ding, Z., Kang, L., & Cao, X. (2014). Application of docking methods for metal chelate affinity precipitation of endo-glucanase using pH-response polymer. Colloids and Surfaces B: Biointerfaces, 113, 412–420.
Ding, Z., Zheng, K., & Cao, X. (2014). Lipase purification by affinity precipitation with a thermo-responsive polymer immobilized Cibacron Blue F3GA ligand. Biotechnology and Bioprocess Engineering, 19, 892–899.
El-Hofi, M., Ismail, A., & Maher Nour, O. I. (2014). Isolation, purification and characterization of transglutaminase from rosemary (Rosmarinus officinalis l.) Leaves. Acta Scientairum Polonorum, Technologia Alimentaria, 13, 267–278.
Ewing, T. J. A., & Kuntz, I. D. (1997). Critical evaluation of search algorithms for automated molecular docking and database screening. Journal of Computational Chemistry, 18, 1175–1189.
Gerber, U., Jucknischke, U., Putzien, S., & Fuchsbauer, H.-L. (1994). A rapid and simple method for the purification of transglutaminase from Streptoverticillium mobaraense. The Biochemical Journal, 299, 825.
Grossowicz, N., Wainfan, E., Borek, E., & Waelsch, H. (1950). The enzymatic formation of hydroxamic acids from glutamine and asparagine. The Journal of Biological Chemistry, 187, 111–125.
Gupta, M. N., Kaul, R., Guoqiang, D., Dissing, U., Mattiasson, B., & Scouten, W. H. (1996). Affinity precipitation of proteins. Journal of Molecular Recognition, 9, 356–359.
Kumar, A., Khalil, A. A. M., Galaev, I. Y., & Mattiasson, B. (2003). Metal chelate affinity precipitation: purification of (his)6-tagged lactate dehydrogenase using poly(vinylimidazole-co-N-isopropylacrylamide) copolymers. Enzyme and Microbial Technology, 33, 113–117.
Kuntz, I. D., Blaney, J. M., Oatley, S. J., Langridge, R., & Ferrin, T. E. (1982). A geometric approach to macromolecule-ligand interactions. Journal of Molecular Biology, 161, 269–288.
Laemmli, U. K. (1970). Cleavage of structural proteins during the assembly of the head of bacteriophage T4. Nature, 227, 680–685.
Lang, P. T., Brozell, S. R., Mukherjee, S., Pettersen, E. F., Meng, E. C., Thomas, V., Rizzo, R. C., Case, D. A., James, T. L., & Kuntz, I. D. (2009). DOCK 6: combining techniques to model RNA-small molecule complexes. RNA (New York, N.Y.), 15, 1219–1230.
Liu, Y., Xiao, L., Joo, K.-I., Hu, B., Fang, J., & Wang, P. (2014). In situ modulation of dendritic cells by injectable thermosensitive hydrogels for cancer vaccines in mice. Biomacromolecules, 15, 3836–3845.
Macedo, J. A., Sette, L. D., & Sato, H. H. (2011). Purification and characterization of a new transglutaminase from Streptomyces sp. Isolated in Brazilian soil. Journal of Food Biochemistry, 35, 1361–1372.
Memmedova, T., Armutcu, C., Uzun, L., & Denizli, A. (2015). Polyglycidyl methacrylate based immunoaffinity cryogels for insulin adsorption. Materials Science and Engineering: C, 52, 178–185.
Meng, E. C., Shoichet, B. K., & Kuntz, I. D. (1992). Automated docking with grid-based energy evaluation. Journal of Computational Chemistry, 13, 505–524.
Motoki, M., & Seguro, K. (1998). Transglutaminase and its use for food processing. Trends if Food Science and Technology, 9, 204–210.
Pencheva, T., Soumana, O. S., Pajeva, I., & Miteva, M. A. (2010). Post-docking virtual screening of diverse binding pockets: comparative study using DOCK, AMMOS, X-Score and FRED scoring functions. European Journal of Medicinal Chemistry, 45, 2622–2628.
Pettersen, E. F., Goddard, T. D., Huang, C. C., Couch, G. S., Greenblatt, D. M., Meng, E. C., & Ferrin, T. E. (2004). UCSF Chimera—a visualization system for exploratory research and analysis. Journal of Computational Chemistry, 25, 1605–1612.
Porath, J., Carlsson, J. A. N., Olsson, I., & Belfrage, G. (1975). Metal chelate affinity chromatography, a new approach to protein fractionation. Nature, 258, 598–599.
Sheth, R. D., Madan, B., Chen, W., & Cramer, S. M. (2013). High-throughput screening for the development of a monoclonal antibody affinity precipitation step using ELP-z stimuli responsive biopolymers. Biotechnology and Bioengineering, 110, 2664–2676.
Shi, Y. G., Qian, L., Zhang, N., Han, C. R., Liu, Y., Zhang, Y. F., & Ma, Y. Q. (2012). Study on separation and purification of the transglutaminase. Applied Mechanics and Materials, 121, 443–447.
Strop, P. (2014). Versatility of microbial transglutaminase. Bioconjugate Chemistry, 25, 855–862.
Subramanian, S. and Ross, P. D. (1984) Dye-ligand affinity chromatography: the interaction of Cibacron Blue F3GA® with proteins and enzyme. Critical Reviews Biochemistry Molecular,16, 169-205.
Totrov, M., & Abagyan, R. (2008). Flexible ligand docking to multiple receptor conformations: a practical alternative. Current Opinion in Structural Biology, 18, 178–184.
Vaidya, A. A., Lele, B. S., Deshmukh, M. V., & Kulkarni, M. G. (2001). Design and evaluation of new ligands for lysozyme recovery by affinity thermoprecipitation. Chemical Engineering Science, 56, 5681–5692.
Wilhelm, B., Meinhardt, A., & Seitz, J. (1996). Transglutaminases: purification and activity assays. Journal of Chromatography B: Biomedical Sciences and Applications, 684, 163–177.
Yan, B., & Cao, X. (2012). Preparation of aqueous two-phase systems composed of two pH-response polymers and liquid–liquid extraction of demeclocycline. Journal of Chromatography. A, 1245, 39–45.
Yılmaz, M., Bayramoǧlu, G., & Arıca, M. Y. (2005). Separation and purification of lysozyme by Reactive Green 19 immobilized membrane affinity chromatography. Food Chemistry, 89, 11–18.
Yokoyama, K., Nio, N., & Kikuchi, Y. (2004). Properties and applications of microbial transglutaminase. Appl Microbiol Biot, 64, 447–454.
Zhu, Y., & Tramper, J. (2008). Novel applications for microbial transglutaminase beyond food processing. Trends in Biotechnology, 26, 559–565.
Author information
Authors and Affiliations
Corresponding author
Glossary
- AM1-BCC
-
Semi-empirical (AM1) with bond charge correction (BCC)
- ANTECHAMBER
-
Antechamber is a set of auxiliary programs for molecular mechanical studies
- DMS
-
DMS is an open-source program written in C for computing the molecular surface of a molecule
- GRID
-
Grid creates the grid files necessary for rapid score evaluation in DOCK
- Mol2
-
The Mol2 file format is a popular method for specifying chemical structures, including atom types, positions, and bonding
- SHOWBOX
-
Showbox is an interactive program for specifying the location and the size of the grids that will be calculated by the program grid
- sphere_selector
-
Sphere_selector filters the output from sphgen selecting all spheres within a user-defined radius of a target molecule
- SPHGEN
-
Sphgen generates sets of overlapping spheres to describe the shape of a molecule or molecular surface
- sphgen_cpp
-
Sphgen_cpp has the same function as Sphgen
Rights and permissions
About this article
Cite this article
Ding, Z., Li, S. & Cao, X. Microbial Transglutaminase Separation by pH-Responsive Affinity Precipitation with Crocein Orange G as the Ligand. Appl Biochem Biotechnol 177, 253–266 (2015). https://doi.org/10.1007/s12010-015-1742-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12010-015-1742-8